Published online Dec 7, 2016. doi: 10.3748/wjg.v22.i45.10053
Peer-review started: August 2, 2016
First decision: August 19, 2016
Revised: September 7, 2016
Accepted: October 26, 2016
Article in press: October 26, 2016
Published online: December 7, 2016
Processing time: 126 Days and 21.6 Hours
To determine the influence of Smoc2 on hepatocellular carcinoma (HCC) cell proliferation and to find a possible new therapeutic target for preventing HCC progression.
We detected expression of Smoc2 in HCC tissues and corresponding non-tumor liver (CNL) tissues using PCR, western blot, and immunohistochemistry methods. Subsequently, we down-regulated and up-regulated Smoc2 expression using siRNA and lentivirus transfection assay, respectively. Then, we identified the effect of Smoc2 on cell proliferation and cell cycle using CCK-8 and flow cytometry, respectively. The common cell growth signaling influenced by Smoc2 was detected by western blot assay.
The expression of Smoc2 was significantly higher in HCC tissues compared with CNL tissues. Overexpression of Smoc2 promoted HCC cell proliferation and cell cycle progression. Down-regulation of Smoc2 led to inhibition of cell proliferation and cell cycle progression. Smoc2 had positive effect on ERK and AKT signaling.
Smoc2 promotes the proliferation of HCC cells through accelerating cell cycle progression and might act as an anti-cancer therapeutic target in the future.
Core tip: In our study, we confirmed that Smoc2 was up-regulated in hepatocellular carcinoma (HCC) tissues and played an important role in regulating liver cancer cell proliferation. Besides, we verified that Smoc2 participated in promoting HCC cell proliferation mainly through regulation of cell cycle progression. We have not investigated the promotive role of Smoc2 in regulating cell proliferation, whether it is through cell cycle regulation only or involves regulation of cell apoptosis as well. Moreover, the exact mechanism of how Smoc2 regulates cell cycle remains unclear. The core contents of our study included Smoc2 promotion of HCC cell proliferation via accelerating cell cycle progression.
- Citation: Su JR, Kuai JH, Li YQ. Smoc2 potentiates proliferation of hepatocellular carcinoma cells via promotion of cell cycle progression. World J Gastroenterol 2016; 22(45): 10053-10063
- URL: https://www.wjgnet.com/1007-9327/full/v22/i45/10053.htm
- DOI: https://dx.doi.org/10.3748/wjg.v22.i45.10053
Hepatocellular carcinoma (HCC) is one of the most common malignancies worldwide, with high mortality rate and low early diagnostic rate[1]. HBV infection, alcohol abuse, aflatoxin exposure and HCV infection are identified as major causes of HCC. The current therapies available for HCC include surgery, interventional therapy, radio frequency therapy, radiotherapy, biological target therapy and so on[2]. All of these treatments have certain curative effects, but have inherent limitations and adverse effects, especially for HCC patients at the advanced stage[3]. Thus, it is urgent to find new treatment target for the sake of enhancing curative effect and reducing adverse effects, especially in advanced HCC patients.
Apart from the common etiologies of HCC listed above, certain oncogenes, cytokines, neurotransmitters, chemokines, extracellular secretory proteins and tumor microenvironment are thought to play important roles in origin and progression of HCC[4]. Therefore, oncogenes and tumor microenvironment, which facilitate HCC progression, can be chosen as therapeutic targets for HCC treatment[5].
The secreted protein acidic and rich in cysteine (SPARC; alternative names: osteonectin; ON or basement membrane-40; BM-40) family is recognized as extracellular matrix proteins[6]. A differential expression of SPARC in tumor tissue and its surrounding stroma compared to normal tissues has been reported for many different types of cancer[7]. And, SPARC was found to be up-regulated in several solid tumors and to facilitate tumor metastasis[8].
Secreted modular calcium-binding protein-2 (Smoc2) is a novel member of the SPARC family[9]. Previous study confirmed that Smoc2 could promote cell cycle progression of human umbilical vein endothelial cells by inducing the expression of transcripts required for cell cycle[10]. Other studies have shown that Smoc2 is necessary for DNA synthesis in the cell cycle and is likely to impact cell growth in vitro and in vivo[11]. However, to the best of our knowledge, no study has been conducted on the role of Smoc2 in HCC.
Thus, the present study was designed to investigate the effect of Smoc2 on proliferation of HCC cells and the impact on cell cycle. Our study showed that Smoc2 promoted proliferation of HCC cells and accelerated cell cycle progression. The results suggested that Smoc2 might act as an anti-cancer therapeutic target for HCC treatment.
A total of 20 pairs of HCC tissues and corresponding non-tumor liver (CNL) tissues were obtained from the Liver Surgery Department of the corresponding hospital. All the human liver tissues were obtained with informed consent and the study was approved by the Ethical Committee of the corresponding hospital.
The paraffin-embedded human liver tissues were incised into 4-5 μm thickness slices and then dewaxed using xylene and ethanol, in a stepwise manner. After dewaxing, the slices were rehydrated for subsequent staining. For immunohistochemistry (IHC) staining, the slices were treated with hydrogen peroxide and boiled for 15 min in citrate solution for antigen retrieval. When the slices had cooled naturally to room temperature, we added goat serum for blocking of unrelated antigens. Afterwards, the slices were incubated with Smoc2 antibody (Abcam) at 4 °C overnight. The following day, the slices were washed three times with phosphate buffer and incubated with horseradish peroxidase-labelled secondary antibody for 1 h at room temperature. The slices were then developed using DAB substrate liquid (Thermo) and dehydrated by ethanol, in a stepwise manner, and finally sealed with neutral balsam.
The tissue protein was isolated from human liver tisseus using T-PER Tissue Protein Extraction Reagent (Pierce Biotechnology) according to protocols provided by the manufacturer. The cell protein was lysed using lysis buffer that contained Tris-HCl, NaCl, Triton-X 100, MgCl2, PMSF and so on. The cell lysate, which was used for cell signaling detection, was obtained by IP cell lysis solution (Beyotime Biotechnology). All the proteins were separated by SDS-PAGE and transferred to a nitrocellulose membrane under constant current condition. Then, the membrane was blocked using 5% bovine serum albumin at room temperature for 1 h. The nitrocellulose membrane was then incubated using antibodies for Smoc2 (Abcam), extracellular regulated protein kinase (ERK; Cell Signaling Technology, CST), phospho-ERK (CST), AKT (CST), phospho-AKT (CST), Src (CST), phospho-Src (CST), FAK (CST), phospho-FAK (CST) and GAPDH (Sigma) at 4 °C overnight. The next day, the nitrocellulose membrane was incubated with the fluorescence-conjugated secondary antibodies at room temperature for 1 h. All the fluorescence signals were captured and saved by the Odyssey imaging system (LI-COR). The images of western blots were analyzed using ImageJ software for gray value calculation.
For cell staining, SMMC-7721 cells transfected with Smoc2 vector using lentivirus method were seeded on rounded slides in 24-well plates and incubated at 37 °C and a 5% CO2 atmosphere overnight. The next day, cells were fixed using paraformaldehyde and washed three times by phosphate buffer before staining. Then, the sildes were incubated with Smoc2 antibody (Abcam) for 75 min at room temperature. After that, the slides were washed three times using phosphate buffer and incubated with Alexa Fluor 488nm-conjugated secondary antibody. The nuclei were stained using DAPI (Sigma) and the immunofluorescence stain images were record using fluorescence microscope (Carl Zeiss).
Total RNA was isolated from fresh liver tissues using the Trizol Reagent (Takara). The isolated RNA was then immediately applied to the reverse transcription reaction. Primers used for quantitative PCR reaction were: forward primer, 5’-GCTCACGTTCTTGAGAGTCG-3’; and reverse primer, 5’-TGTAGCTGTGACACTGGACC-3’. The PCR reaction conditions were 30 s at 94 °C, 1 min at 57 °C and 1 min and 30 s at 72 °C for 35 cycles. The Amplitaq polymerase and related reagents were supplied by Takara.
The full-length cDNA cloning vector was purchased from GeneCopoeia and used for amplification template. The Smoc2 full-length cDNA was amplified with NheI and BamHI restriction sites using KOD-PLUS Neo polymerase. Then, the PCR-amplified cDNA was cloned into the pcDNA3.1 vector using restriction enzymes and DNA ligase. The recombined vector was packed with VSV, REV and GAG vectors and transfected into 293T cells using the Lipofectamine 2000 Reagent (Invitrogen) to obtain virus supernatent. The virus supernatent was added into SMMC7721 cells and Huh7 cells with polybrene. The transfected SMMC7721 cells and Huh7 cells were cultured under puromycin condition and verified by western blot assay.
MHCC-97H cells and HCC-LM3 cells were transfected with siRNA duplexes against Smoc2 or with control RNA duplex oligonucleotides using Lipofectamine 2000, following the manufacturer’s instructions. The Smoc2 interference target sequence was TTAAGAGGTTCCTGCGAAA.
MHCC-97H and HCC-LM3 cells transfected with Smoc2-SiRNA were seeded into 96-well plates at density of 3000 cells per 100 μL and cultured for 5 d in vitro. In addition, the SMMC-7721 and HCC-LM3 cells transfected with Smoc2 lentivirus were seeded into 96-well plates at density of 2000 cells per 100 μL and cultured for 5 d at 37 °C and in 5% CO2 atmosphere. The CCK-8 reagent was added to each well at 10 μL volume and reacted for 1 h in light-free condition, at 37 °C, and in 5% CO2 atmosphere. Then, the absorbance value was detected using a microplate reader. The absorbance value at 450 nm was recorded.
The cells were seeded into 6-well plates and collected at logarithmic growth phase. The cells were cultured in serum-free condition overnight before collection. Liver cancer cells were collected and washed three times using phosphate buffer. Then, the cells were centrifuged at 1000 r/min for 5 min and resuspended in 1 mL phosphate buffer. Next, the cells were added to 9 mL 70% cool ethanol, slowly and carefully. The fixed cells were then stored at -20 °C for at least 24 h. For cell cycle assay, the fixed cells were centrifuged at 1000 r/min for 10 min and washed three times using cool phosphate buffer. Then, we added 0.5 mL RNase (50 μg/mL) to each tube and incubated for 20 min at 37 °C. Lastly, we added propidium iodide at 50 μg/mL to each tube and stained on ice for 30 min in light-free condition. The stained cells were detected using flow cytometry (BD Biosciences).
We used SPSS 16.0 software to analyze the statistical significance of differences in our study. Statistic differences were calculated using two-tailed Student’s t-test. P < 0.05 was considered statistically significant and P < 0.01 was considered very statistically significant.
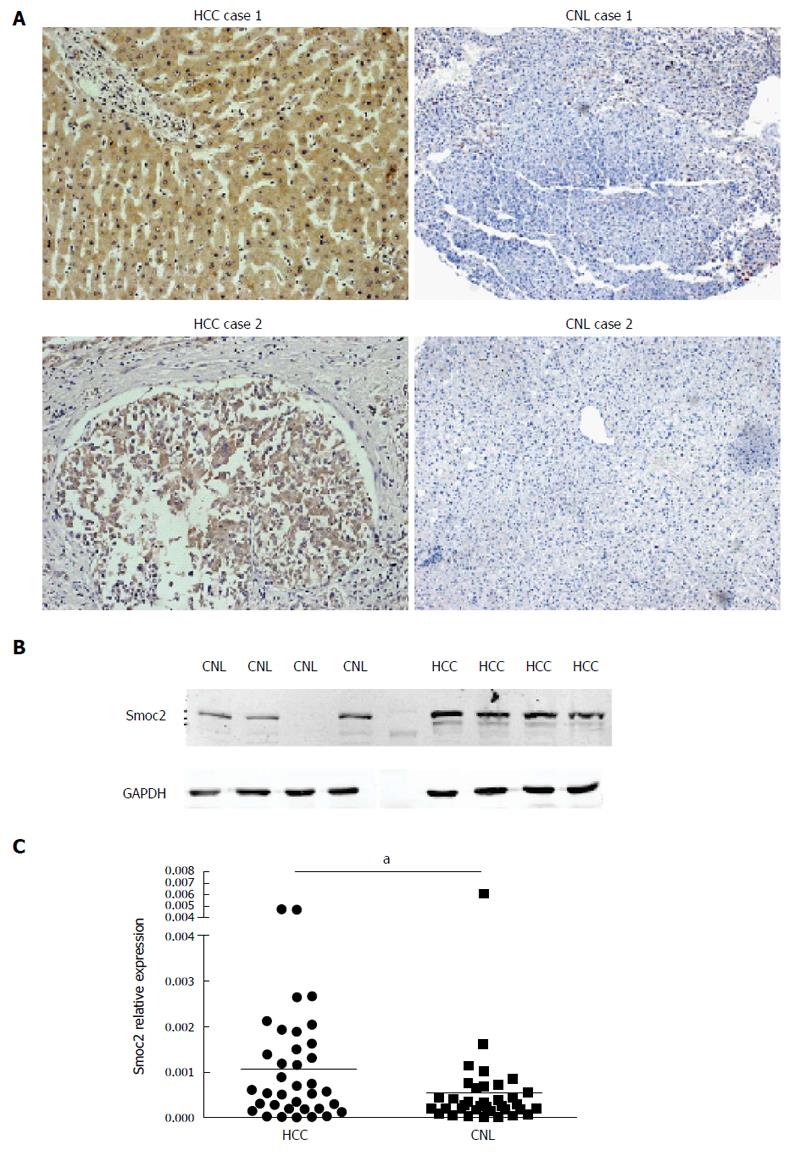
The expression of Smoc2 was significantly up-regulated in HCC tissues, compared to CNL tissues, as evidenced by IHC (Figure 1A). IHC results showed that expression of Smoc2 was mainly located in the cytoplasm of HCC cells and the extracellular lesion of liver tissues. Western blot assay showed that protein expression level of Smoc2 was significantly higher in human HCC tissues, compared to CNL tissues (Figure 1B). The real-time quantitative PCR result indicated that mRNA expression level of Smoc2 in HCC tissues was remarkably higher than in CNL tissues. All the results above revealed that expression of Smoc2 was up-regulated in HCC tissues, compared to CNL tissues, at both protein and mRNA levels (Figure 1C).
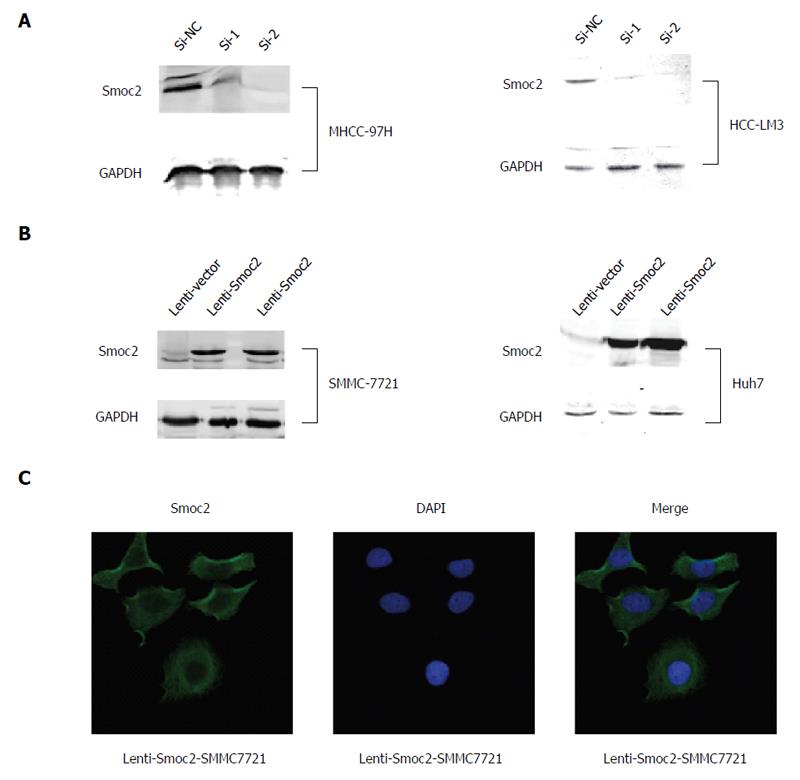
We carried out siRNA transfection for silencing of Smoc2 in MHCC-97H and HCC-LM3 cells, and verified the silencing effect using western blot assay (Figure 2A). We induced overexpression of Smoc2 in SMMC-7721 and Huh7 cells using the lentivirus transfection method and identified the overexpressing effect using western blot assay (Figure 2B). The immunofluorescence stain results showed that expression of Smoc2 induced by lentivirus transfection can be found in cytoplasm of SMMC-7721 cells (Figure 2C).
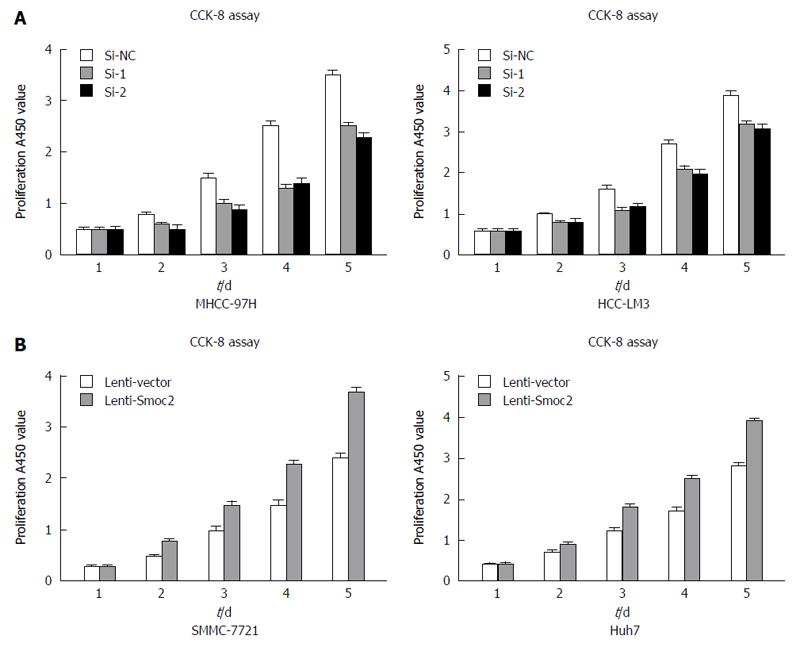
CCK-8 assay directly reflects cell viability and can be used for evaluating cell proliferation. We collected MHCC-97H and HCC-LM3 cells transfected with Smoc2-siRNA at 48 h and detected the cell viability for 5 d in vitro using CCK-8 assay. The results showed that the proliferation capacity of MHCC-97H and HCC-LM3 cells transfected with Smoc2-siRNA were significantly worse than that in the control group (Figure 3A). Besides, the proliferative capacity in Smoc2 overexpressing SMMC-7721 and Huh7 cells was remarkably stronger than in control groups in vitro (Figure 3B). These results suggested that Smoc2 plays an important role in promoting HCC cells proliferation in vitro.
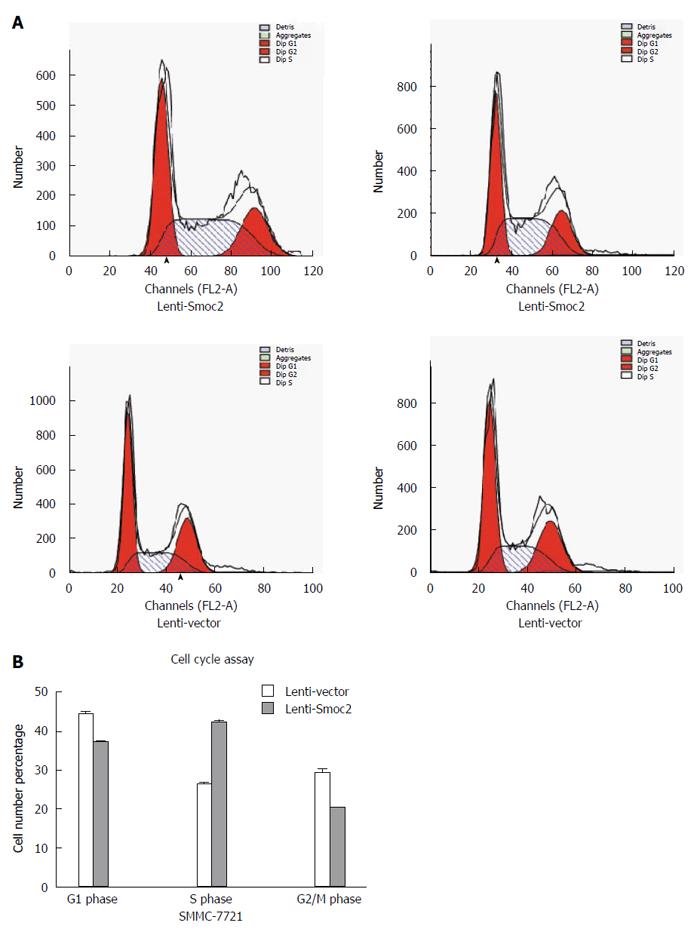
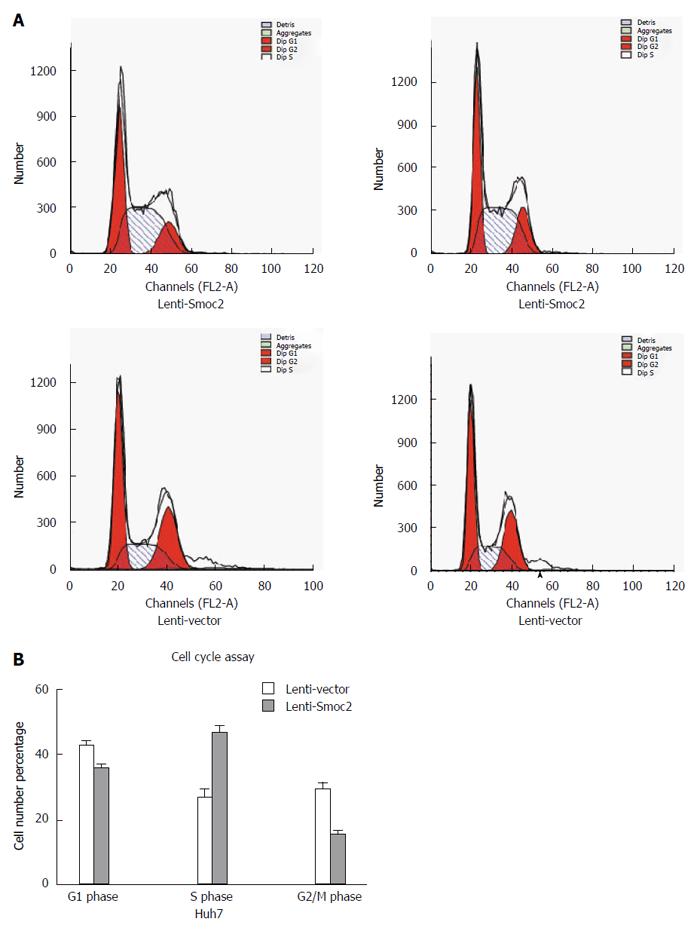
Cell cycle directly reflects cell proliferation capacity and is composed of G0/G1 phase, S phase and G2/M phase. The cell proliferation capacity can be reflected by proliferation index (PI) and S phase cell fraction (SPF). PI equates to the percentage of sum of S phase and G2/M phase cells in total cell cycle cells. SPF equates to the percentage of S phase cells in total cell cycle (G0/G1 + S + G2/M). Our results showed that both PI and SPF values in overexpressing Smoc2 cells were higher than those in the control group, which indicated the promotive role of Smoc2 in cell cycle progression and cell proliferation (Figure 4A and B; Figure 5A and B).
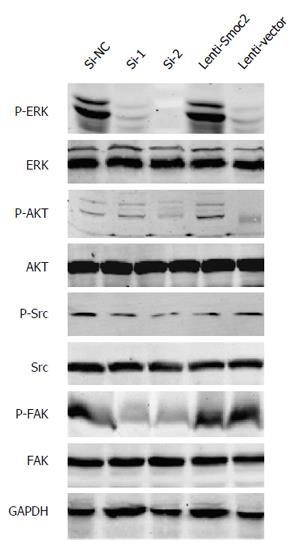
The ERKs consist of ERK1 and ERK2, having molecular weights of 44 kDa and 42 kDa respectively[12]. Phosphorylation of ERK allows translocation from the cytoplasm to the nucleus, and mediates transcriptional activation of Elk-1, ATF, NF-κB, Ap-1, c-fos and c-Jun. Activation of ERK signaling participates in regulating cell proliferation and differentiation, construction of the cell skeleton, and the processes of cell apoptosis and cell canceration[13]. Our results showed that phosphorylation of ERK was up-regulated in Smoc2 overexpressed cells and down-regulated in Smoc2 siRNA interfered cells (Figure 6).
Signaling by AKT, also known as protein kinase B, plays an important role in regulating cell survival and apoptosis[14]. Our results indicated that phosphorylation of AKT was up-regulated in Smoc2 overexpressed cells and down-regulated in Smoc2 siRNA interfered cells (Figure 6).
FAK, also known as focal adhesion kinase, and its activation is related to cell proliferation and cell cycle[15]. Inhibition of FAK signaling can cause decrease of S phase cell ratio in cell cycle. Thus, inhibition of FAK signaling lead to inhibition of cell proliferation and enhancement of cell apoptosis. Our results illustrated that down-regulation of Smoc2 by siRNA transfection caused inhibition of FAK phosphorylation (Figure 6).
As one of the most frequent human cancers worldwide, the incidence and progression of HCC are affected by oncogenes and tumor microenvironment[16,17]. Cytokines, chemokines, pH value, hypoxia, fibroblast and extracellular matrix proteins are important components of the tumor microenvironment[18-21]. The tumor microenvironment plays an important role in regulating tumor progression[22,23]. It has been reported that many extracellular matrix proteins exert significant influence on tumor progression. Thus, we focused on finding a new secretory protein which can cause tumor development.
The Smoc2 gene, encoding a protein belonging to the SPARC family and identified as an extracellular matrix protein, is up-regulated during embryogenesis and wound healing[24]. The Smoc2 gene encoded protein promotes extracellular matrix assembly and potentiates endothelial cell proliferation and migration[25], and has pro-angiogenic activity[26]. It was reported that the Smoc2 gene is indispensable for VEGF-induced vascular endothelial cells mitogenesis and tube formation[27,28]. Due to its promotive effect on angiogenesis, the Smoc2 gene may represent a useful target for inhibition of tumor progress.
The cell cycle consists of G0/G1 phase, S phase and G2/M phase in mammalian cells[29,30]. The cell cycle is mainly regulated by various kinds of growth factors, such as platelet-derived growth factor[31,32]. Cyclin D1 expression is induced by mitogens and is crucial for G1 progression[33]. Previous results showed that Smoc2 plays an important role in promoting G1/S progression through participating in the maintenance of cyclin D1 expression[34-36]. In other words, cyclin D1 can be considered as an effector of Smoc2-induced DNA synthesis[37,38].
It was also demonstrated that Smoc2 was necessary for DNA synthesis in response to PDGF and other growth factors during the cell cycle[39-41]. Our study demonstrated that Smoc2 had a promotive effect on HCC cell G1/S phase progression, as well as on cell proliferation. The S phase cell ratio was significantly elevated in Smoc2 overexpressing HCC cells, and this result was in accordance with those from a previous study using human umbilical vein endothelial cells transduced with Ad-Smoc2[42,43].
Compared with Swiss 3T3 cells transfected with Sicontrol, the cells transfected with SiSmoc2 oligonucleotide duplexes showed no significant change in phospho-MAPK and phospho-AKT levels[44-47]. Unlike the results in Swiss 3T3 cells, our results indicated that Smoc2 had promotive influence on MAPK/ERK and AKT signaling pathways in HCC cells. MAPK/ERK and AKT are identified as PDGF β receptor effector kinases and positively regulate the expression of cyclin D1[48]. Therefore, we speculated that Smoc2 could potentiate PDGF-induced mitogenesis and the subsequent cell proliferation. Nevertheless, another study based on Swiss 3T3 cells showed that the promotion of Smoc2 in DNA synthesis was independent of PDGF-binding activity but dependent on integrin-activated protein kinase[49].
Previous studies revealed that Smoc2 promotes growth factor-induced DNA synthesis and differs from SPARC, which was identified as an anti-mitogenic factor. It was reported that SPARC could bind various kinds of mitogens (such as VEGF, PDGF, FGF and so on) but inhibit the signaling mediated by these growth factors. Unlike SPARC, Smoc2 could facilitate growth factor-induced DNA synthesis[50]. The exact mechanism of how Smoc2 promotes growth factor-induced DNA synthesis in HCC cells remains unclear and needs to be studied further.
Our research focused on the proliferation promotion effect of Smoc2 in HCC cells. However, Smoc2 also has influence on cell migration. It has been reported that extracellular matrix protein Smoc2 can promote keratinocyte migration in vitro. Accordingly, the role of Smoc2 on HCC cell migration and invasion need to be further studied. The influences of Smoc2 on HCC cell biological behaviors may be extensive.
In conclusion, our results demonstrated that Smoc2 is an important regulator of cell mitogenesis, and plays a promotive role in growth factor signaling and cell cycle progression in HCC. Based on our study, Smoc2 might represent a promising therapeutic target for HCC treatment.
The secreted modular calcium-binding protein-2 (Smoc2) gene, belonging to the secreted protein acidic and rich in cysteine (SPARC) family, was identified as encoding an extracellular matrix protein and as being up-regulated during embryogenesis and wound healing. The Smoc2 gene encoded protein promotes extracellular matrix assembly and potentiates endothelial cell proliferation and migration, as well as has pro-angiogenic activity. Due to the promotive effect on angiogenesis, the Smoc2 gene can be chosen as a target for inhibition of tumor progression.
Smoc2 is a novel member of the SPARC family. Current studies have confirmed that Smoc2 can promote cell cycle progression by inducing the expression of transcripts required for the cell cycle. Other studies have shown that Smoc2 is necessary for DNA synthesis in the cell cycle and is likely to impact cell growth in vitro and in vivo. These studies also revealed that Smoc2 plays an important role in regulating the cell cycle. However, to the best of our knowledge, no study has investigated the role of Smoc2 in hepatocellular carcinoma (HCC).
The expression of Smoc2 was significantly higher in HCC tissues, as compared to CNL tissues. Overexpression of Smoc2 promoted HCC cell proliferation and cell cycle progression. Down-regulation of Smoc2 led to inhibition of cell proliferation and cell cycle progression. Smoc2 had a positive effect on ERK and AKT signaling.
According to our studies, Smoc2 was identified as pro-proliferative gene in HCC. Besides, blockage of Smoc2 could result in inhibition of liver cancer cell proliferation. Therefore, Smoc2 might be chosen as a target gene for preventing HCC progression. In the future, a neutralizing antibody for Smoc2 could be produced and used in animal experiments to further test its efficacy for preventing HCC proliferation. Blockage of Smoc2, by using neutralizing antibody or another approach, may contribute to HCC treatment and improve therapeutic efficacy.
The SPARC family was recognized as extracellular matrix proteins. SPARC has been found to be up-regulated in several solid tumor tissues and to facilitate tumor metastasis. Smoc2, a novel member of the SPARC family, was identified as an extracellular matrix protein.
This study is very interesting. The authors determined the influence of Smoc2 on HCC cell proliferation and aimed to find a possible new therapeutic target for preventing HCC progression. The authors detected expression of Smoc2 in HCC tissues and corresponding non-tumor liver (CNL) tissues using PCR, western blot, immunohistochemistry methods and down-regulation and up-regulation of Smoc2 expression using small interfering RNA and lentivirus transfection assay respectively. The effect of Smoc2 on cell proliferation and cell cycle was identified using CCK-8 and flow cytometry respectively. The common cell growth signaling influenced by Smoc2 was detected. The authors found that Smoc2 promotes the proliferation of HCC cells through accelerating cell cycle progression and might act as an anti-cancer therapeutic target in the future. Overall, this study is well designed and the manuscript is well written.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: China
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): B, B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Deepak P, Pratschke S S- Editor: Qi Y L- Editor: Filipodia E- Editor: Wang CH
| 1. | de Castro Brás LE, Toba H, Baicu CF, Zile MR, Weintraub ST, Lindsey ML, Bradshaw AD. Age and SPARC change the extracellular matrix composition of the left ventricle. Biomed Res Int. 2014;2014:810562. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 32] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 2. | Zhu GC, Gao L, He J, Long Y, Liao S, Wang H, Li X, Yi W, Pei Z, Wu M. CD90 is upregulated in gastric cancer tissues and inhibits gastric cancer cell apoptosis by modulating the expression level of SPARC protein. Oncol Rep. 2015;34:2497-2506. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 14] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 3. | Gao YY, Han RB, Wang X, Ge SH, Li HL, Deng T, Liu R, Bai M, Zhou LK, Zhang XY. Change of SPARC expression after chemotherapy in gastric cancer. Cancer Biol Med. 2015;12:33-40. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |
| 4. | Yang L, Rong W, Xiao T, Zhang Y, Xu B, Liu Y, Wang L, Wu F, Qi J, Zhao X. Secretory/releasing proteome-based identification of plasma biomarkers in HBV-associated hepatocellular carcinoma. Sci China Life Sci. 2013;56:638-646. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 23] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 5. | Won C, Kim BH, Yi EH, Choi KJ, Kim EK, Jeong JM, Lee JH, Jang JJ, Yoon JH, Jeong WI. Signal transducer and activator of transcription 3-mediated CD133 up-regulation contributes to promotion of hepatocellular carcinoma. Hepatology. 2015;62:1160-1173. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 148] [Cited by in RCA: 151] [Article Influence: 15.1] [Reference Citation Analysis (0)] |
| 6. | Deng B, Qu L, Li J, Fang J, Yang S, Cao Z, Mei Z, Sun X. MiRNA-211 suppresses cell proliferation, migration and invasion by targeting SPARC in human hepatocellular carcinoma. Sci Rep. 2016;6:26679. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 38] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 7. | Lindner JL, Loibl S, Denkert C, Ataseven B, Fasching PA, Pfitzner BM, Gerber B, Gade S, Darb-Esfahani S, Sinn BV. Expression of secreted protein acidic and rich in cysteine (SPARC) in breast cancer and response to neoadjuvant chemotherapy. Ann Oncol. 2015;26:95-100. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 58] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 8. | Nian Q, Zhang Z, Wei C, Kuang X, Wang X, Wang L. Gene expression profiling in myelodysplastic syndrome after SPARC overexpression associated with Ara-C. Oncol Rep. 2015;34:2072-2082. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 9. | Pazin DE, Albrecht KH. Developmental expression of Smoc1 and Smoc2 suggests potential roles in fetal gonad and reproductive tract differentiation. Dev Dyn. 2009;238:2877-2890. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 40] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 10. | Liu P, Pazin DE, Merson RR, Albrecht KH, Vaziri C. The developmentally-regulated Smoc2 gene is repressed by Aryl-hydrocarbon receptor (Ahr) signaling. Gene. 2009;433:72-80. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 13] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 11. | Plentz RR, Malek NP. Early Detection of Hepatocellular Carcinoma: How to Screen and Follow up Patients with Liver Cirrhosis According to the GERMAN S3 Guideline? Diagnostics (Basel). 2015;5:497-503. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 9] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 12. | Birlea SA, Gowan K, Fain PR, Spritz RA. Genome-wide association study of generalized vitiligo in an isolated European founder population identifies SMOC2, in close proximity to IDDM8. J Invest Dermatol. 2010;130:798-803. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 65] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 13. | Lin CY, Hu CT, Cheng CC, Lee MC, Pan SM, Lin TY, Wu WS. Oxidation of heat shock protein 60 and protein disulfide isomerase activates ERK and migration of human hepatocellular carcinoma HepG2. Oncotarget. 2016;7:11067-11082. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 21] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 14. | Wen S, Liu Y, Yang M, Yang K, Huang J, Feng D. Increased NEK2 in hepatocellular carcinoma promotes cancer progression and drug resistance by promoting PP1/Akt and Wnt activation. Oncol Rep. 2016;36:2193-2199. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 23] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 15. | Mattos AA, Marcon Pdos S, Araújo FS, Coral GP, Tovo CV. Hepatocellular carcinoma in a non-cirrhotic patient with sustained virological response after hepatitis c treatment. Rev Inst Med Trop Sao Paulo. 2015;57:519-522. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 6] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 16. | Liu D, Staveley-O’Carroll KF, Li G. Immune-based Therapy Clinical Trials in Hepatocellular Carcinoma. J Clin Cell Immunol. 2015;6. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 16] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 17. | He M, Qin H, Poon TC, Sze SC, Ding X, Co NN, Ngai SM, Chan TF, Wong N. Hepatocellular carcinoma-derived exosomes promote motility of immortalized hepatocyte through transfer of oncogenic proteins and RNAs. Carcinogenesis. 2015;36:1008-1018. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 171] [Cited by in RCA: 208] [Article Influence: 20.8] [Reference Citation Analysis (0)] |
| 18. | Shahab J, Baratta C, Scuric B, Godt D, Venken KJ, Ringuette MJ. Loss of SPARC dysregulates basal lamina assembly to disrupt larval fat body homeostasis in Drosophila melanogaster. Dev Dyn. 2015;244:540-552. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 31] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 19. | Ehninger A, Boch T, Medyouf H, Müdder K, Orend G, Trumpp A. Loss of SPARC protects hematopoietic stem cells from chemotherapy toxicity by accelerating their return to quiescence. Blood. 2014;123:4054-4063. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 21] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 20. | Qiu F, Sun R, Deng N, Guo T, Cao Y, Yu Y, Wang X, Zou B, Zhang S, Jing T. miR-29a/b enhances cell migration and invasion in nasopharyngeal carcinoma progression by regulating SPARC and COL3A1 gene expression. PLoS One. 2015;10:e0120969. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 50] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 21. | Nagaraju GP, Dontula R, El-Rayes BF, Lakka SS. Molecular mechanisms underlying the divergent roles of SPARC in human carcinogenesis. Carcinogenesis. 2014;35:967-973. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 102] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 22. | Liao CY, Lee CC, Tsai CC, Hsueh CW, Wang CC, Chen IH, Tsai MK, Liu MY, Hsieh AT, Su KJ. Novel Investigations of Flavonoids as Chemopreventive Agents for Hepatocellular Carcinoma. Biomed Res Int. 2015;2015:840542. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 34] [Cited by in RCA: 49] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 23. | Aseer KR, Kim SW, Choi MS, Yun JW. Opposite Expression of SPARC between the Liver and Pancreas in Streptozotocin-Induced Diabetic Rats. PLoS One. 2015;10:e0131189. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 13] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 24. | Shi D, Jiang K, Fu Y, Fang R, Liu XI, Chen J. Overexpression of SPARC correlates with poor prognosis in patients with cervical carcinoma and regulates cancer cell epithelial-mesenchymal transition. Oncol Lett. 2016;11:3251-3258. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 25] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 25. | Cheng CT, Chu YY, Yeh CN, Huang SC, Chen MH, Wang SY, Tsai CY, Chiang KC, Chen YY, Ma MC. Peritumoral SPARC expression and patient outcome with resectable intrahepatic cholangiocarcinoma. Onco Targets Ther. 2015;8:1899-1907. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 6] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 26. | Villarreal G, Chatterjee A, Oh SS, Oh DJ, Rhee DJ. Pharmacological regulation of SPARC by lovastatin in human trabecular meshwork cells. Invest Ophthalmol Vis Sci. 2014;55:1657-1665. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 18] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 27. | Stenzel A, Goss KU, Endo S. Prediction of partition coefficients for complex environmental contaminants: Validation of COSMOtherm, ABSOLV, and SPARC. Environ Toxicol Chem. 2014;33:1537-1543. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 31] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 28. | Nakashima S, Kobayashi S, Sakai D, Tomokuni A, Tomimaru Y, Hama N, Wada H, Kawamoto K, Marubashi S, Eguchi H. Prognostic impact of tumoral and/or peri-tumoral stromal SPARC expressions after surgery in patients with biliary tract cancer. J Surg Oncol. 2014;110:1016-1022. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 12] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 29. | Han W, Cao F, Chen MB, Lu RZ, Wang HB, Yu M, Shi CT, Ding HZ. Prognostic Value of SPARC in Patients with Pancreatic Cancer: A Systematic Review and Meta-Analysis. PLoS One. 2016;11:e0145803. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 32] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 30. | Joo I, Lee JM. Recent Advances in the Imaging Diagnosis of Hepatocellular Carcinoma: Value of Gadoxetic Acid-Enhanced MRI. Liver Cancer. 2016;5:67-87. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 51] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 31. | Alfawaz S, Fong F, Plagnol V, Wong FS, Fearne J, Kelsell DP. Recessive oligodontia linked to a homozygous loss-of-function mutation in the SMOC2 gene. Arch Oral Biol. 2013;58:462-466. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 32] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 32. | Hua HW, Jiang F, Huang Q, Liao ZJ, Ding G. Re-sensitization of 5-FU resistance by SPARC through negative regulation of glucose metabolism in hepatocellular carcinoma. Tumour Biol. 2015;36:303-313. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 9] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 33. | Bradshaw AD. The role of secreted protein acidic and rich in cysteine (SPARC) in cardiac repair and fibrosis: Does expression of SPARC by macrophages influence outcomes? J Mol Cell Cardiol. 2016;93:156-161. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 42] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 34. | Morgan SS, Nagle RB, Cranmer LD. Serum protein acidic and rich in cysteine (SPARC) as a prognostic marker in soft tissue sarcomas. Clin Sarcoma Res. 2014;4:2. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 35. | Alkhateeb A, Al-Dain Marzouka N, Qarqaz F. SMOC2 gene variant and the risk of vitiligo in Jordanian Arabs. Eur J Dermatol. 2010;20:701-704. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 17] [Reference Citation Analysis (0)] |
| 36. | Mommaerts H, Esguerra CV, Hartmann U, Luyten FP, Tylzanowski P. Smoc2 modulates embryonic myelopoiesis during zebrafish development. Dev Dyn. 2014;243:1375-1390. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 20] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 37. | Peixoto E, Atorrasagasti C, Aquino JB, Militello R, Bayo J, Fiore E, Piccioni F, Salvatierra E, Alaniz L, García MG. SPARC (secreted protein acidic and rich in cysteine) knockdown protects mice from acute liver injury by reducing vascular endothelial cell damage. Gene Ther. 2015;22:9-19. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 19] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 38. | Salvatierra E, Alvarez MJ, Leishman CC, Rivas Baquero E, Lutzky VP, Chuluyan HE, Podhajcer OL. SPARC Controls Melanoma Cell Plasticity through Rac1. PLoS One. 2015;10:e0134714. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 12] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 39. | Sinn M, Sinn BV, Striefler JK, Lindner JL, Stieler JM, Lohneis P, Bischoff S, Bläker H, Pelzer U, Bahra M. SPARC expression in resected pancreatic cancer patients treated with gemcitabine: results from the CONKO-001 study. Ann Oncol. 2014;25:1025-1032. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 65] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 40. | Zhang J, Wang P, Zhu J, Wang W, Yin J, Zhang C, Chen Z, Sun L, Wan Y, Wang X. SPARC expression is negatively correlated with clinicopathological factors of gastric cancer and inhibits malignancy of gastric cancer cells. Oncol Rep. 2014;31:2312-2320. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 23] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 41. | Peixoto E, Atorrasagasti C, Malvicini M, Fiore E, Rodriguez M, Garcia M, Finocchieto P, Poderoso JJ, Corrales F, Mazzolini G. SPARC gene deletion protects against toxic liver injury and is associated to an enhanced proliferative capacity and reduced oxidative stress response. Oncotarget. 2016; Epub ahead of print. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 3] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 42. | Rossi MK, Gnanamony M, Gondi CS. The ‘SPARC’ of life: Analysis of the role of osteonectin/SPARC in pancreatic cancer (Review). Int J Oncol. 2016;48:1765-1771. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 20] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 43. | Morrissey MA, Jayadev R, Miley GR, Blebea CA, Chi Q, Ihara S, Sherwood DR. SPARC Promotes Cell Invasion In Vivo by Decreasing Type IV Collagen Levels in the Basement Membrane. PLoS Genet. 2016;12:e1005905. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 54] [Cited by in RCA: 52] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 44. | Alachkar H, Santhanam R, Maharry K, Metzeler KH, Huang X, Kohlschmidt J, Mendler JH, Benito JM, Hickey C, Neviani P. SPARC promotes leukemic cell growth and predicts acute myeloid leukemia outcome. J Clin Invest. 2014;124:1512-1524. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 48] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 45. | Nian Q, Xiao Q, Wang L, Luo J, Chen LP, Yang ZS, Liu L. SPARC silencing inhibits the growth of acute myeloid leukemia transformed from myelodysplastic syndrome via induction of cell cycle arrest and apoptosis. Int J Mol Med. 2014;33:856-862. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 9] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 46. | López-Murcia FJ, Terni B, Llobet A. SPARC triggers a cell-autonomous program of synapse elimination. Proc Natl Acad Sci U S A. 2015;112:13366-13371. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 37] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 47. | Vaz J, Ansari D, Sasor A, Andersson R. SPARC: A Potential Prognostic and Therapeutic Target in Pancreatic Cancer. Pancreas. 2015;44:1024-1035. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 87] [Cited by in RCA: 77] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 48. | Hartley PS, Motamedchaboki K, Bodmer R, Ocorr K. SPARC-Dependent Cardiomyopathy in Drosophila. Circ Cardiovasc Genet. 2016;9:119-129. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 25] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 49. | Rayne S, Forest K. Use of the SPARC software program to calculate hydrolysis rate constants for the polymeric brominated flame retardants BC-58 and FR-1025. J Environ Sci Health A Tox Hazard Subst Environ Eng. 2016;51:509-513. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 50. | Alkhateeb A, Marzouka NA, Tashtoush R. Variants in PTPN22 and SMOC2 genes and the risk of thyroid disease in the Jordanian Arab population. Endocrine. 2013;44:702-709. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 16] [Article Influence: 1.3] [Reference Citation Analysis (0)] |