Published online Jul 7, 2017. doi: 10.3748/wjg.v23.i25.4491
Peer-review started: February 27, 2017
First decision: April 7, 2017
Revised: April 14, 2017
Accepted: May 19, 2017
Article in press: May 19, 2017
Published online: July 7, 2017
Processing time: 131 Days and 14.8 Hours
Antibiotics are usually prescribed to cure infections but they also have significant modulatory effects on the gut microbiota. Several alterations of the intestinal bacterial community have been reported during antibiotic treatment, including the reduction of beneficial bacteria as well as of microbial alpha-diversity. Although after the discontinuation of antibiotic therapies it has been observed a trend towards the restoration of the original condition, the new steady state is different from the previous one, as if antibiotics induced some kind of irreversible perturbation of the gut microbial community. The poorly absorbed antibiotic rifaximin seem to be different from the other antibiotics, because it exerts non-traditional effects additional to the bactericidal/bacteriostatic activity on the gut microbiota. Rifaximin is able to reduce bacterial virulence and translocation, has anti-inflammatory properties and has been demonstrated to positively modulate the gut microbial composition. Animal models, culture studies and metagenomic analyses have demonstrated an increase in Bifidobacterium, Faecalibacterium prausnitzii and Lactobacillus abundance after rifaximin treatment, probably consequent to the induction of bacterial resistance, with no major change in the overall gut microbiota composition. Antibiotics are therefore modulators of the symbiotic relationship between the host and the gut microbiota. Specific antibiotics, such as rifaximin, can also induce eubiotic changes in the intestinal ecosystem; this additional property may represent a therapeutic advantage in specific clinical settings.
Core tip: The traditional use of antibiotics in the clinical practice is to antagonize local or systemic infections. However, antibiotics induce several alterations of the gut microbiota, including the reduction of beneficial bacteria as well as of microbial diversity. Rifaximin is a non-absorbable antibiotic with broad-spectrum activity and non-traditional antimicrobial effects. Rifaximin has the potential to induce a positive modulation of the gut microbiota, favoring the growth of bacteria beneficial to the host without altering its overall composition. Therefore, rifaximin can be defined not only as an antibiotic but also as an “eubiotic”, namely a positive modulator of the gut ecosystem.
- Citation: Ponziani FR, Zocco MA, D’Aversa F, Pompili M, Gasbarrini A. Eubiotic properties of rifaximin: Disruption of the traditional concepts in gut microbiota modulation. World J Gastroenterol 2017; 23(25): 4491-4499
- URL: https://www.wjgnet.com/1007-9327/full/v23/i25/4491.htm
- DOI: https://dx.doi.org/10.3748/wjg.v23.i25.4491
The human gut microbiota has been the main target of scientific research in the very recent years. Studies based on metagenomic techniques have revealed the multifaceted abilities of gut microbes, ranging from metabolic functions to immunomodulation, from anti-pathogen activity to behavior conditioning[1-5]. As the gut microbiota plays a crucial role in maintaining humans’ health, less or more specific gut microbiota alterations have been associated with various gastrointestinal diseases[6-20]. These findings strongly support the use of gut microbiota modulators such as antibiotics, prebiotics and probiotics as the treatment of choice in almost all gastrointestinal disorders.
In addition to the speculative and intriguing value of these physiopathological findings, it is also interesting to address which modifications of the gut microbiota may occur after a therapeutic intervention, such as antibiotic treatment. In particular, recent metagenomic studies have highlighted a positive modulation of gut bacteria associated with the administration of rifaximin, one of the antibiotics most frequently used for the treatment of digestive diseases. This has opened the scenario to the new concept of antibiotic-related gut microbiota modulation, which overcomes the traditional bactericidal and bacteriostatic effects and involves the possibility of more complex interactions, resulting in a modulation of the gut flora favorable to the host.
In this paper, we will focus on the effects of antibiotics, and in particular rifaximin, on gut bacteria, discussing both its traditional and non-traditional properties, and defining the new concept of “eubiotic modulation” of the gut microbiota.
The traditional use of antibiotics in the clinical practice is to antagonize local or systemic infections. However, after the discovery of the gut microbiota potential, several studies have focused on the effects of antibiotic therapies on commensal gut microbes[21-27]. Beta-lactamics, fluoro-quinolones, glycylcyclines, lincosamide, nitroimidazole, and various combinations of antibiotics are able to produce a profound alteration of the gut microbiota composition, mainly characterized by the reduction of autochthonous taxa and by the increase of potentially pathogenic bacteria, such as Enterobacteriaceae. In contrast, Bifidobacteria, Faecalibacterium prausnitzii and Lactobacilli, which usually exert beneficial effects to the gut, seem to be reduced after antibiotic treatment. Bifidobacteria are included into several probiotic preparations and have intestinal and systemic anti-inflammatory effects, being also protective against antibiotic associated diarrhea and infective colitis[28-53]. Faecalibacterium prausnitzii is a butyrate-producing bacterium with a high metabolic activity[54]; butyrate is involved in intestinal cells life cycle, has immunomodulatory effects and protects against infections caused by pathogens[55,56]. Finally, Lactobacilli are known for their anti-inflammatory, immunomodulatory, anti-oxidant, anti-bacterial and anti-viral properties[57].
Besides the effect on specific bacterial strains, antibiotic treatment decreases taxonomic richness, diversity, and evenness of the gut microbial community, producing a shift to an alternative state that is different from the baseline[27,58]. These alterations seem to recover after the end of antibiotic administration; however, the gut microbial composition does not exactly return to the pre-treatment condition, but rather it acquires a new connotation, similar (but not identical) to the original one. All these modifications are highly variable among individuals, and the consequences of antibiotics-associated gut microbiota perturbation in humans remain unknown.
Morgun et al[59] analyzed metagenomic and metatranscriptomic changes in a mouse model of microbiota depletion after the administration of an antibiotic cocktail (ampicillin, vancomycin, neomycin and metronidazole). The main registered effects, in addition to the drop in total bacterial mass and the emergence of resistan strains, were (1) the impairment of local mucosal immunity, with a depletion of immune cells in the lamina propria and in the villous epithelium, as well as of IgA-producing plasma cells; and (2) mitochondrial toxicity, leading to increased apoptosis and cell death in the intestinal epithelium of treated animals. The authors concluded that gut microbiota depletion as well as the development of resistance among the remaining bacteria and the consequent effects on host tissues were the major consequences produced by antibiotics on the gut.
Therefore, two main properties of antibiotics should be recognized: the classic effect against pathogens, which is the main indication for their use in clinical practice, and the modulation of the commensal gut microbiota, which is a “collateral” effect. Although the consequences of this last feature on the host are still not clear, the reduction of beneficial bacteria may cause the loss of the favorable influence exerted by the gut microbiota on human health.
This may reasonably induce to suppose a detrimental effect of antibiotics on intestinal ecology. However, this is not the case of all antibiotics. Recent findings have pointed out that rifaximin can positively modulate the gut microbiota. This peculiar characteristic may differentiate rifaximin from the other systemic antibiotics.
Rifaximin is an antibiotic with broad-spectrum activity against Gram positive and Gram negative aerobic and anaerobic bacteria, effective for the treatment of travelers’ diarrhea and other gastrointestinal infective conditions such as Clostridium difficile colitis[60-62]. As a consequence of the negligible systemic absorption (< 0.4%)[63], rifaximin reaches high tissue concentrations, as high as 8000 mcg/g in fecal samples after a 3-d regimen of 800 mg daily[64], far beyond the minimal inhibitory concentrations observed for local microbial isolates. Another characteristic of rifaximin is bile acids-dependent solubility; this makes the drug more effective in the small intestine while colonic bacteria are poorly inhibited[65]. Indeed, changes in the colonic gut microbiota composition are progressively but completely reversed after rifaximin interruption, differently from the effects on duodenal bacteria that appear more stable[66].
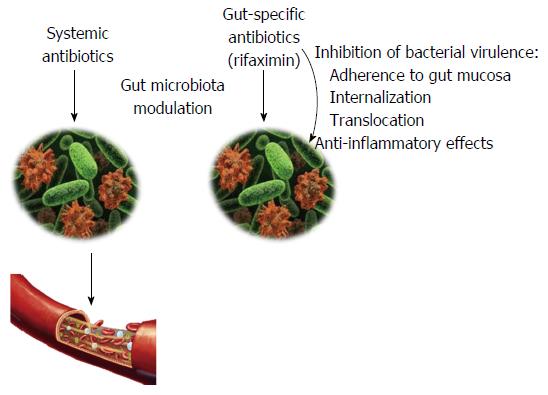
In addition to the bactericidal and bacteriostatic activity, which is typical of an antibiotic, rifaximin can also exert non-traditional effects on the gut microbiota (Figure 1). In particular, rifaximin can down-regulate the inflammatory response triggered by the gut microbes by inhibiting the activation of the nuclear factor (NF)-κB via the pregnane X receptor (PXR) and by reducing the expression of the pro-inflammatory cytokines interleukin (IL)-1B and tumor necrosis factor-alpha (TNFα)[67-71].
Moreover, rifaximin alters bacterial virulence through the inhibition of adhesion, internalization and translocation[72-75], and can modify bacterial metabolism[76].
Rifaximin modulation of gut microbial composition also belongs to the non-traditional features of this poorly absorbed antibiotic.
In addition to the aforementioned activity against bacterial adherence, internalization and translocation and to the anti-inflammatory effects, there is evidence of a positive modulation of the gut microbiota as a consequence of rifaximin treatment (Table 1).
| Ref. | Patients/model | Technique | Rifaximin dose | Changes in gut microbiota after rifaximin |
| Brigidi et al[77], 2002 | 12 pts UC | Standard bacteriological procedures | 1800 mg/d, 3 cycles of 10 d followed by 25 d of wash-out | Enterococci: < |
| Coliform: = | ||||
| Bifidobacteria: > | ||||
| Lactobacilli: < | ||||
| Clostridium perfrigens: > than < | ||||
| Bacteroides: | ||||
| unpredictable variations | ||||
| Candida: = | ||||
| Overall composition: | ||||
| not explored | ||||
| Maccaferri et al[78], 2010 | 4 pts colonic active CD | Continuous | 1800 mg/d | Bifidobacterium: > |
| culture colonic model system, FISH, quantitative PCR, PCR-denaturing gradient gel electrophoresis | Atopobium: > | |||
| Faecalibacterium prausnitzii: > | ||||
| Overall composition: = | ||||
| Bajaj et al[76], 2013 | 20 pts HE | 454 pyrosequencing | 1100 mg/d | Overall composition: = |
| Xu et al[79], 2014 | Rat model of visceral hyperalgesia | Quantitative PCR, 454 pyrosequencing | 150 mg/kg, twice daily | Lactobacillus: > |
| Clostridiaceae, Erysipelotrichaceae, and Peptostreptococcaceae: < | ||||
| Overall composition: 84% reduction in bacterial load | ||||
| Soldi et al[80], 2015 | 15 pts non-C IBS | Real-time PCR, Illumina pyrosequencing | 1650 mg/d for 14 d | Faecalibacterium prausnitzii: > |
| Clostridiaceae, Streptococcaceae: < | ||||
| Bacteroidaceae, Prevotellaceae: > | ||||
| Overall composition: = | ||||
| Ponziani et al[81], 2016 | 20 pts CD, UC, non-C IBS, DD, HE | 454 pyrosequencing | 1200 mg/d for 14 d | Lactobacillus: > |
| Roseburia, Haemophilus, Veillonella and | ||||
| Streptococcus: < | ||||
| Overall composition: = |
The pioneering study that demonstrated for the first time the increase in beneficial bacterial strains associated with rifaximin intake was conducted more than ten years ago on twelve patients affected by ulcerative colitis[77]. Rifaximin was administered at the dose of 1800 mg daily for 3 treatment cycles, each lasting for 10 d, followed by a wash-out period of 25 d. By means of standard bacteriological analysis, the Authors observed an increase in the concentration of Bifidobacteria after treatment, which tended to decrease in the interval periods. A subsequent study by the same group using a continuous culture model of the gut microbiota of four patients with active Crohn’s disease (CD)[78] also confirmed this preliminary result. Indeed, rifaximin administration at the same dose of 1800 mg daily did not alter the overall composition of the gut microbiota, but promoted the growth of Bifidobacterium, Atopobium and Faecalibacterium prausnitzii.
More recently, Xu et al[79] demonstrated that Lactobacilli could grow in response to rifaximin administration. In particular, in the mouse model of visceral hyperalgesia used in the study, the administration of rifaximin increased the abundance of Lactobacilli in the ileum; this effect was restricted only to this antibiotic, since in the same experimental conditions the administration of another poorly absorbed antibiotic, neomycin, produced just an increase in Proteobacteria.
Finally, two metagenomic analyses of the gut microbiota of human subjects affected by gastrointestinal and liver diseases have been recently published.
In the first one[80], patients with non-constipated irritable bowel syndrome were treated with rifaximin 550 mg tid for 14 d. Fecal specimens were collected before starting the treatment, at the end of it, and after a 6-wk wash-out period. As one of the most intriguing result of the study, the Authors observed an increase in Faecalibacterium prausnitzii abundance at the end of treatment, without any major modification of the overall gut microbiota composition.
In the second study[81], rifaximin was administered at the dose of 1200 mg daily for 10 d to patients with different gastrointestinal diseases, including irritable bowel syndrome, CD, ulcerative colitis and diverticular disease, and also to patients with liver cirrhosis complicated by hepatic encephalopathy. Stool samples were analyzed before, at the end and 1 mo after the end of rifaximin treatment. Lactobacillus abundance was increased at the end of treatment with rifaximin and 1 mo thereafter, while no modification of the overall composition of the gut microbiota was observed, even stratifying patients according to treatment timepoints and considering the original disease.
In conclusion, rifaximin is able to increase the abundance of beneficial intestinal bacteria, while keeping stable the overall composition of the gut microbial community. Although this peculiar behavior has been described so far only in association with rifaximin, it is related to one of the most common consequences of antibiotic therapies: the development of microbial resistance. Indeed, members of Bifidobacterium, Lactobacillus, Bacteroides, Clostridium, Eubacterium, Atopobium and Collinsella genera may develop mechanisms of resistance after rifaximin exposure, becoming able to grow at rifaximin concentrations higher than 1024 mg/L[82,83].
In the previously reported study, Brigidi et al[77] demonstrated that the increase in Bifidobacteria consequent to rifaximin intervention was associated with the increase of minimal inhibitory concentrations (MIC) values from 16 mcg/mL to 40 mcg/mL. Notably, rifaximin inhibits bacterial RNA synthesis acting on the β subunit of the RNA polymerase, and missense mutations of the core region of the rpoB gene encoding the β subunit of RNA polymerase have been described in Bifidobacterium infantis following rifaximin exposure[84,85]. Therefore, in the case of rifaximin, the acquisition of resistance may represent an advantage for the host rather than a detrimental consequence. However, the development of rifaximin resistance requires chromosomal mutation, which is vertically transmitted and has, therefore, a rare diffusion among intestinal microorganisms[61]; as a consequence, almost all resistant bacteria disappear within weeks after treatment discontinuation[86]. Data from metagenomic studies reflect this characteristic. Indeed, immediately after the beginning of rifaximin treatment, bacterial alpha diversity shows a trend towards reduction, reverting to values similar to pre-treatment after a wash-out period[80,81].
Based on these considerations, and also on the preliminary evidence that rifaximin effects do not abruptly disappear after treatment interruption, lasting in the short time period, further studies are needed to investigate if the positive modulation of the gut microbiota produced by rifaximin could persist over time.
In addition to the favorable modulation of the gut microbiota, rifaximin presents other features that make its use safely applicable in clinical practice.
Rifaximin is poorly absorbed and a negligible concentration is retrieved in urine after oral administration. This feature is typical of the branded formulation of the drug, which contains only the crystal polymorph -α, while a higher systemic bioavailability has been demonstrated for the generic formulation containing an amorphous form of the molecule[87]. Considerations about careful use of rifaximin in conditions increasing intestinal permeability, such as liver impairment, should also been provided. Rifaximin systemic exposure is increased by 10, 13 and 20 folds in Child A, B and C cirrhotic patients, respectively; although no dose adjustment is required by the manufacturer, a careful use is advisable in this clinical setting[88].
Rifaximin drug interactions are also expected to be rare, as confirmed by two studies including healthy individuals on treatment with CYP450 substrates[89,90]. P-glycoprotein is also involved in the metabolism of rifaximin but the clinical consequences of the co-administration of p-glycoprotein inhibitors are unknown.
Rifaximin has also been safely used in elderly subjects[91] and children[92]. However, teratogenic effects on the fetus have been demonstrated in animal models, and the benefits of rifaximin in pregnant women should be weighed against this risk[88].
A final consideration should be drawn about rifaximin tolerability in clinical practice. In the major studies reporting on rifaximin use in gastrointestinal diseases and hepatic encephalopathy, a low rate of adverse events has been registered, mainly of mild/low grade and with a similar frequency compared to the placebo[93]. The most frequent ones have been headache, nausea, dizziness, dyspepsia, abdominal discomfort, abdominal distention, diarrhea, constipation and flatulence. The incidence of infections was low too, and the few cases of C. difficile colitis reported during rifaximin treatment were generally associated with predisposing conditions, such as hospitalization and prolonged antibiotic use.
Therefore, rifaximin eubiotic properties are associated with a good safety and tolerability profile, which makes possible a wide use of this molecule in different clinical settings.
The increase in bacteria beneficial to the gut associated with rifaximin treatment, together with its additional properties and its local action, reasonably support a reclassification of this molecule, which can be defined not only as an antibiotic but also as an “eubiotic”, namely a positive modulator of the gut ecosystem.
Usually, humans live in a commensal and mutualistic symbiotic relationship with their own gut microbiota, which is a prolonged and close association resulting in a benefit for one or both the organisms[94].
However, the term “symbiosis” does not completely describe the exact nature of this cooperation, failing to depict the dynamism that is the culprit of this relationship. Indeed, while the human host can be considered a relatively stable system, it is extremely difficult to take a static picture of the gut microbiota. However, trying to partially summarize the high variability of the microbial ecosystem, two main conditions may be outlined: “dysbiosis” and “eubiosis”. Dysbiosis has been defined as a condition characterized by “imbalanced intestinal microbial community with quantitative and qualitative changes in the composition of the microbiota itself, in its metabolic activities or in the local distribution of its members”[94]; in contrast, a proper definition of eubiosis has not been provided yet. Intuitively, a quantitative and qualitative harmonic balance of the gut microbial components, resulting in a healthy metabolic and immunologic cooperation with the host, should be the main feature of eubiosis. However, apart from infections caused by opportunistic and non-opportunistic pathogens, it is difficult to figure-out which could be the main change conditioning the transition from eubiosis to dysbiosis in other pathologic settings (e.g., irritable bowel syndrome, inflammatory bowel diseases etc.). Probably dysbiosis may occur transiently in healthy individuals, usually resolving spontaneously without any clinical manifestation or presenting with mild symptoms. Conversely, changes in the composition of the gut microbiota observed in patients with gastrointestinal diseases may be configured as a the persistence of dysbiotic alterations that fails to recover, thus contributing to the onset and to the progression of the pathologic condition in predisposed individuals, as well as to the worsening of clinical symptoms. This explains the efficacy of eubiotic compounds such as rifaximin in these clinical settings: the beneficial modulation of the gut microbiota produced by rifaximin, together with its other non-traditional effects, result in a beneficial anti-inflammatory and trophic action on the intestine.
The symbiotic host-gut microbiota relationship is a condition of relational harmony fluctuating between eubiosis and dysbiosis. Since dysbiosis plays a role in the pathogenesis and in the progression of several gastrointestinal and systemic diseases, the use of eubiotics, such as rifaximin, should be reconsidered in these settings in the light of the positive modulating effects on the gut microbial community, and their use should be further supported and integrated with the other available medical treatments.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: Italy
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Giamarellos-Bourboulis EJ, Tosetti C S- Editor: Qi Y L- Editor: A E- Editor: Zhang FF
| 1. | Nieuwdorp M, Gilijamse PW, Pai N, Kaplan LM. Role of the microbiome in energy regulation and metabolism. Gastroenterology. 2014;146:1525-1533. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 266] [Cited by in RCA: 298] [Article Influence: 27.1] [Reference Citation Analysis (0)] |
| 2. | Hollister EB, Gao C, Versalovic J. Compositional and functional features of the gastrointestinal microbiome and their effects on human health. Gastroenterology. 2014;146:1449-1458. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 300] [Cited by in RCA: 326] [Article Influence: 29.6] [Reference Citation Analysis (0)] |
| 3. | Sommer F, Bäckhed F. The gut microbiota--masters of host development and physiology. Nat Rev Microbiol. 2013;11:227-238. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2011] [Cited by in RCA: 2407] [Article Influence: 200.6] [Reference Citation Analysis (0)] |
| 4. | Sekirov I, Russell SL, Antunes LC, Finlay BB. Gut microbiota in health and disease. Physiol Rev. 2010;90:859-904. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2501] [Cited by in RCA: 2749] [Article Influence: 183.3] [Reference Citation Analysis (1)] |
| 5. | Tremaroli V, Bäckhed F. Functional interactions between the gut microbiota and host metabolism. Nature. 2012;489:242-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2779] [Cited by in RCA: 3085] [Article Influence: 237.3] [Reference Citation Analysis (0)] |
| 6. | Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR. Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci USA. 2007;104:13780-13785. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3731] [Cited by in RCA: 3432] [Article Influence: 190.7] [Reference Citation Analysis (1)] |
| 7. | Sokol H, Seksik P. The intestinal microbiota in inflammatory bowel diseases: time to connect with the host. Curr Opin Gastroenterol. 2010;26:327-331. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 107] [Cited by in RCA: 116] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 8. | Elson CO, Cong Y. Host-microbiota interactions in inflammatory bowel disease. Gut Microbes. 2012;3:332-344. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 95] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 9. | Machiels K, Joossens M, Sabino J, De Preter V, Arijs I, Eeckhaut V, Ballet V, Claes K, Van Immerseel F, Verbeke K. A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut. 2014;63:1275-1283. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1036] [Cited by in RCA: 1337] [Article Influence: 121.5] [Reference Citation Analysis (3)] |
| 10. | Hansen R, Russell RK, Reiff C, Louis P, McIntosh F, Berry SH, Mukhopadhya I, Bisset WM, Barclay AR, Bishop J. Microbiota of de-novo pediatric IBD: increased Faecalibacterium prausnitzii and reduced bacterial diversity in Crohn’s but not in ulcerative colitis. Am J Gastroenterol. 2012;107:1913-1922. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 206] [Cited by in RCA: 219] [Article Influence: 16.8] [Reference Citation Analysis (0)] |
| 11. | Lepage P, Häsler R, Spehlmann ME, Rehman A, Zvirbliene A, Begun A, Ott S, Kupcinskas L, Doré J, Raedler A. Twin study indicates loss of interaction between microbiota and mucosa of patients with ulcerative colitis. Gastroenterology. 2011;141:227-236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 419] [Cited by in RCA: 452] [Article Influence: 32.3] [Reference Citation Analysis (1)] |
| 12. | Rajilić-Stojanović M, Biagi E, Heilig HG, Kajander K, Kekkonen RA, Tims S, de Vos WM. Global and deep molecular analysis of microbiota signatures in fecal samples from patients with irritable bowel syndrome. Gastroenterology. 2011;141:1792-1801. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 685] [Cited by in RCA: 773] [Article Influence: 55.2] [Reference Citation Analysis (0)] |
| 13. | Jalanka-Tuovinen J, Salojärvi J, Salonen A, Immonen O, Garsed K, Kelly FM, Zaitoun A, Palva A, Spiller RC, de Vos WM. Faecal microbiota composition and host-microbe cross-talk following gastroenteritis and in postinfectious irritable bowel syndrome. Gut. 2014;63:1737-1745. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 226] [Cited by in RCA: 248] [Article Influence: 22.5] [Reference Citation Analysis (0)] |
| 14. | Lin HC. Small intestinal bacterial overgrowth: a framework for understanding irritable bowel syndrome. JAMA. 2004;292:852-858. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 283] [Cited by in RCA: 265] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 15. | Riordan SM, Kim R. Bacterial overgrowth as a cause of irritable bowel syndrome. Curr Opin Gastroenterol. 2006;22:669-673. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 24] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 16. | Bouhnik Y, Alain S, Attar A, Flourié B, Raskine L, Sanson-Le Pors MJ, Rambaud JC. Bacterial populations contaminating the upper gut in patients with small intestinal bacterial overgrowth syndrome. Am J Gastroenterol. 1999;94:1327-1331. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 126] [Cited by in RCA: 121] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 17. | Sánchez E, Donat E, Ribes-Koninckx C, Fernández-Murga ML, Sanz Y. Duodenal-mucosal bacteria associated with celiac disease in children. Appl Environ Microbiol. 2013;79:5472-5479. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 120] [Cited by in RCA: 121] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 18. | Olivares M, Neef A, Castillejo G, Palma GD, Varea V, Capilla A, Palau F, Nova E, Marcos A, Polanco I. The HLA-DQ2 genotype selects for early intestinal microbiota composition in infants at high risk of developing coeliac disease. Gut. 2015;64:406-417. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 203] [Cited by in RCA: 222] [Article Influence: 22.2] [Reference Citation Analysis (1)] |
| 19. | Gevers D, Kugathasan S, Denson LA, Vázquez-Baeza Y, Van Treuren W, Ren B, Schwager E, Knights D, Song SJ, Yassour M. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe. 2014;15:382-392. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1945] [Cited by in RCA: 2358] [Article Influence: 214.4] [Reference Citation Analysis (0)] |
| 20. | Kostic AD, Xavier RJ, Gevers D. The microbiome in inflammatory bowel disease: current status and the future ahead. Gastroenterology. 2014;146:1489-1499. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1145] [Cited by in RCA: 1296] [Article Influence: 117.8] [Reference Citation Analysis (0)] |
| 21. | Penders J, Thijs C, Vink C, Stelma FF, Snijders B, Kummeling I, van den Brandt PA, Stobberingh EE. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics. 2006;118:511-521. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1513] [Cited by in RCA: 1528] [Article Influence: 80.4] [Reference Citation Analysis (0)] |
| 22. | Donskey CJ, Hujer AM, Das SM, Pultz NJ, Bonomo RA, Rice LB. Use of denaturing gradient gel electrophoresis for analysis of the stool microbiota of hospitalized patients. J Microbiol Methods. 2003;54:249-256. [PubMed] |
| 23. | De La Cochetière MF, Durand T, Lepage P, Bourreille A, Galmiche JP, Doré J. Resilience of the dominant human fecal microbiota upon short-course antibiotic challenge. J Clin Microbiol. 2005;43:5588-5592. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 219] [Cited by in RCA: 228] [Article Influence: 11.4] [Reference Citation Analysis (0)] |
| 24. | Young VB, Schmidt TM. Antibiotic-associated diarrhea accompanied by large-scale alterations in the composition of the fecal microbiota. J Clin Microbiol. 2004;42:1203-1206. [PubMed] |
| 25. | Bassis CM, Theriot CM, Young VB. Alteration of the murine gastrointestinal microbiota by tigecycline leads to increased susceptibility to Clostridium difficile infection. Antimicrob Agents Chemother. 2014;58:2767-2774. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 60] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 26. | Jernberg C, Löfmark S, Edlund C, Jansson JK. Long-term impacts of antibiotic exposure on the human intestinal microbiota. Microbiology. 2010;156:3216-3223. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 696] [Cited by in RCA: 718] [Article Influence: 47.9] [Reference Citation Analysis (0)] |
| 27. | Dethlefsen L, Relman DA. Incomplete recovery and individualized responses of the human distal gut microbiota to repeated antibiotic perturbation. Proc Natl Acad Sci USA. 2011;108 Suppl 1:4554-4561. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1850] [Cited by in RCA: 1652] [Article Influence: 118.0] [Reference Citation Analysis (0)] |
| 28. | Corrêa NB, Péret Filho LA, Penna FJ, Lima FM, Nicoli JR. A randomized formula controlled trial of Bifidobacterium lactis and Streptococcus thermophilus for prevention of antibiotic-associated diarrhea in infants. J Clin Gastroenterol. 2005;39:385-389. [PubMed] |
| 29. | Weizman Z, Asli G, Alsheikh A. Effect of a probiotic infant formula on infections in child care centers: comparison of two probiotic agents. Pediatrics. 2005;115:5-9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 365] [Cited by in RCA: 321] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 30. | Kajander K, Myllyluoma E, Rajilić-Stojanović M, Kyrönpalo S, Rasmussen M, Järvenpää S, Zoetendal EG, de Vos WM, Vapaatalo H, Korpela R. Clinical trial: multispecies probiotic supplementation alleviates the symptoms of irritable bowel syndrome and stabilizes intestinal microbiota. Aliment Pharmacol Ther. 2008;27:48-57. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 234] [Cited by in RCA: 222] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 31. | Guyonnet D, Chassany O, Ducrotte P, Picard C, Mouret M, Mercier CH, Matuchansky C. Effect of a fermented milk containing Bifidobacterium animalis DN-173 010 on the health-related quality of life and symptoms in irritable bowel syndrome in adults in primary care: a multicentre, randomized, double-blind, controlled trial. Aliment Pharmacol Ther. 2007;26:475-486. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 247] [Cited by in RCA: 232] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 32. | Agrawal A, Houghton LA, Morris J, Reilly B, Guyonnet D, Goupil Feuillerat N, Schlumberger A, Jakob S, Whorwell PJ. Clinical trial: the effects of a fermented milk product containing Bifidobacterium lactis DN-173 010 on abdominal distension and gastrointestinal transit in irritable bowel syndrome with constipation. Aliment Pharmacol Ther. 2009;29:104-114. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 209] [Cited by in RCA: 207] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 33. | Guyonnet D, Schlumberger A, Mhamdi L, Jakob S, Chassany O. Fermented milk containing Bifidobacterium lactis DN-173 010 improves gastrointestinal well-being and digestive symptoms in women reporting minor digestive symptoms: a randomised, double-blind, parallel, controlled study. Br J Nutr. 2009;102:1654-1662. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 65] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 34. | Marteau P, Guyonnet D, Lafaye de Micheaux P, Gelu S. A randomized, double-blind, controlled study and pooled analysis of two identical trials of fermented milk containing probiotic Bifidobacterium lactis CNCM I-2494 in healthy women reporting minor digestive symptoms. Neurogastroenterol Motil. 2013;25:331-e252. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 35] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 35. | Tillisch K, Labus J, Kilpatrick L, Jiang Z, Stains J, Ebrat B, Guyonnet D, Legrain-Raspaud S, Trotin B, Naliboff B. Consumption of fermented milk product with probiotic modulates brain activity. Gastroenterology. 2013;144:1394-1401, 1401.e1-1401.e4. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 733] [Cited by in RCA: 758] [Article Influence: 63.2] [Reference Citation Analysis (0)] |
| 36. | Waller PA, Gopal PK, Leyer GJ, Ouwehand AC, Reifer C, Stewart ME, Miller LE. Dose-response effect of Bifidobacterium lactis HN019 on whole gut transit time and functional gastrointestinal symptoms in adults. Scand J Gastroenterol. 2011;46:1057-1064. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 121] [Cited by in RCA: 135] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 37. | Lin HC, Hsu CH, Chen HL, Chung MY, Hsu JF, Lien RI, Tsao LY, Chen CH, Su BH. Oral probiotics prevent necrotizing enterocolitis in very low birth weight preterm infants: a multicenter, randomized, controlled trial. Pediatrics. 2008;122:693-700. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 359] [Cited by in RCA: 334] [Article Influence: 19.6] [Reference Citation Analysis (0)] |
| 38. | Kajander K, Hatakka K, Poussa T, Färkkilä M, Korpela R. A probiotic mixture alleviates symptoms in irritable bowel syndrome patients: a controlled 6-month intervention. Aliment Pharmacol Ther. 2005;22:387-394. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 252] [Cited by in RCA: 222] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 39. | Ishikawa H, Matsumoto S, Ohashi Y, Imaoka A, Setoyama H, Umesaki Y, Tanaka R, Otani T. Beneficial effects of probiotic bifidobacterium and galacto-oligosaccharide in patients with ulcerative colitis: a randomized controlled study. Digestion. 2011;84:128-133. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 140] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 40. | Almeida CC, Lorena SL, Pavan CR, Akasaka HM, Mesquita MA. Beneficial effects of long-term consumption of a probiotic combination of Lactobacillus casei Shirota and Bifidobacterium breve Yakult may persist after suspension of therapy in lactose-intolerant patients. Nutr Clin Pract. 2012;27:247-251. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 69] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 41. | O'Mahony L, McCarthy J, Kelly P, Hurley G, Luo F, Chen K, O’Sullivan GC, Kiely B, Collins JK, Shanahan F. Lactobacillus and bifidobacterium in irritable bowel syndrome: symptom responses and relationship to cytokine profiles. Gastroenterology. 2005;128:541-551. [PubMed] |
| 42. | Whorwell PJ, Altringer L, Morel J, Bond Y, Charbonneau D, O’Mahony L, Kiely B, Shanahan F, Quigley EM. Efficacy of an encapsulated probiotic Bifidobacterium infantis 35624 in women with irritable bowel syndrome. Am J Gastroenterol. 2006;101:1581-1590. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 562] [Cited by in RCA: 517] [Article Influence: 27.2] [Reference Citation Analysis (0)] |
| 43. | Groeger D, O’Mahony L, Murphy EF, Bourke JF, Dinan TG, Kiely B, Shanahan F, Quigley EM. Bifidobacterium infantis 35624 modulates host inflammatory processes beyond the gut. Gut Microbes. 2013;4:325-339. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 261] [Cited by in RCA: 320] [Article Influence: 26.7] [Reference Citation Analysis (0)] |
| 44. | Lin HC, Su BH, Chen AC, Lin TW, Tsai CH, Yeh TF, Oh W. Oral probiotics reduce the incidence and severity of necrotizing enterocolitis in very low birth weight infants. Pediatrics. 2005;115:1-4. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 428] [Cited by in RCA: 409] [Article Influence: 20.5] [Reference Citation Analysis (0)] |
| 45. | Bin-Nun A, Bromiker R, Wilschanski M, Kaplan M, Rudensky B, Caplan M, Hammerman C. Oral probiotics prevent necrotizing enterocolitis in very low birth weight neonates. J Pediatr. 2005;147:192-196. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 391] [Cited by in RCA: 364] [Article Influence: 18.2] [Reference Citation Analysis (0)] |
| 46. | Miele E, Pascarella F, Giannetti E, Quaglietta L, Baldassano RN, Staiano A. Effect of a probiotic preparation (VSL#3) on induction and maintenance of remission in children with ulcerative colitis. Am J Gastroenterol. 2009;104:437-443. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 350] [Cited by in RCA: 357] [Article Influence: 22.3] [Reference Citation Analysis (0)] |
| 47. | Guandalini S, Magazzù G, Chiaro A, La Balestra V, Di Nardo G, Gopalan S, Sibal A, Romano C, Canani RB, Lionetti P. VSL#3 improves symptoms in children with irritable bowel syndrome: a multicenter, randomized, placebo-controlled, double-blind, crossover study. J Pediatr Gastroenterol Nutr. 2010;51:24-30. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 210] [Cited by in RCA: 174] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 48. | Sood A, Midha V, Makharia GK, Ahuja V, Singal D, Goswami P, Tandon RK. The probiotic preparation, VSL#3 induces remission in patients with mild-to-moderately active ulcerative colitis. Clin Gastroenterol Hepatol. 2009;7:1202-1209, 1209.e1. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 387] [Cited by in RCA: 355] [Article Influence: 22.2] [Reference Citation Analysis (0)] |
| 49. | Tursi A, Brandimarte G, Papa A, Giglio A, Elisei W, Giorgetti GM, Forti G, Morini S, Hassan C, Pistoia MA. Treatment of relapsing mild-to-moderate ulcerative colitis with the probiotic VSL#3 as adjunctive to a standard pharmaceutical treatment: a double-blind, randomized, placebo-controlled study. Am J Gastroenterol. 2010;105:2218-2227. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 391] [Cited by in RCA: 354] [Article Influence: 23.6] [Reference Citation Analysis (0)] |
| 50. | Gionchetti P, Rizzello F, Helwig U, Venturi A, Lammers KM, Brigidi P, Vitali B, Poggioli G, Miglioli M, Campieri M. Prophylaxis of pouchitis onset with probiotic therapy: a double-blind, placebo-controlled trial. Gastroenterology. 2003;124:1202-1209. [PubMed] |
| 51. | Mimura T, Rizzello F, Helwig U, Poggioli G, Schreiber S, Talbot IC, Nicholls RJ, Gionchetti P, Campieri M, Kamm MA. Once daily high dose probiotic therapy (VSL#3) for maintaining remission in recurrent or refractory pouchitis. Gut. 2004;53:108-114. [PubMed] |
| 52. | Pronio A, Montesani C, Butteroni C, Vecchione S, Mumolo G, Vestri A, Vitolo D, Boirivant M. Probiotic administration in patients with ileal pouch-anal anastomosis for ulcerative colitis is associated with expansion of mucosal regulatory cells. Inflamm Bowel Dis. 2008;14:662-668. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 147] [Cited by in RCA: 153] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 53. | Selinger CP, Bell A, Cairns A, Lockett M, Sebastian S, Haslam N. Probiotic VSL#3 prevents antibiotic-associated diarrhoea in a double-blind, randomized, placebo-controlled clinical trial. J Hosp Infect. 2013;84:159-165. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 70] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 54. | Flint HJ, Scott KP, Duncan SH, Louis P, Forano E. Microbial degradation of complex carbohydrates in the gut. Gut Microbes. 2012;3:289-306. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1231] [Cited by in RCA: 1468] [Article Influence: 112.9] [Reference Citation Analysis (1)] |
| 55. | Macfarlane GT, Macfarlane S. Fermentation in the human large intestine: its physiologic consequences and the potential contribution of prebiotics. J Clin Gastroenterol. 2011;45 Suppl:S120-S127. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 265] [Cited by in RCA: 276] [Article Influence: 19.7] [Reference Citation Analysis (0)] |
| 56. | Plöger S, Stumpff F, Penner GB, Schulzke JD, Gäbel G, Martens H, Shen Z, Günzel D, Aschenbach JR. Microbial butyrate and its role for barrier function in the gastrointestinal tract. Ann N Y Acad Sci. 2012;1258:52-59. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 259] [Cited by in RCA: 327] [Article Influence: 25.2] [Reference Citation Analysis (0)] |
| 57. | Di Cerbo A, Palmieri B, Aponte M, Morales-Medina JC, Iannitti T. Mechanisms and therapeutic effectiveness of lactobacilli. J Clin Pathol. 2016;69:187-203. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 133] [Cited by in RCA: 182] [Article Influence: 18.2] [Reference Citation Analysis (0)] |
| 58. | Dethlefsen L, Huse S, Sogin ML, Relman DA. The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biol. 2008;6:e280. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1907] [Cited by in RCA: 1739] [Article Influence: 102.3] [Reference Citation Analysis (1)] |
| 59. | Morgun A, Dzutsev A, Dong X, Greer RL, Sexton DJ, Ravel J, Schuster M, Hsiao W, Matzinger P, Shulzhenko N. Uncovering effects of antibiotics on the host and microbiota using transkingdom gene networks. Gut. 2015;64:1732-1743. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 192] [Cited by in RCA: 212] [Article Influence: 21.2] [Reference Citation Analysis (0)] |
| 60. | Scarpignato C, Pelosini I. Experimental and clinical pharmacology of rifaximin, a gastrointestinal selective antibiotic. Digestion. 2006;73 Suppl 1:13-27. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 81] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 61. | Scarpignato C, Pelosini I. Rifaximin, a poorly absorbed antibiotic: pharmacology and clinical potential. Chemotherapy. 2005;51 Suppl 1:36-66. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 135] [Cited by in RCA: 174] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 62. | Marchese A, Salerno A, Pesce A, Debbia EA, Schito GC. In vitro activity of rifaximin, metronidazole and vancomycin against Clostridium difficile and the rate of selection of spontaneously resistant mutants against representative anaerobic and aerobic bacteria, including ammonia-producing species. Chemotherapy. 2000;46:253-266. [PubMed] |
| 63. | Chang YY, Lan MY, Wu HS, Huang SH, Chen SS, Liu JS. Decreased muscular radionuclide uptake in Tc-99m MIBI scintigraphy during paralytic phase of thyrotoxic periodic paralysis. Clin Nucl Med. 2008;33:297-298. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 64. | Jiang ZD, Ke S, Palazzini E, Riopel L, Dupont H. In vitro activity and fecal concentration of rifaximin after oral administration. Antimicrob Agents Chemother. 2000;44:2205-2206. [PubMed] |
| 65. | Darkoh C, Lichtenberger LM, Ajami N, Dial EJ, Jiang ZD, DuPont HL. Bile acids improve the antimicrobial effect of rifaximin. Antimicrob Agents Chemother. 2010;54:3618-3624. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 76] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 66. | Kim MS, Morales W, Hani AA, Kim S, Kim G, Weitsman S, Chang C, Pimentel M. The effect of rifaximin on gut flora and Staphylococcus resistance. Dig Dis Sci. 2013;58:1676-1682. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 29] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 67. | Mencarelli A, Migliorati M, Barbanti M, Cipriani S, Palladino G, Distrutti E, Renga B, Fiorucci S. Pregnane-X-receptor mediates the anti-inflammatory activities of rifaximin on detoxification pathways in intestinal epithelial cells. Biochem Pharmacol. 2010;80:1700-1707. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 86] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 68. | Hirota SA. Understanding the Molecular Mechanisms of Rifaximin in the Treatment of Gastrointestinal Disorders--A Focus on the Modulation of Host Tissue Function. Mini Rev Med Chem. 2015;16:206-217. [PubMed] |
| 69. | Cheng J, Shah YM, Ma X, Pang X, Tanaka T, Kodama T, Krausz KW, Gonzalez FJ. Therapeutic role of rifaximin in inflammatory bowel disease: clinical implication of human pregnane X receptor activation. J Pharmacol Exp Ther. 2010;335:32-41. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 145] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 70. | Ma X, Shah YM, Guo GL, Wang T, Krausz KW, Idle JR, Gonzalez FJ. Rifaximin is a gut-specific human pregnane X receptor activator. J Pharmacol Exp Ther. 2007;322:391-398. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 105] [Cited by in RCA: 103] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 71. | Mencarelli A, Renga B, Palladino G, Claudio D, Ricci P, Distrutti E, Barbanti M, Baldelli F, Fiorucci S. Inhibition of NF-κB by a PXR-dependent pathway mediates counter-regulatory activities of rifaximin on innate immunity in intestinal epithelial cells. Eur J Pharmacol. 2011;668:317-324. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 99] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 72. | Jiang ZD, Ke S, Dupont HL. Rifaximin-induced alteration of virulence of diarrhoea-producing Escherichia coli and Shigella sonnei. Int J Antimicrob Agents. 2010;35:278-281. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 67] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 73. | Brown EL, Xue Q, Jiang ZD, Xu Y, Dupont HL. Pretreatment of epithelial cells with rifaximin alters bacterial attachment and internalization profiles. Antimicrob Agents Chemother. 2010;54:388-396. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 79] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 74. | Schrodt C, McHugh EE, Gawinowicz MA, Dupont HL, Brown EL. Rifaximin-mediated changes to the epithelial cell proteome: 2-D gel analysis. PLoS One. 2013;8:e68550. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 18] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 75. | Fiorucci S, Distrutti E, Mencarelli A, Barbanti M, Palazzini E, Morelli A. Inhibition of intestinal bacterial translocation with rifaximin modulates lamina propria monocytic cells reactivity and protects against inflammation in a rodent model of colitis. Digestion. 2002;66:246-256. [PubMed] |
| 76. | Bajaj JS, Heuman DM, Sanyal AJ, Hylemon PB, Sterling RK, Stravitz RT, Fuchs M, Ridlon JM, Daita K, Monteith P. Modulation of the metabiome by rifaximin in patients with cirrhosis and minimal hepatic encephalopathy. PLoS One. 2013;8:e60042. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 254] [Cited by in RCA: 339] [Article Influence: 28.3] [Reference Citation Analysis (0)] |
| 77. | Brigidi P, Swennen E, Rizzello F, Bozzolasco M, Matteuzzi D. Effects of rifaximin administration on the intestinal microbiota in patients with ulcerative colitis. J Chemother. 2002;14:290-295. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 64] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 78. | Maccaferri S, Vitali B, Klinder A, Kolida S, Ndagijimana M, Laghi L, Calanni F, Brigidi P, Gibson GR, Costabile A. Rifaximin modulates the colonic microbiota of patients with Crohn’s disease: an in vitro approach using a continuous culture colonic model system. J Antimicrob Chemother. 2010;65:2556-2565. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 120] [Cited by in RCA: 143] [Article Influence: 9.5] [Reference Citation Analysis (1)] |
| 79. | Xu D, Gao J, Gillilland M 3rd, Wu X, Song I, Kao JY, Owyang C. Rifaximin alters intestinal bacteria and prevents stress-induced gut inflammation and visceral hyperalgesia in rats. Gastroenterology. 2014;146:484-496.e4. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 205] [Article Influence: 18.6] [Reference Citation Analysis (0)] |
| 80. | Soldi S, Vasileiadis S, Uggeri F, Campanale M, Morelli L, Fogli MV, Calanni F, Grimaldi M, Gasbarrini A. Modulation of the gut microbiota composition by rifaximin in non-constipated irritable bowel syndrome patients: a molecular approach. Clin Exp Gastroenterol. 2015;8:309-325. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 61] [Cited by in RCA: 73] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 81. | Ponziani FR, Scaldaferri F, Petito V, Paroni Sterbini F, Pecere S, Lopetuso LR, Palladini A, Gerardi V, Masucci L, Pompili M. The Role of Antibiotics in Gut Microbiota Modulation: The Eubiotic Effects of Rifaximin. Dig Dis. 2016;34:269-278. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 92] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 82. | Finegold SM, Molitoris D, Väisänen ML. Study of the in vitro activities of rifaximin and comparator agents against 536 anaerobic intestinal bacteria from the perspective of potential utility in pathology involving bowel flora. Antimicrob Agents Chemother. 2009;53:281-286. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 29] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 83. | Gillis JC, Brogden RN. Rifaximin. A review of its antibacterial activity, pharmacokinetic properties and therapeutic potential in conditions mediated by gastrointestinal bacteria. Drugs. 1995;49:467-484. [PubMed] |
| 84. | Vitali B, Turroni S, Serina S, Sosio M, Vannini L, Candela M, Guerzoni ME, Brigidi P. Molecular and phenotypic traits of in-vitro-selected mutants of Bifidobacterium resistant to rifaximin. Int J Antimicrob Agents. 2008;31:555-560. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 16] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 85. | Vitali B, Turroni S, Dal Piaz F, Candela M, Wasinger V, Brigidi P. Genetic and proteomic characterization of rifaximin resistance in Bifidobacterium infantis BI07. Res Microbiol. 2007;158:355-362. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 13] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 86. | De Leo C, Eftimiadi C, Schito GC. Rapid disappearance from the intestinal tract of bacteria resistant to rifaximin. Drugs Exp Clin Res. 1986;12:979-981. [PubMed] |
| 87. | Blandizzi C, Viscomi GC, Marzo A, Scarpignato C. Is generic rifaximin still a poorly absorbed antibiotic? A comparison of branded and generic formulations in healthy volunteers. Pharmacol Res. 2014;85:39-44. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 29] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 88. | Highlights of prescribing information. Available from: https://www.accessdata.fda.gov/drugsatfda_docs/label/2014/021361s013lbl.pdf. |
| 89. | Trapnell CB, Connolly M, Pentikis H, Forbes WP, Bettenhausen DK. Absence of effect of oral rifaximin on the pharmacokinetics of ethinyl estradiol/norgestimate in healthy females. Ann Pharmacother. 2007;41:222-228. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 31] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 90. | Pentikis HS, Connolly M, Trapnell CB, Forbes WP, Bettenhausen DK. The effect of multiple-dose, oral rifaximin on the pharmacokinetics of intravenous and oral midazolam in healthy volunteers. Pharmacotherapy. 2007;27:1361-1369. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 35] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 91. | Bass NM, Mullen KD, Sanyal A, Poordad F, Neff G, Leevy CB, Sigal S, Sheikh MY, Beavers K, Frederick T. Rifaximin treatment in hepatic encephalopathy. N Engl J Med. 2010;362:1071-1081. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 868] [Cited by in RCA: 869] [Article Influence: 57.9] [Reference Citation Analysis (0)] |
| 92. | Scarpellini E, Giorgio V, Gabrielli M, Filoni S, Vitale G, Tortora A, Ojetti V, Gigante G, Fundarò C, Gasbarrini A. Rifaximin treatment for small intestinal bacterial overgrowth in children with irritable bowel syndrome. Eur Rev Med Pharmacol Sci. 2013;17:1314-1320. [PubMed] |
| 93. | Ponziani FR, Pecere S, Lopetuso L, Scaldaferri F, Cammarota G, Gasbarrini A. Rifaximin for the treatment of irritable bowel syndrome - a drug safety evaluation. Expert Opin Drug Saf. 2016;15:983-991. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 18] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 94. | Stecher B, Maier L, Hardt WD. ‘Blooming’ in the gut: how dysbiosis might contribute to pathogen evolution. Nat Rev Microbiol. 2013;11:277-284. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 227] [Cited by in RCA: 263] [Article Influence: 21.9] [Reference Citation Analysis (0)] |