Published online Dec 14, 2017. doi: 10.3748/wjg.v23.i46.8217
Peer-review started: August 5, 2017
First decision: August 30, 2017
Revised: September 13, 2017
Accepted: November 7, 2017
Article in press: November 7, 2017
Published online: December 14, 2017
Processing time: 131 Days and 16.6 Hours
To explore the possible relationship between fecal microbial communities and non-anastomotic stricture (NAS) after liver transplantation (LT).
A total of 30 subjects including 10 patients with NAS, 10 patients with no complications after LT, and 10 non-LT healthy individuals were enrolled. Fecal microbial communities were assessed by the 16S rRNA gene sequencing technology.
Different from the uncomplicated and healthy groups, unbalanced fecal bacterium ratio existed in patients with NAS after LT. The results showed that NAS patients were associated with a decrease of Firmicutes and Bacteroidetes and an increase of Proteobacteria at the phylum level, with the proportion-ratio imbalance between potential pathogenic families including Enterococcaceae, Streptococcaceae, Enterobacteriaceae, Pseudomonadaceae and dominant families including Bacteroidaceae.
The compositional shifts of the increase of potential pathogenic bacteria as well as the decrease of dominant bacteria might contribute to the incidence of NAS.
Core tip: This study is the first attempt to investigate the possible relationship between gut microbiota and post-liver transplantation (LT) biliary complication based on the 16S rRNA sequencing technology. Our results showed unbalanced ratio of pathogenic bacteria to dominant bacteria really existed in patients with non-anastomotic stricture after LT. The shifts of fecal microbial communities may be involved in or exacerbate the process of bile duct injury, which may contribute to the mechanism research and prevention in future.
- Citation: Zhang J, Ren FG, Liu P, Zhang HK, Zhu HY, Feng Z, Zhang XF, Wang B, Liu XM, Zhang XG, Wu RQ, Lv Y. Characteristics of fecal microbial communities in patients with non-anastomotic biliary strictures after liver transplantation. World J Gastroenterol 2017; 23(46): 8217-8226
- URL: https://www.wjgnet.com/1007-9327/full/v23/i46/8217.htm
- DOI: https://dx.doi.org/10.3748/wjg.v23.i46.8217
As Thomas Starzl performed the first human liver transplantation in 1963, orthotopic liver transplantation (OLT) has been regarded as the standard therapy for patients with end-stage liver diseases. In the past three decades, the postoperative complications of OLT decreased markedly due to the improvement of surgical techniques and immunosuppressive treatment[1,2]. However, the morbidity of biliary stricture after OLT is still high, ranging from 5% to 20%[3]. Non-anastomotic stricture (NAS), also known as ischemic type biliary stricture, is a lethal complication for recipients and severely affects their long-term prognosis[4]. Factors including poor liver graft, ABO-incompatibility, cytomegalovirus (CMV) infection may contribute to the development of NAS, and ischemic reperfusion related inflammatory injury is commonly regarded as an inducer of this pathologic process[5-8]. But up to date, the definite mechanisms of NAS remain unknown.
Gut microbiota is the general term for all microorganisms (mainly for bacteria) living in the human intestine, with a microbial density larger than 1014 cells/g, containing 100 times more genes than human’s[9,10]. Current studies have titled the gut bacteria as another human organ for its enormous influences on human metabolic activity, barrier function, and immunity development. However, endotoxemia caused by dysbacteriosis was also connected to obesity, diabetes, nonalcoholic fatty liver diseases (NAFLD), and autoimmune disorders[11,12], and even played a key role in ischemic reperfusion injury[13]. While for patients who underwent liver transplantation, complex factors like portal vein blocking, ischemic reperfusion injury, antibiotics or immunosuppression use can seriously impair recipient’s immune function, destroy the intestinal barrier, and finally increase the risk of dysbacteriosis. These changes of microbiota may directly injury host liver parenchyma through the “gut-liver” axis[14]. Actually, the relationship between dysbacteriosis and postoperative complications including acute rejection, early-stage infection, and graft loss is under investigation[15,16]. To account for all of these, we hypothesized that quantitative or qualitative alterations of gut microbiota may be involved in or exacerbate graft’s ischemic reperfusion injury, which eventually leads to NAS. But so far, the detailed relationship between them has never been explored. Furthermore, whether the changes of gut microbiota contribute to the occurrence of NAS after OLT is still obscure.
In this study, we explored the potential relationship between gut microbiota and NAS by investigating the changes in microbial communities in patients diagnosed with NAS.
All subjects in this study came from the First Affiliated Hospital of Xi’an Jiaotong University, with no history of the use of systemic antibiotics or probiotics within previous 3 mo. We excluded patients accompanied by other digestive comorbidities, autoimmune disorders, NAFLD, obesity, or diabetes mellitus, and those who suffered from diarrhea or constipation within 1 mo were not included either. Patients with NAS were defined as suffering from repeated cholangitis, and the magnetic resonance cholangio-pancreatography (MRCP) or endoscopic retrograde cholangio-pancreatography (ERCP) results suggesting multiple strictures located in the donor biliary system with/without anastomotic stricture. To eliminate arterial factors, those accompanied with hepatic artery thrombosis were not included. For patients in an uncomplicated group, they had no obvious complications after OLT, and the regular reexaminations (symptoms, physical examinations, B-ultrasound, CT scan, biochemical tests, and plasma concentration of immunosuppressive drugs) were normal. The healthy controls were those non-LT individuals who came to hospital for a routine health examination, with no digestive diseases or surgical history and their routine tests indexes were in normal ranges. Finally, a total of 30 patients meeting the inclusion criteria were enrolled, including 20 post-LT patients (10 in the NAS group and 10 in the uncomplicated group) and 10 healthy controls.
All participants were totally informed of the related matters prior to entering in and signed the informed consent form. This study was performed in accordance with the ethical guidelines of the 1975 Declaration of Helsinki and was approved by the institutional review board of the First Affiliated Hospital of Xi’an Jiaotong University.
All post-LT patients underwent OLT at the First Affiliated Hospital of Xi’an Jiaotong University. Organ donation or transplantation in this study was strictly implemented under the regulation of the China Organ Donation Committee (CODC), Organ Transplant Committee (OTC), and the Declaration of Helsinki. Recipients were carefully evaluated before operation, while candidates diagnosed with hepatocellular carcinoma (HCC) totally accorded with the Milan criteria[17]. Operations were performed with an ABO-compatible liver graft by the same group of doctors. All grafts derived from donors of cardiac death (DCD) and preserved in University of Wisconsin solution at 4 °C before LT. During the operation, graft’s common bile duct were bonded to recipient’s by means of duct to duct anastomosis, interruptedly suturing for the anterior wall and continuously for the posterior wall with 6-0 absorbable strings. A T-tube was applied just as necessarily required. After operation, they were given the triple regimen anti-rejection therapy consisting of tacrolimus, mycophenolate mofetil, and methylprednisone.
We documented individual’s basic characteristics, including age, gender, body mass index (BMI), current state of smoking or drinking, blood routine test, and liver function indexes within 48 h before sample collecting. For post-LT patients, graft related factors (warm and cold ischemic time) and perioperative characteristics (including Child-Pugh classification, total duration of operation, anhepatic phase, bleeding volume, T-tube inserted or not) were reviewed. The duration from LT to diagnosis and the duration from LT to sample collecting were also respectively recorded.
All fecal samples were carefully collected to avoid the pollution by urine, accurately weighed, sub-packaged into a 2 mL micro-centrifuge tube (180-200 mg per tube), and immediately stored at -80 °C before analysis. All these stages were finished within 30 min.
The fecal DNA was extracted according to the manufacturer’s instructions of a testing kit (QIAamp DNA Stool Mini Kit, Qiagen, Valencia, CA, United States). For one aliquot, a little bit of stool was scraped into a 2 mL microcentrifuge tube on ice, and 1.4 mL of buffer ASL (from the QIAamp DNA Stool Mini Kit) was added before the sample thawed. The tube was then vortexed continuously for 1 min until the sample was thoroughly homogenized. After incubation in a water bath for 5 min at 70 °C, the tube was vortexed for 15 s and centrifuged at 2000 g for 1 min. The sediment was then discarded, and 1.2 mL of the supernatant was pipetted into a new 2 mL microcentrifuge tube. An inhibitEX tablet (from the kit) was added and vortexed for 1 min until the tablet was completely suspended. After incubation of the suspension for 1 min at room temperature and centrifugation for 3 min, all the supernatant was pipetted into a new 1.5 mL microcentrifuge tube and centrifuged for 3 min. Above 200 μL supernatant was pipetted into a new 1.5 mL microcentrifuge tube which had already contained 15 μL proteinase K. Then, 200 μL of Buffer AL (from the kit) was added, and the tube was vortexed for 15 s and incubated at 70 °C for 10 min. Following the addition of 200 μL of anhydrous ethanol to the lysate, the tube was vortexed thoroughly. Subsequently, the lysate was carefully applied to the QIAamp spin column. After centrifugation for 1 min, the QIAamp spin column was transferred into a new 2 mL collection tube, and the tube containing filtrate was discarded. Then, 500 μL of Buffer AW1 (from the kit) was added. After centrifugation for 1 min and discarding the filtrate, 500 μL of Buffer AW2 (from the kit) was added. Following centrifugation for 3 min, the spin column was placed into a new 2 mL collection tube and centrifuged for 1 min. The spin column was transferred into a new 1.5 mL tube, and 200 μL of Buffer AE was pipetted onto the QIAamp membrane. The tube was incubated at room temperature for 1 min and then centrifuged for 1 min to elute DNA. Finally, the filtrate (containing DNA) was stored at -20 °C.
The DNA isolated from fecal samples was used as the template for the amplification of the 16S rRNA V3-V4 region. The universal primers used were F (5’-NNNNNNN ACTCCTACGGGAGGCAGCA-3’) and R (5’-NNNNNNN GGACTACVSGGGTATCTAAT-3’), with the NNNNNNN being unique seven-base barcode used to tag each PCR product. The PCR reaction was performed according to the touchdown protocol[18] in a system of 25 μL containing 5.0 μL 5 × reaction buffer (TaKaRa, Dalian, China), 5.0 μL 5 × high GC buffer (TaKaRa, Dalian, China), 0.5 μL dNTPs (10 mmol/L) mixture , 1.0 µL forward primer (10 µmol/L), 1.0 μL reverse primer (10 µmol/L), 0.25 μL Q5 high-fidelity DNA polymerase (5 U/uL, TaKaRa, Dalian, China), and 1 μL DNA template. Each PCR product was purified by 2% agarose gel electrophoresis. DNA was isolated using the Axygen Axy Prep DNA Gel Extraction kit (Axygen, Shanghai, China). The sequencing was finished with the help of the Illumina Miseq System (Illumina).
The sequencing data of samples were analyzed using pyrosequencing pipeline tools at RDP 10 (http://pyro.cme.msu.edu/). Bacterial diversity was determined by sampling-based analysis of operational taxonomic units (OTUs), α-diversity index (including rarefaction curves, Chao1 index, ACE index, Shannon index, and Simpson index, estimated at a distance of 5%), as well as principal component analysis (PCA). The OTU is an operational definition referring to those closely related individuals, in the system of biological classification, and it is defined based on a similarity threshold to classify microbial species into different taxonomic levels (97% similarity equal to the level of species)[19,20]. Species accumulation curve is applied to assess species richness based on the results of species and individual sampling. It can only be compared when the species richness has reached a clear asymptote[21]. PCA is mathematically defined as an orthogonal linear transformation which transforms the original data to a new system defined as principal component. Hence, the greatest variance by some projection of the data comes to lie on the corresponding principal component, which makes it easier to investigate the correlation between multiple variables[22].
Diversity indexes and the species accumulation curve were calculated by QIIME. PCA plots of the bacterial communities were created using pcaMethods (Stacklies et al, 2007) in R (R Development Core Team, 2012). Differences of categorical variables among groups were analyzed by Chi-square or Fisher’s exact test, and final results are expressed as percentage (%). For continuous variables, ANOVA test was used if data met the normal distribution or Mann-Whitney test if not, with corresponding results expressed as mean ± SD or median (range). Statistical analyses were performed with SPSS version 18.0 (SPSS Inc., Chicago, IL, United States). P-values < 0.05 were considered statistically significant.
As Table 1 shows, patients in the three groups shared the similar age distribution, gender proportion, and BMI (P > 0.05 for all). Results of blood routine tests were generally in normal ranges and showed no differences among the groups (P > 0.05, Table 1). While for liver function, all median or mean values were obviously abnormal for patients diagnosed with NAS, but no differences existed between the uncomplicated and healthy control groups. Notably, for patients with NAS, biliary tract associated indexes like ALP and GGT were elevated as nearly 4 times as healthy controls’ (P < 0.05), while ALB level was seriously decreased with a mean value of 34.14 g/L (41.1 g/L for healthy and 41.9 g/L for uncomplicated, P < 0.05).
| Healthy,(n = 10) | Post-LT | ||
| Uncomplicated, (n = 10) | NAS,(n = 10) | ||
| Age (yr) | 38 ± 12 | 43 ± 11 | 42 ± 9 |
| Male | 9 (90.0) | 8 (80.0) | 8 (80.0) |
| BMI (kg/m2) | 23.3 ± 2.5 | 22.1 ± 2.6 | 22.4 ± 2.7 |
| Current smoking | 3 (30.0) | 2 (10.0) | 0 |
| Current drinking | 2 (20.0) | 0 | 0 |
| Blood routine test | |||
| HB (g/L) | 122.5 ± 12.7 | 129.0 ± 20.0 | 127.4 ± 9.0 |
| WBC (×109) | 6.0 ± 1.7 | 5.1 ± 2.2 | 5.2 ± 2.5 |
| Neu (%) | 59.6 ± 14.8 | 66.4 ± 16.4 | 64.3 ± 20.0 |
| Liver function | |||
| AST (U/L) | 21.3 (7.9-39.6) | 41.0 (13.0-93.0) | 57.1 (17.0-107.0)ac |
| ALT (U/L) | 20.1 (14.6-34.4) | 49.3 (12.0-89.1) | 57.3 (18.0-111.0)ac |
| ALP (U/L) | 77.3 ± 31.7 | 93.9 ± 17.2 | 332.8 ± 52.4ac |
| GGT (U/L) | 27.2 ± 8.2 | 53.3 ± 35.6 | 226.4 ± 83.4ac |
| TB (μmol/L) | 13.7 ± 6.7 | 27.4 ± 17.6 | 104.43 ± 47.8ac |
| DB (μmol/L) | 5.4 ± 3.1 | 12.5 ± 8.6 | 43.8 ± 6.8ac |
| ALB (g/L) | 41.1 ± 2.9 | 41.9 ± 5.3 | 34.1 ± 5.0ac |
For all patients who underwent LT, the main inducers were HBV-related cirrhosis (80.00% vs 80.00%, P = 0.568, Table 2), and others including subacute liver failure (SALF), hepatocellular carcinoma (HCC), and drug-induced liver injury (DILI) were relatively few in this study. Distributions of preoperative Child-Pugh scores between two groups were similar also, with the percentage of patients having Child-Pugh A or B were 50% vs 50% (P = 0.834, Table 2). In addition, other factors such as liver grafts’ ischemic time, the mean duration of anhepatic phase, total operation duration, intraoperative bleeding volume, and the proportion of T-tube application were all equally distributed (P > 0.05 for all, Table 2). The median duration from LT to final diagnosis of NAS was 9 months, and those from LT to sample collecting in two post-LT groups were 21 and 15 months, respectively (P = 0.129).
| Uncomplicated, (n = 10) | NAS,(n = 10) | P value | |
| Primary disease | |||
| HBV cirrhosis | 8 (80.0) | 8 (80.0) | 0.568 |
| HBV SALF | 0 (0.0) | 1 (10.0) | |
| HCC | 1 (10.0) | 1 (10.0) | |
| DILI | 1 (10.0) | 0 (0.0) | |
| Child-Pugh classification | |||
| A | 1 (10.0) | 1 (10.0) | 0.834 |
| B | 4 (40.0) | 4 (40.0) | |
| C | 5 (50.0) | 5 (50.0) | |
| WIT (min) | 7 ± 2 | 8 ± 0 | 0.108 |
| CIT (h) | 7 ± 1 | 6 ± 1 | 0.291 |
| Total operation duration (min) | 366 ± 80 | 377 ± 62 | 0.893 |
| Anhepatic phase (min) | 46 ± 10 | 49 ± 7 | 0.513 |
| Bleeding Volume (mL) | 1760 ± 347 | 1311 ± 268 | 0.329 |
| T-tube insertion | 8 (80.00) | 7 (70.00) | 0.906 |
| Median time from LT to NAS (m) | - | 9 (5-13) | - |
| Median time from LT to SC (m) | 15 (6-36) | 21 (13-32) | 0.129 |
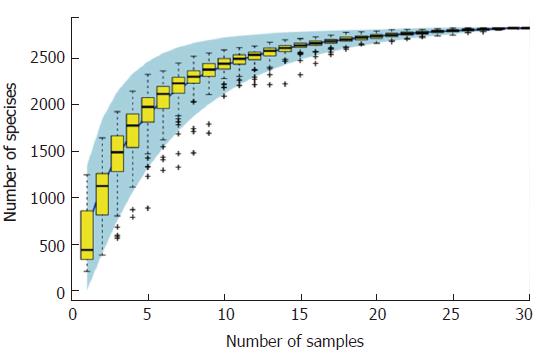
According to the sample number and species OTUs, we calculated the species accumulation curve of all participants (Figure 1). In this study, the curve had reached a plateau, and the species had no more obvious increase as the sample number increased, which indicated that the sample volume in our study was relatively large enough to reflect the species richness.
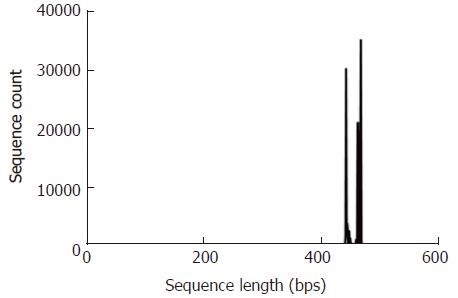
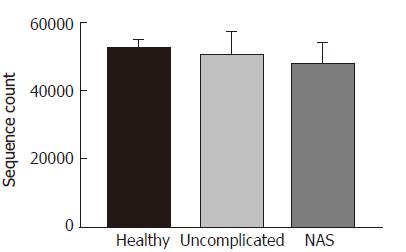
To ensure the validity, we excluded those rare OTUs of which the richness was less than 0.001% of the total, and also took a flattening process to eliminate the bias of sequencing depth. Finally, we got a total of 1,494,713 valid sequences, with an average sequence length of 468 bps. For these three groups, the mean valid sequence numbers were 52222, 49947, and 47302, respectively (P > 0.05, Figures 2 and 3).
As for the microbial community diversity, the OTUs number at the phylum level in the healthy control group was 969 ± 43, while in the two post-LT groups, the numbers were 443 ± 75 and 568 ± 122, respectively, obviously smaller than that of healthy controls (P < 0.05 for both, Table 3). It seemed that there were more OTUs in the NAS group than in the uncomplicated group, but the difference was not significant. Similarly, these manifestations were also applicable to the OTUs distributions at the order/family/genus/species levels (Table 3). Meanwhile, both two post-LT groups showed smaller α-diversity index (including Chao1, ACE, Simpson, and Shannon indexes) than the healthy controls (P < 0.01, Table 4). All of these indicated that patients who underwent LT had a lower gut microbiota diversity (including richness and species number) than healthy controls. Furthermore, despite no significant differences, gut microbiota of patients with NAS after LT was more diverse than that of the uncomplicated group. We surmise that it was mainly due to the increase of potentially pathogenic bacteria (details will be described later).
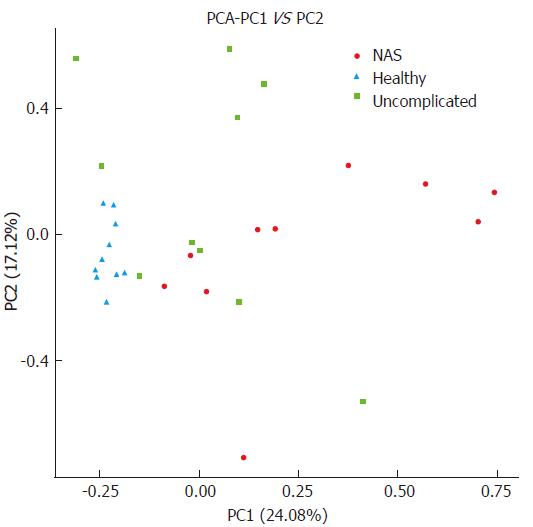
About the PCA of different groups, the healthy controls were shown to well aggregate and not overlap with the two post-LT groups. Post-LT individuals of the two groups were partially overlapped, but they still had their own trend to aggregate separately. Therefore, we can still distinguish the NAS cluster from the uncomplicated group (Figure 4). Collectively, we can conclude that the variation among groups was larger than that within groups, and clustering in our study was actually feasible (PC1 = 24.08%, PC2 = 17.12%).
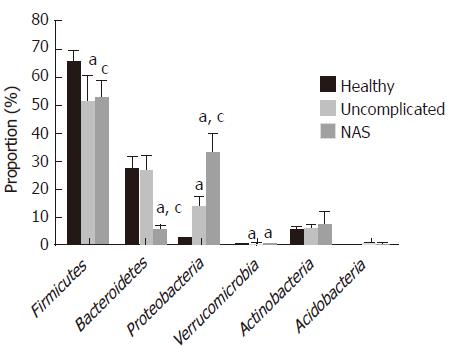
As shown in Figure 5, gut microbiota in this study was mainly composed of six phyla, including Firmicutes, Bacteroidetes, Proteobacteria, Actinobacteria, Acidobacteria, and Verrucomicrobia. Firmicutes and Bacteroidetes, as the main bacteria coexisting in human intestine, contributed to 92.32% of the total microbiota in the healthy control group, while the proportions were 77.11% in the uncomplicated group and 57.40% in the NAS group, which were significantly smaller than that of healthy controls (P < 0.05 for both). Specifically, the change of Firmicutes in post-LT patients was mostly due to the decrease of Lachnospiraceae and Ruminococcaceae at the family level, accompanied by the increase of Enterococcaceae and Streptococcaceae (all owned to Bacilli class, Table 5). Especially for the NAS group, the proportions of the latter two were significantly larger than those in the uncomplicated group (2.60% vs 1.20%, 8.60% vs 3.90%, P < 0.05 for both, Table 5). For Bacteroidetes, uncomplicated patients after LT shared the similar proportion to the healthy group (P > 0.05). While further analyzing, this phenomenon was caused by the increase of Bacteroidaceae and equivalent decrease of Prevotellaceae at the family level. However, phylum of Bacteroidetes was substantially decreased in the NAS group, with a constituent ratio of only 5.11%, nearly one fifth of that in the healthy group (P < 0.05, Table 5). The decrease of Bacteroidaceae and Prevotellaceae at the family level played the inducing role in this change, from the normal 11.60% and 11.60% to 2.70% and 0.70%, respectively (P < 0.05 for both, Table 5). As for the phylum of Proteobacteria, it increased obviously in the two post-LT groups, especially for patients with NAS, in whom the proportion of Proteobacteria was up to nearly 30 times than that in the healthy group (32.44% ± 7.32% vs 1.99% ± 0.25%, P < 0.05, Figure 5). The proportions of family of Enterobacteriaceae in the three groups were 0.70%, 12.80%, and 27.60%, respectively, and those of Pseudomonadaceae were 0.00%, 0.00% and 5.90%, respectively (P < 0.05 for all, Table 4). Similarly, phylum of Verrucomicrobia also increased in post-LT patients (P < 0.05, Figure 4). Besides these, the proportions of Actinobacteria and Acidobacteria were relatively balanced, and no significant differences existed among the three groups.
| Phylum | Class | Family | Healthy (n = 10) | Post-LT | |
| Uncomplicated (n = 10) | NAS (n = 10) | ||||
| Bacteroidetes | Bacteroidia | Bacteroidaceae | 11.60% ± 5.33% | 16.20% ± 3.20% | 2.70% ± 0.97%ac |
| Prevotellaceae | 11.60% ± 4.56% | 0.00% ± 0.00%a | 0.70% ± 0.08%a | ||
| Firmicutes | Bacilli | Enterococcaceae | 0.00% ± 0.00% | 1.20% ± 0.45%a | 2.60% ± 0.87%ac |
| Leuconostocaceae | 0.00% ± 0.00% | 0.70% ± 0.20% | 0.40% ± 0.05% | ||
| Streptococcaceae | 0.30% ± 0.11% | 3.90% ± 1.05%a | 8.60% ± 4.10%ac | ||
| Lachnospiraceae | 21.50% ± 6.78% | 9.80% ± 2.45%a | 10.50% ± 3.44%a | ||
| Ruminococcaceae | 30.90% ± 6.78% | 7.00% ± 3.16%a | 11.20% ± 2.33%a | ||
| Proteobacteria | γ-proteobacteria | Enterobacteriaceae | 0.70% ± 0.35% | 12.80% ± 2.56%a | 27.60% ± 7.06%ac |
| Pseudomonadaceae | 0.00% ± 0.00% | 0.00% ± 0.00% | 5.90% ± 3.16%ac | ||
| Verrucomicrobia | Verrucomicrobiae | Verrucomicrobiaceae | 0.10% ± 0.09% | 0.40% ± 0.16%a | 0.40% ± 0.05%a |
Nowadays, more and more studies have suggested the potential relationship between gut microbiota and liver diseases. Bacterial overgrowth or dysbacteriosis has also been proved to contribute to recipient’s post-LT complications[23]. In this study, we investigated the fecal microbial communities in patients diagnosed with NAS by pyrosequencing of the 16S rRNA V3-V4 region, taking the well-recovered recipients (uncomplicated) after OLT as negative controls and normal non-LT individuals as healthy controls, to explore the possible relationship between post-LT biliary complications and host’s gut microbiota.
According to our results, a structural change of fecal microbial communities was observed in patients who underwent LT, especially for those diagnosed with NAS. As α-diversity indexes reflected, post-LT patients presented with a significantly lower gut microbial diversity than healthy individuals, with the decrease of Firmicutes and Bacteroidetes and increase of Proteobacteria and Verrucomicrobia at the phylum level. Firmicutes and Bacteroidetes were intestinal dominant bacteria, playing a key role in maintaining host’s intestinal homeostasis. A decrease of these two bacteria always indicated the destruction of intestinal barrier function and increased risk of bacterial translocation[24]. In fact, the decrease of these two phyla was partially attributed to the increase of Proteobacteria and Verrucomicrobia, which usually contributed to a very small portion of human gut microbiota[25,26]. Similar changes had also been reported in cirrhotic patients waiting for OLT[27]. However, the shifts in our study were more obvious. At the family level, we found that the proportions of Prevotellaceae, Bacteroidaceae, Lachnospiraceae, and Ruminococcaceae were lower in post-LT patients, accompanied with an increase of Enterococcaceae, Streptococcaceae, Enterobacteriaceae, and Pseudomonadaceae. In previous studies, families of Lachnospiraceae and Ruminococcaceae were suggested to participate in the metabolism of short-chain fatty acids (SCFAs), while SCFAs have been regarded as a molecular link between the microbiota and inflammation by acting on their specific G protein-coupled receptors 43 (GPR 43). Exogenous supplement of SCFAs can inhibit oxidative stress and inflammatory response induced by high glucose and bacterial endotoxins (LPS)[28-30]. Therefore, loss of these potentially beneficial bacteria during the perioperative period may aggravate systemic inflammatory reaction and finally lead to liver injury[31]. Meanwhile, families of Enterococcaceae, Streptococcaceae, Enterobacteriaceae, and Pseudomonadaceae were commonly regarded as pathogenic bacteria, and their overgrowth has been found to participate in various kinds of human diseases, and even linearly correlated to patient’s Child-Pugh score[27,32-34]. Moreover, bacterial translocation and elevation of LPS have been estimated in rats with liver ischemia-reperfusion injury or post-LT acute rejection[35-37]. Ren et al[38] also found that liver ischemic preconditioning can improve intestinal barrier function and promote the restorations of intestinal microbiota following OLT.
Compared with patients without complications after liver transplantation, patients diagnosed with NAS in our study showed a more significant decrease of Bacteroidetes and increase of Proteobacteria at the phylum level, with higher proportions of Enterococcaceae, Streptococcaceae, Enterobacteriaceae, and Pseudomonadaceae. This dramatic shift in the ratio between phyla or the expansion of Proteobacteria is often referred to as dysbacteriosis. Outgrowth of Enterococcaceae, Streptococcaceae, Enterobacteriaceae, and Pseudomonadaceae will lead to a large release of LPS and peptidoglycan. When recognized by human immune system via Toll-like receptors (TLRs) or nucleotide-binding oligomerization domain like receptors (NLRs), LPS and peptidoglycan would trigger the pro-inflammatory NF-κB cascade and directly stimulate hepatic stellate cells, which finally contributed to liver damage and liver disease progression[14,39,40]. For patients who underwent hepatic inflow occlusion and immunosuppressive treatment during or after OLT, these overgrown pathogenic bacteria may easily penetrate through the intestinal barrier and translocate in the bloodstream, finally aggravating the ischemic reperfusion injury. While bile ducts are susceptible to inflammatory damage, so serious gut dysbacteriosis may exacerbate the cholangiocyte apoptosis and eventually lead to bile duct strictures[41,42]. Whereas, the proportions of Lachnospiraceae and Ruminococcaceae were similar between the NAS group and uncomplicated group, indicating that the overgrowth of the former four pathogenic bacteria contributed more effect to the pathologic process. Nevertheless, the detailed relationship between bacterial shifts and NAS is not clear.
NAS is a serious and progressive complication after OLT. Since graft associated factors are commonly uncontrollable, seeking new breakthrough from recipients themselves is quite important for its prevention. Interestingly, adjustment of microbial structure has been recommended in the treatment of inflammatory bowel disease and metabolic diseases[43]. Inhibition of pathogenic bacteria with antibiotics or probiotics has also been proved to improve cirrhosis patient’s prognosis, preventing the early-stage infection and acute rejection after OLT[44-46]. Therefore, targeted interventions to result in microbial compositional shift in NAS may contribute to its treatment in future.
As we know, this study is the first attempt to investigate the possible relationship between gut microbiota and post-LT biliary complication. With all possible influencing factors including preoperative characteristics and postoperative intervention equally distributed between all subjects, unbalanced ratio between pathogenic bacteria to dominant bacteria existed in patients with non-anastomotic biliary strictures after liver transplantation. This finding might indicate the shifts of fecal microbial communities participate in or exacerbate the process of bile duct injury. However, we admitted that this is a small-volume study from a single-center experience, and gut microbial changes related to NAS remain obscure. To verify the possible mechanisms, larger-scale, multicenter studies are necessary in the future.
In conclusion, our findings show that fecal microbial composition of patients with nonanastomotic biliary stricture is distinct from that of patients with no complications after orthotopic liver transplantation. These compositional shifts of the increase of potential pathogenic bacteria (e.g., Enterococcaceae, Streptococcaceae, Enterobacteriaceae, and Pseudomonadaceae) as well as the decrease of dominant bacteria (e.g., Bacteroidaceae) might contribute to the incidence of NAS. However, the underlying mechanism warrants further investigation.
Non-anastomotic biliary stricture (NAS) is a lethal disorder after liver transplantation (LT), but the mechanisms are still obscure. Gut microbiota has been shown to participate in the pathogenesis of some post-LT complications, while the characteristics of microbial communities in patients with NAS have never been investigated.
The purpose of this study was to explore the possible relationship between fecal microbial communities and NAS after OLT.
To perform possible mechanism research about NAS after LT to shed some light on its prevention in future.
A total of 30 subjects including 10 patients with NAS, 10 patients with no complications after LT, and 10 non-LT healthy individuals were enrolled. Fecal microbial communities were assessed by the 16S rRNA gene sequencing technology. Diversity indexes and the species accumulation curve were calculated by QIIME. PCA plots of the bacterial communities were created using pcaMethods. Other data analysis was finished by Chi-square or Fisher’s exact test or ANOVA test using SPSS software.
Different from the uncomplicated and healthy groups, unbalanced fecal bacterium ratio existed in patients with non-anastomotic biliary strictures after liver transplantation. The results showed that NAS patients were associated with a decrease of Firmicutes and Bacteroidetes and an increase of Proteobacteria at the phylum level, with the proportion-ratio imbalance between potentially pathogenic families including Enterococcaceae, Streptococcaceae, Enterobacteriaceae, and Pseudomonadaceae and dominant families including Bacteroidaceae.
The compositional shifts of the increase of potential pathogenic bacterium as well as the decrease of dominant bacterium might contribute to the incidence of NAS. Gut microbiota may participate in the pathological process of NAS. Factors including poor liver graft, ABO-incompatibility, cytomegalovirus (CMV) infection contribute to the development of NAS.
Dysbacteriosis may be another inducer contributing to the development of NAS. The shifts of fecal microbial communities may participate in or exacerbate the process of bile duct injury. Unbalanced ratio of pathogenic bacteria to dominant bacteria really existed in patients with NAS after liver transplantation. What are the implications of this? Bacterial intervention may be a new therapy for preventing the occurence of NAS.
According to our study, shifts of fecal microbial communities may participate in or exacerbate the process of bile duct inflammation. This might be helpful for NAS prevention. While the definite relationship was obscure, more mechanism research about how microbiota affects the pathological process should be carried out in the future. To learn more interaction relationship between microbiota and biliary inflammatory injury, technology based on functional genomics may be used for future research.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: China
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): 0
Grade C (Good): C, C
Grade D (Fair): D
Grade E (Poor): 0
P- Reviewer: Kang KJ, Pompili M, Tsoulfas G S- Editor: Gong ZM L- Editor: Wang TQ E- Editor: Ma YJ
| 1. | Federle MP. Milestones and future trends in solid organ transplantation. Radiol Clin North Am. 1995;33:417-434. [PubMed] |
| 2. | Zitta S, Schaffellner S, Gutschi J, Meinitzer A, Kniepeiss D, Artinger K, Reibnegger G, Rosenkranz AR, Wagner D. The Effect of Mammalian Target of Rapamycin Versus Calcineurin Inhibitor-based Immunosuppression on Measured Versus Estimated Glomerular Filtration Rate After Orthotopic Liver Transplantation. Transplantation. 2015;99:1250-1256. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 10] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 3. | Gastaca M. Biliary complications after orthotopic liver transplantation: a review of incidence and risk factors. Transplant Proc. 2012;44:1545-1549. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 81] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 4. | Seehofer D, Eurich D, Veltzke-Schlieker W, Neuhaus P. Biliary complications after liver transplantation: old problems and new challenges. Am J Transplant. 2013;13:253-265. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 199] [Cited by in RCA: 215] [Article Influence: 17.9] [Reference Citation Analysis (0)] |
| 5. | Buis CI, Verdonk RC, Van der Jagt EJ, van der Hilst CS, Slooff MJ, Haagsma EB, Porte RJ. Nonanastomotic biliary strictures after liver transplantation, part 1: Radiological features and risk factors for early vs. late presentation. Liver Transpl. 2007;13:708-718. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 156] [Cited by in RCA: 160] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 6. | Heidenhain C, Pratschke J, Puhl G, Neumann U, Pascher A, Veltzke-Schlieker W, Neuhaus P. Incidence of and risk factors for ischemic-type biliary lesions following orthotopic liver transplantation. Transpl Int. 2010;23:14-22. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 122] [Cited by in RCA: 132] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 7. | Serrano MT, Garcia-Gil A, Arenas J, Ber Y, Cortes L, Valiente C, Araiz JJ. Outcome of liver transplantation using donors older than 60 years of age. Clin Transplant. 2010;24:543-549. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 54] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 8. | Sundaram V, Jones DT, Shah NH, de Vera ME, Fontes P, Marsh JW, Humar A, Ahmad J. Posttransplant biliary complications in the pre- and post-model for end-stage liver disease era. Liver Transpl. 2011;17:428-435. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 73] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 9. | Ley RE, Turnbaugh PJ, Klein S, Gordon JI. Microbial ecology: human gut microbes associated with obesity. Nature. 2006;444:1022-1023. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7222] [Cited by in RCA: 6396] [Article Influence: 336.6] [Reference Citation Analysis (0)] |
| 10. | Tremaroli V, Bäckhed F. Functional interactions between the gut microbiota and host metabolism. Nature. 2012;489:242-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2779] [Cited by in RCA: 3071] [Article Influence: 236.2] [Reference Citation Analysis (0)] |
| 11. | Shanahan F, Quigley EM. Manipulation of the microbiota for treatment of IBS and IBD-challenges and controversies. Gastroenterology. 2014;146:1554-1563. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 127] [Cited by in RCA: 124] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 12. | Cani PD. Metabolism in 2013: The gut microbiota manages host metabolism. Nat Rev Endocrinol. 2014;10:74-76. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 123] [Article Influence: 11.2] [Reference Citation Analysis (0)] |
| 13. | Wang W, Xu S, Ren Z, Jiang J, Zheng S. Gut microbiota and allogeneic transplantation. J Transl Med. 2015;13:275. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 62] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 14. | Chassaing B, Etienne-Mesmin L, Gewirtz AT. Microbiota-liver axis in hepatic disease. Hepatology. 2014;59:328-339. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 219] [Cited by in RCA: 256] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 15. | Xie Y, Chen H, Zhu B, Qin N, Chen Y, Li Z, Deng M, Jiang H, Xu X, Yang J. Effect of intestinal microbiota alteration on hepatic damage in rats with acute rejection after liver transplantation. Microb Ecol. 2014;68:871-880. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 26] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 16. | Xie YR, Liu SL, Liu X, Luo ZB, Zhu B, Li ZF, Li LJ, He Y, Jiang L, Li H. Intestinal microbiota and innate immunity-related gene alteration in cirrhotic rats with liver transplantation. Transplant Proc. 2011;43:3973-3979. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 23] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 17. | Mazzaferro V, Regalia E, Doci R, Andreola S, Pulvirenti A, Bozzetti F, Montalto F, Ammatuna M, Morabito A, Gennari L. Liver transplantation for the treatment of small hepatocellular carcinomas in patients with cirrhosis. N Engl J Med. 1996;334:693-699. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5110] [Cited by in RCA: 5300] [Article Influence: 182.8] [Reference Citation Analysis (0)] |
| 18. | Muyzer G, de Waal EC, Uitterlinden AG. Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol. 1993;59:695-700. [PubMed] |
| 20. | Blaxter M, Mann J, Chapman T, Thomas F, Whitton C, Floyd R, Abebe E. Defining operational taxonomic units using DNA barcode data. Philos Trans R Soc Lond B Biol Sci. 2005;360:1935-1943. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 544] [Cited by in RCA: 485] [Article Influence: 25.5] [Reference Citation Analysis (0)] |
| 21. | Xu G, Zhong X, Wang Y, Xu H. An approach to detecting species diversity of microfaunas in colonization surveys for marine bioassessment based on rarefaction curves. Mar Pollut Bull. 2014;88:268-274. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 8] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 22. | Jolliffe IT, Cadima J. Principal component analysis: a review and recent developments. Philos Trans A Math Phys Eng Sci. 2016;374:20150202. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2837] [Cited by in RCA: 2177] [Article Influence: 241.9] [Reference Citation Analysis (0)] |
| 23. | Doycheva I, Leise MD, Watt KD. The Intestinal Microbiome and the Liver Transplant Recipient: What We Know and What We Need to Know. Transplantation. 2016;100:61-68. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 38] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 24. | Arumugam M, Raes J, Pelletier E, Le Paslier D, Yamada T, Mende DR, Fernandes GR, Tap J, Bruls T, Batto JM. Enterotypes of the human gut microbiome. Nature. 2011;473:174-180. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5822] [Cited by in RCA: 5010] [Article Influence: 357.9] [Reference Citation Analysis (2)] |
| 25. | Lupp C, Robertson ML, Wickham ME, Sekirov I, Champion OL, Gaynor EC, Finlay BB. Host-mediated inflammation disrupts the intestinal microbiota and promotes the overgrowth of Enterobacteriaceae. Cell Host Microbe. 2007;2:204. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 311] [Cited by in RCA: 370] [Article Influence: 20.6] [Reference Citation Analysis (0)] |
| 26. | Balzan S, de Almeida Quadros C, de Cleva R, Zilberstein B, Cecconello I. Bacterial translocation: overview of mechanisms and clinical impact. J Gastroenterol Hepatol. 2007;22:464-471. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 243] [Cited by in RCA: 272] [Article Influence: 15.1] [Reference Citation Analysis (0)] |
| 27. | Qin N, Yang F, Li A, Prifti E, Chen Y, Shao L, Guo J, Le Chatelier E, Yao J, Wu L. Alterations of the human gut microbiome in liver cirrhosis. Nature. 2014;513:59-64. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1230] [Cited by in RCA: 1533] [Article Influence: 139.4] [Reference Citation Analysis (40)] |
| 28. | Huang W, Guo HL, Deng X, Zhu TT, Xiong JF, Xu YH, Xu Y. Short-Chain Fatty Acids Inhibit Oxidative Stress and Inflammation in Mesangial Cells Induced by High Glucose and Lipopolysaccharide. Exp Clin Endocrinol Diabetes. 2017;125:98-105. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 82] [Cited by in RCA: 127] [Article Influence: 15.9] [Reference Citation Analysis (0)] |
| 29. | Lin MY, de Zoete MR, van Putten JP, Strijbis K. Redirection of Epithelial Immune Responses by Short-Chain Fatty Acids through Inhibition of Histone Deacetylases. Front Immunol. 2015;6:554. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 116] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 30. | Kim CH, Park J, Kim M. Gut microbiota-derived short-chain Fatty acids, T cells, and inflammation. Immune Netw. 2014;14:277-288. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 337] [Cited by in RCA: 476] [Article Influence: 43.3] [Reference Citation Analysis (0)] |
| 31. | Duncan SH, Louis P, Flint HJ. Cultivable bacterial diversity from the human colon. Lett Appl Microbiol. 2007;44:343-350. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 170] [Cited by in RCA: 185] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 32. | Chen Y, Yang F, Lu H, Wang B, Chen Y, Lei D, Wang Y, Zhu B, Li L. Characterization of fecal microbial communities in patients with liver cirrhosis. Hepatology. 2011;54:562-572. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 674] [Cited by in RCA: 794] [Article Influence: 56.7] [Reference Citation Analysis (3)] |
| 33. | Riordan SM, Williams R. The intestinal flora and bacterial infection in cirrhosis. J Hepatol. 2006;45:744-757. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 73] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 34. | Pande C, Kumar A, Sarin SK. Small-intestinal bacterial overgrowth in cirrhosis is related to the severity of liver disease. Aliment Pharmacol Ther. 2009;29:1273-1281. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 97] [Cited by in RCA: 106] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 35. | Xie Y, Luo Z, Li Z, Deng M, Liu H, Zhu B, Ruan B, Li L. Structural shifts of fecal microbial communities in rats with acute rejection after liver transplantation. Microb Ecol. 2012;64:546-554. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 36] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 36. | Xing HC, Li LJ, Xu KJ, Shen T, Chen YB, Sheng JF, Yu YS, Chen YG. Intestinal microflora in rats with ischemia/reperfusion liver injury. J Zhejiang Univ Sci B. 2005;6:14-21. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 28] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 37. | Wu ZW, Ling ZX, Lu HF, Zuo J, Sheng JF, Zheng SS, Li LJ. Changes of gut bacteria and immune parameters in liver transplant recipients. Hepatobiliary Pancreat Dis Int. 2012;11:40-50. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 97] [Cited by in RCA: 92] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 38. | Ren Z, Cui G, Lu H, Chen X, Jiang J, Liu H, He Y, Ding S, Hu Z, Wang W. Liver ischemic preconditioning (IPC) improves intestinal microbiota following liver transplantation in rats through 16s rDNA-based analysis of microbial structure shift. PLoS One. 2013;8:e75950. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 40] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 39. | Fung TC, Olson CA, Hsiao EY. Interactions between the microbiota, immune and nervous systems in health and disease. Nat Neurosci. 2017;20:145-155. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 980] [Cited by in RCA: 1299] [Article Influence: 162.4] [Reference Citation Analysis (0)] |
| 40. | Seki E, De Minicis S, Osterreicher CH, Kluwe J, Osawa Y, Brenner DA, Schwabe RF. TLR4 enhances TGF-beta signaling and hepatic fibrosis. Nat Med. 2007;13:1324-1332. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1361] [Cited by in RCA: 1554] [Article Influence: 86.3] [Reference Citation Analysis (1)] |
| 41. | Imamura H, Brault A, Huet PM. Effects of extended cold preservation and transplantation on the rat liver microcirculation. Hepatology. 1997;25:664-671. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 39] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 42. | Moench C, Moench K, Lohse AW, Thies J, Otto G. Prevention of ischemic-type biliary lesions by arterial back-table pressure perfusion. Liver Transpl. 2003;9:285-289. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 107] [Cited by in RCA: 104] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 43. | Yuan F, Ni H, Asche CV, Kim M, Walayat S, Ren J. Efficacy of Bifidobacterium infantis 35624 in patients with irritable bowel syndrome: a meta-analysis. Curr Med Res Opin. 2017;33:1191-1197. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 55] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 44. | Liu Q, Duan ZP, Ha DK, Bengmark S, Kurtovic J, Riordan SM. Synbiotic modulation of gut flora: effect on minimal hepatic encephalopathy in patients with cirrhosis. Hepatology. 2004;39:1441-1449. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 444] [Cited by in RCA: 454] [Article Influence: 21.6] [Reference Citation Analysis (0)] |
| 45. | Safdar N, Said A, Lucey MR. The role of selective digestive decontamination for reducing infection in patients undergoing liver transplantation: a systematic review and meta-analysis. Liver Transpl. 2004;10:817-827. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 60] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 46. | Rayes N, Seehofer D, Theruvath T, Schiller RA, Langrehr JM, Jonas S, Bengmark S, Neuhaus P. Supply of pre- and probiotics reduces bacterial infection rates after liver transplantation--a randomized, double-blind trial. Am J Transplant. 2005;5:125-130. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 335] [Cited by in RCA: 312] [Article Influence: 15.6] [Reference Citation Analysis (0)] |