Published online Nov 14, 2016. doi: 10.3748/wjg.v22.i42.9288
Peer-review started: June 30, 2016
First decision: August 8, 2016
Revised: August 23, 2016
Accepted: September 14, 2016
Article in press: September 14, 2016
Published online: November 14, 2016
Processing time: 136 Days and 11.4 Hours
Aberrations in protein glycosylation and polysaccharides play a pivotal role in pancreatic tumorigenesis, influencing cancer progression, metastasis, immuno-response and chemoresistance. Abnormal expression in sugar moieties can impact the function of various glycoproteins, including mucins, surface receptors, adhesive proteins, proteoglycans, as well as their effectors and binding ligands, resulting in an increase in pancreatic cancer invasiveness and a cancer-favored microenvironment. Recent advance in glycoproteomics, glycomics and other chemical biology techniques have been employed to better understand the complex mechanism of glycosylation events and how they orchestrate molecular activities in genomics, proteomics and metabolomics implicated in pancreatic adenocarcinoma. A variety of strategies have been demonstrated targeting protein glycosylation and polysaccharides for diagnostic and therapeutic development.
Core tip: Protein glycosylation plays an important role in pancreatic tumorigenesis. Malignance induced changes in protein glycosylation can profoundly impact the function of a protein in multiple ways. One approach for developing better diagnostic and therapeutic strategies in pancreatic cancer involves targeting cancer-associated aberrant glycosylation. This review discusses the recent discoveries in glycoproteomics study of pancreatic cancer.
- Citation: Pan S, Brentnall TA, Chen R. Glycoproteins and glycoproteomics in pancreatic cancer. World J Gastroenterol 2016; 22(42): 9288-9299
- URL: https://www.wjgnet.com/1007-9327/full/v22/i42/9288.htm
- DOI: https://dx.doi.org/10.3748/wjg.v22.i42.9288
Pancreatic cancer is one of the most deadly cancers, in part because detection of pancreatic cancer is difficult at its early stages when surgical and other treatments are most effective[1,2]. In addition, innate or adapted drug-resistance has been a major hurdle in pancreatic cancer chemotherapy[3,4]. Malignance induced changes in protein glycosylation, such N-glycosylation and O-glycosylation, can profoundly impact the function of a protein in multiple ways, including protein maturation, expression, localization, as well as post-translational modifications, influencing a wide spectrum of glycoproteins and their binding ligands. One approach for developing better diagnostic and therapeutic strategies in pancreatic cancer involves targeting cancer-associated aberrant glycosylation. Recent developments of technology in proteomics and chemical biology have thus stimulated growing interest in elucidating the complex glycosylation events involved in pancreatic adenocarcinoma.
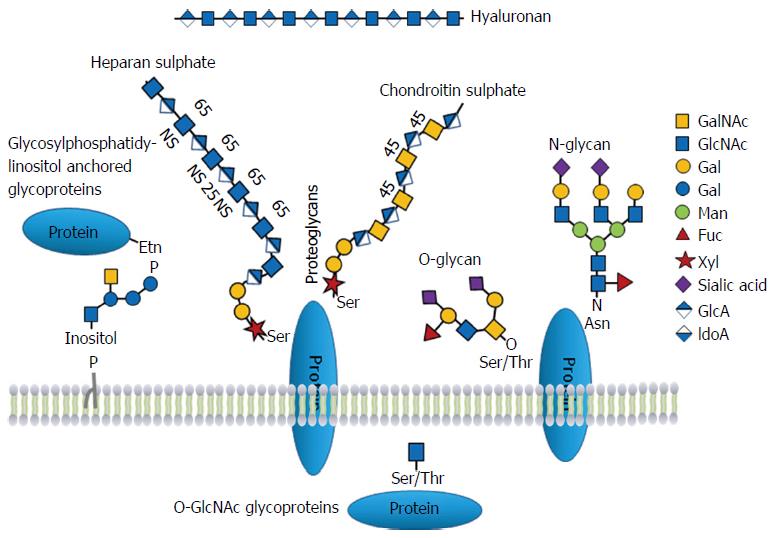
Glycosylation is one of the most complex and common forms of protein post-translational modifications[5,6]. It plays a pivotal role in many biological processes, such as protein folding, cell adhesion and trafficking, cell signaling, pathogen recognition and immune response[7-11]. Protein glycosylation occurs in the endoplasmic reticulum and Golgi apparatus in multiple enzymatic steps. As illustrated in Figure 1, the most common protein glycosylations are N-linked and O-linked glycosylation. N-linked glycans are attached to the amide group of asparagine residues in a consensus Asn-X-Ser/Thr sequence (X can be any amino acid except proline)[12]. O-linked glycans are linked to the hydroxyl group on serine or threonine residues[13]. One unique subclass of O-glycosylation is the phosphorylation-like, reversible O-GlcNAcylation[14]. Less common forms of glycosylation include glycosylphosphatidylinositol anchors attached to protein carboxyl terminus, C-glycosylation that occurs on tryptophan residues[15] and S-linked glycosylation through a sulfur atom on cysteine or methionine[16]. In addition to protein glycosylation, proteoglycans and hyaluronan are major components of the extracellular matrix (ECM), which are implicated in cell proliferation and migration.
Most secretory and membrane-bound proteins produced by mammalian cells contain covalently linked sugar chains with diverse structures. The glycosylation form and density of glycans on a protein can be altered significantly in association with changes in cellular pathways and processes resulted from diseases, such as malignancy. In fact, altered glycosylation patterns have long been recognized as hallmarks in epithelial cancer[17-22], including pancreatic ductal adenocarcinoma (PDAC), which accounts for about 90% of pancreatic cancer. Glycosylation abnormalities can be characterized by one or both of the following changes: (1) composition and structural alterations of glycan; and (2) change in the density of glycosylation at protein sites (hyper, hypo or neo-glycosylation). Ultimately, malignant transformation is usually associated with one or both of these types of glycosylation alterations, leading to the expressional and functional changes of tumor-specific glycoproteins. Malignancy associated glycosylation abnormalities can influence cancer cell proliferation, invasion and viability, as well as interactions with tumor micro environment. Disruption or inhibition of glycosylation and carbohydrate-dependent cellular pathways may represent potential modalities for cancer therapies[23,24]. Receptor tyrosine kinases (RTKs), which are transmembrane glycoproteins that play important roles in malignancy and drug resistance, have been targets of anti-cancer drug development for various malignancies, including pancreatic adenocarcinoma[25,26]. In addition, many of the current blood-based tumor markers are glycoproteins, including CA 19-9 for pancreatic cancer, CA 125 for ovarian cancer, CA 15-3 for breast cancer, and CA 242 for gastrointestinal cancer. CA 19-9, which detects the epitope of sialyl Lewis (a) on mucins and other adhesive molecules such as carcinoembryonic antigen[27-29], is widely used for monitoring the clinical course of pancreatic cancer patients[30]. To date, while implication of aberrant protein glycosylation in malignancy has been well recognized[21,31], limited information is available describing the site specific glycoproteome changes associated with pancreatic cancer.
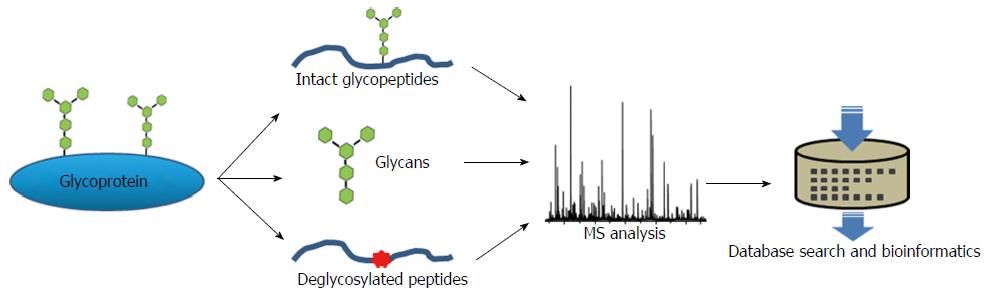
A number of proteomics studies in pancreatic cancer have been reported[32-39]. As a subfield of proteomics, glycoproteomics uniquely focuses on analyzing glycosylated proteins to reveal glycoproteome alterations associated with pancreatic cancer. The major challenge for a comprehensive glycoproteomics analysis in a clinical sample arises from the biological intricacy within the molecule of a glycoprotein, including the variety in glycan composition and structure, as well as the complex linkage to the corresponding protein. Mass spectrometry has been the most effective and versatile instrument platform for both glycan and protein analysis. Although various sample preparation strategies may be applied to collect glycoproteins or glycans from different biological specimens, a glycoproteomics pipeline typically consists of glyco-enrichment, MS analysis and bioinformatics interpretation. The technical details of global analysis of glycoprotein can be found in a number of reviews on the subject of glycomics (analysis of glycans)[40-43] and glycoproteomics (analysis of glycoproteins and glycosites)[15,44-52]. Figure 2 illustrates the overall approaches for MS analysis of glycoproteins. Prior to MS acquisition, glyco-enrichment strategies, including lectin affinity[53-56], hydrazide chemistry[57-60], boronic acid[61], size-exclusion chromatography[62], and hydrophilic interaction[63], may be applied to enrich glycoproteins from complex biological samples, and thus, enhance analytical sensitivity. The direct analysis of intact glycopeptides with carbohydrate attachments is analytically challenging, but allows complementary identification of the peptide backbone and the glycan structure in a single measurement, providing site-specific glycosylation characterization directly. However, this approach is complicated by the mixed information obtained from the MS signals from the peptide backbone, the carbohydrate group and the combinations of both, and therefore, is largely limited for analyzing purified glycoproteins or simple systems. Alternatively, glycans, especially N-linked glycans, can be enzymatically or chemically cleaved from proteins or peptides and analyzed separately by MS. Using glycan databases and bioinformatics tools, MS analysis enables global identification of glycan species in a complex biological sample. On the other hand, de-glycosylated glycopeptides can also be profiled in a global fashion using shotgun proteomics approach to identify the amino acid sequence of the backbone peptides. The N-glycosylation sites can be precisely mapped using the consensus sequence of Asn-X-Ser/Thr, in which asparagine is converted to aspartic acid after PNGase F enzymatic cleavage, which introduces a mass difference of 0.9840 Dalton for MS identification. By defining the glycan structures and profiling the glycoproteins in complex clinical samples, disease associated aberrant glycan forms and site-specific occupancy on proteins can be revealed. For quantitative analysis, additional steps, such as differential stable isotope labeling of the sample and controls, may be required. Ultimately, to comprehensively address disease associated aberrant glycosylation, all the data obtained from different aspects of the workflow need to be integrated, so that the full extent of glycosylation changes with site-specific information can be better revealed.
Identification and detection of abnormal protein glycosylation associated with pancreatic cancer in bodily fluids may present meaningful targets for cancer detection. A variety of carbohydrates and glycoproteins have been investigated for pancreatic cancer detection. Currently, CA19-9 is the only clinical biomarker test for management of pancreatic cancer[64]. While CA19-9 is widely used for monitoring the clinical course of cancer patients, it does not provide adequate accuracy for pancreatic cancer diagnosis and early detection, underscoring the importance of obtaining molecular details on specific glycosylation events involved in neoplastic progression. Mucin (MUC) proteins, including MUC1, MUC5AC, and MUC16 are major protein carriers of CA 19-9, and play important roles in pancreatic cancer tumorigenesis, invasiveness and metastasis, in part through their characteristic glycoforms[28,65-67]. Changes of MUC1 and MUC5AC in pancreatic cancer serum involved distinct glycan alterations, including Thomsen-Friedenreich antigen and fucose and Lewis antigens[28]. The measurement of CA 19-9 antigen on MUC1, MUC5AC and MUC16 individually did not improve the performance of cancer detection, owing to the biological heterogeneity of the patients in their CA 19-9 protein carriers[68]. However, the combined measurement of standard CA 19-9 assay and the detection of the CA 19-9 antigen on MUC5AC and MUC16 did improve the performance of pancreatic cancer detection[68].
In addition to CA19-9, other aberrant protein glycosylations associated with pancreatic cancer have also been investigated in bodily fluids. The glycosylation of serum ribonuclease 1 (RNASE1) - another well-studied pancreas associated protein, showed a 40% increase in core fucosylation in pancreatic cancer[69]. Using Concanavalin A lectin affinity chromatography for N-glycopeptide enrichment and LC MS/MS, one study identified 92 individual glycosylation sites and 105 unique carbohydrate structures in serum, and observed increased branching of N-linked oligosaccharides, as well as increased protein fucosylation and sialylation in the sera from pancreatic cancer patients[70]. Increased level of sialyl Lewis X of major serum acute-phase proteins, including alpha-1-acid glycoprotein (AGP1 or ORM1), haptoglobin (HP), fetuin (AHSG), alpha-1-antitrypsin (SERPINA1) and transferrin (TF) were observed in the sera from patients with advanced pancreatic cancer and chronic pancreatitis - an alteration possibly associated with inflammatory response[71]. In addition, the observation of an increase in core fucosylation on AGP1 and HP in the serum of advanced pancreatic cancer may represent a potential cancer associated signal[71]. Although the detection of increase level of fucosylated HP alone does not provide sufficient accuracy for pancreatic cancer diagnosis, it is possible that fucosylated HP might be used as an indication of liver metastasis if the biomarker undergoes further validation[72,73]. The changes in protein fucosylation and sialylation in pancreatic cancer were also investigated by analyzing intact glycopeptides. Using immunoprecipitation, partial deglycosylation and LC MS/MS, one study suggested that the core-fucosylation levels at site N396 and N1424 in alpha-2-macroglobulin (A2M) were decreased in serum of both pancreatic cancer and chronic pancreatitis compared to non-diseased controls[74]. The investigation of sialylated N-glycopeptide levels in sera from pancreatic cancer patients in comparison to non-diseased controls and acute pancreatitis patients identified 13 glycoforms, mainly from high-abundant serum proteins, with changes associated with pancreatic cancer group[75]. Mucinous cystic neoplasms (MCN) and intraductal papillary mucinous neoplasms (IPMN) are pancreatic cysts that are subject to high risk of malignant transformation. Proteomic and glycomic investigation of cyst fluids collected from patients with MCN and IPMN led to the identification of 80 N-linked glycans, and several hyper-fucosylated glycoproteins, including triacylglycerol lipase and pancreatic α-amylase[76].
Known tumor-specific glycoproteins, such as mucins and carcinoembryonic antigen-related cell adhesion molecules, have been extensively studied for their roles in neoplastic progression and metastasis of pancreatic cancer[77-81]. The emerging technology of glycoproteomics has been recently applied to interrogate broader changes of glycoproteome in pancreatic cancer cells and tissue. A large number of cell surface proteins are transmembrane glycoproteins, including many of cell-surface receptors such as RTKs, which play pivotal roles in signaling, trafficking and cell-cell interactions. These cell-surface receptors, such as epithelial growth factor receptor (EGFR), integrins, and TGF β receptor (TGFβR) have been important targets for anti-cancer therapy, and their glycosylation forms impact their functionality[82-85]. Using a biocytin hydrazide cell surface capturing technique[86], or azido sugar based bioorthogonal chemical reporter for metabolic glycan labeling[87] for glycopeptide enrichment, studies were carried out to profile N-linked glycopeptides derived from surface glycoproteins of pancreatic cancer cells using LC MS/MS[88,89]. The studies indicated the overexpression of CD109[88] and ecto-50-nucleotidase[89] in pancreatic cancer cells and tissues. Using multi-lectin affinity chromatography and LC-MS/MS, another study investigated the differential glycoproteins associated with pancreatic cancer CD24+CD44+ stem-like cells in comparison with CD24-CD44+ cells[90]. The study indicated that the high expression and high positive rate of CD24 was significantly associated with late-stage pancreatic adenocarcinomas, while CD13 expression and positive rate were negatively associated with tumor progression. By manipulating exogenous substrate supply, a study reported that increases in metabolic flux through the sialic acid pathway could dramatically enhance the sialylation of certain N-linked glycoproteins to influence cancer cell adhesive and mobility properties of SW1990 pancreatic cancer cells[91].
Glycoproteomic techniques were also applied to investigate the glycoproteome of pancreatic cancer tissues. In our study, we observed an overall increase in N-glycosylation level on many glycoproteins in PDAC tissue in comparison with normal pancreas[92]. Supplemental Table 1 summarizes some of the glycoproteins with at least one N-glycopeptide overexpressed (≥ 2 fold) in pancreatic cancer, including many pancreatic cancer associated proteins, such as MUC5AC, carcinoembryonic antigen-related cell adhesion molecule 5, insulin-like growth factor binding protein (IGFBP3), cathepsin D (CTSD), as well as a number of CD antigens (including CD44 - a marker of pancreatic cancer stem-like cells) and integrins. Pathway analysis suggested that increased N-glycosylation activities of these proteins were implicated in several pancreatic cancer pathways, including TGF-β, TNF, and NF-kappa-B[92]. Other glycoproteins, such as Thy-1 membrane glycoprotein (THY1), which was recently developed into an ultrasound molecular imaging marker for pancreatic cancer detection[93], was found heavily N-glycosylated in pancreatic cancer tissues. Further mapping of N-glycosylation sites revealed that the change of N-glycosylation level in pancreatic cancer was not only protein specific, but also glycosylation site specific. Specific N-glycosylation sites within certain individual proteins can have significantly altered glycosylation occupancy (e.g., a change in glycan density) in pancreatic cancer, reflecting the complex nature of glycosylation events underlying pancreatic tumorigenesis. Notably, the increase of N-glycosylation of many of these proteins was also found in chronic pancreatitis tissue, supporting the notion that pancreatic cancer and chronic pancreatitis share many common clinical and molecular features[94-98]. It is also noteworthy to mention that in contrast to many glycoproteins with increased N-glycosylation, pancreatic secretory granule membrane major glycoprotein (GP2) - a pancreas specific glycoprotein, showed a reduced N-glycosylated level in both cancer and chronic pancreatitis tissues. While the global data have revealed the aberrant N-glycosylation changes of many relevant proteins in pancreatic cancer tissues, the orchestrated glycosylation mechanism underlying pancreatic tumorigenesis, immune response and pancreatic functional changes, remains poorly understood and warrant further investigation.
Mucins are high molecular weight glycoproteins produced by various epithelial cells, and have 21 family members. The mucins are heavily glycosylated in O- and N-linked glycosylation and implicated in PDAC through their characteristic glycoforms influencing tumorigenicity, invasiveness, metastasis and drug resistance. Mucins have been extensively studied in PDAC, and showed various expressional and glycosylation changes not only in pancreatic carcinoma, but also in pancreatic intraepithelial neoplasia (PanIN), IPMN and MCN[66,67,99]. Several mucins, including MUC1, MUC4, MUC5AC and MUC16, are frequently upregulated in PDAC. Mucin core protein expression and the differential localization in PDAC and its precursor lesions have been well documented in the literature[66,67,100-103]. In addition, mucin glycoforms also play an important role in modulating their functionality in tumorigenesis as well as cancer cell interaction with the tumor microenvironment. In fact, the glycan component can make up more than 50% of the molecular weight of a mucin glycoprotein.
The glycosylation of cancer associated mucins is largely associated with Tn antigen, sialyl Tn and fucosylated core 1 structures, forming the so-called tumor-associated antigens[104]. Altered glycoforms of MUC1, MUC4 and MUC5AC were observed early in pancreatic cancer progression (PanINs) to late stage metastatic disease[105]. The elevation of fucosylated core structures, fucose and Lewis antigen have frequently been detected on MUC1 and MUC5AC in the blood from patients with pancreatic cancer[28]. Additionally, MUC16 and its sialofucosylated structures were reported overexpressed in pancreatic cancer cell and acted as a functional ligand for E- and L-selectin to enhance cancer cell metastatic spread[106]. By stimulating pancreatic cancer cells with pro-inflammatory conditions, such as oxidative stress and cytokines, mucin glycosylation can be significantly altered in specific pancreatic cancer cell lines, suggesting a possible molecular link between inflammation, glycosylation alteration and adaptive responses of those pancreatic cancer cells[107]. Efforts have also been made to use proteomic approaches to prolife mucins in cyst fluids to enhance the discrimination of malignant pancreatic cyst lesions from those that are benign[108].
In our proteomic study, we observed a large group of ECM associated proteins overexpressed in pancreatic cancer and chronic pancreatitis tissues[97]. Many of these proteins are glycoproteins and are involved in stellate cell activation and ECM organizational and structural changes, which regulate pancreatic fibrosis - one of the fundamental histological abnormalities observed in pancreatic adenocarcinoma and chronic pancreatitis. ECM components, including matrix proteins, proteoglycan proteins, galectins and hyaluronan, which interact with each other and form supramolecular complexes, are subjected to alterations during cancer progression, leading to cancer associated ECM[109-115]. Abnormal protein glycosylation can significantly affect the mechanical properties of ECM, enhancing tumor cell migration[110,116]. Studies have shown that ECM components associated with integrin-ECM axis are highly up-regulated in pancreatic cancer[117]. The glycoforms of integrins, such as the presence of N-linked oligosaccharides, can regulate integrin function, affecting the cell-ECM interactions. In pancreatic cancer tissues, we observed increased levels of N-glycosylation, not only on several integrins (both α and β subunits), but also on ECM adhesion proteins, including collagens, fibronectin, vitronectin, and laminin (Supplemental Table 1)[92]. These observations warrant mechanistic study to better understand how aberrant glycosylation of integrins and ECM adhesion ligands influence pancreatic cancer migration and malignant phenotypes. Galectins and fibulins play a role in organization of ECM supramolecular structure, such as basement membranes, by forming intramolecular bridges, binding to complex carbohydrates and ECM adhesive proteins[118-120]. The core protein expression and N-glycosylation level of fibulin 1 were both found up-regulated in pancreatic cancer tissues[92,97]. Galectin 1 (LGALS1) is a human extracellular lectin that specifically binds to β-galactoside sugars, including N- and O-linked glycans. Galectin 1 was overexpressed in the stroma of both pancreatic cancer and PanINs lesions[121], and its expression was related to pancreatic cancer survival[122,123]. Concurrently, the N-glycosylation level of endogenous ligands of galectin-1 in the ECM, including fibronectin (FN1), laminins and galectin-3-binding protein (LGALS3BP), were all up-regulated in pancreatic cancer tissue (Supplemental Table 1), implying an intensified interaction of galectins and their major binding partners in pancreatic cancer[92]. Periostin (POSTN), an ECM protein involved in cell mobility and neovascularization[124], has both up-regulated core protein expression and N-glycosylation levels in pancreatic cancer tissue[92,97]. Cathepsins are proteases that are implicated in cancer invasion by degrading ECM, including proteoglycans and collagens. We observed up-regulation of both core protein expression and N-glycosylation level of cathepsins (CTSD, CTSL) (Supplemental Table 1) in pancreatic cancer tissue, suggesting its possible functional role in pancreatic tumorigenesis[92,94].
Proteoglycans are heavily glycosylated proteins with serine attached glycosaminoglycans (GAGs), such as heparan sulphate and chondroitin sulphate (Figure 1). Proteoglycans are an important component of ECM and affect multiple biological processes, including cell differentiation and proliferation, binding to cytokines, growth factors and morphogens. During tumorigenesis, the expression and glycosylation patterns of proteoglycans change in the stroma surrounding cancer, influencing tumor growth and neoplastic progression[125]. In proteomics and other studies, the increased expression of proteoglycan proteins, including lumican, decorin, versican, and biglycan, has been observed in pancreatic cancer tissues or cells[97,121,126-130]. Since GAGs are large, linear polysaccharides, to a certain extent, the biological function of proteoglycans can be governed by the interaction of the attached GAGs with other proteins. Most of proteoglycans also contain N- and O-linked glycans. In a quantitative glycoproteomics study, N-glycosylation levels of several major ECM proteoglycans, including decorin (DCN), biglycan (BGN), lumican (LUM), versican (VCAN), and aggrecan (ACAN), were found elevated in pancreatic cancer tissues (Supplemental Table 1)[92].
Hyaluronan is a non sulfated glycosaminoglycan that is not covalently attached to proteoglycans and can have a very high molecule weight[131]. CD44 and receptor for HA-mediated motility are the two main receptors for the anchorage of hyaluronan-rich ECM to the cell surface[132,133]. Although it has relatively simple chemical composition, as one of the major components of ECM, hyaluronan is involved in promoting pancreatic cancer progression and chemoresistance[132-134]. Aberrant production and deposition of hyaluronan provide a favorable microenvironment to enhance cancer cell proliferation, migration, invasion, angiogenesis, and limit the delivery of anti-cancer agents[134-137]. Studies have also shown that the interaction between hyaluronan and its CD44 receptor is involved in the stemness and survival of cancer stem cells[138], and may be relevant to pancreatic cancer CD24+CD44+ stem-like cells.
Protein glycosylation has become a prominent target for drug development. One strategy involves disruption of the protein glycosylation process, such as inhibition of glycosylation enzymes and hexosamine biosynthetic pathway, to reduce pancreatic cancer progression and tumor growth[23,24]. Silencing O-GlcNAc transferase has shown to inhibit pancreatic cancer growth[139]. Inhibition of N-glycosylation can influence the maturation and surface expression of RTKs (e.g., EGFR, IGF1R), and enhance chemosensitivity of drug-resistant pancreatic cancer cells[82]. Lewis-Y carbohydrate antigen is expressed by many epithelial cancers, including pancreatic cancer[24,140], and has been a target for cancer vaccines and immunoconjugated chemotherapy[141,142].
Mucins (especially MUC1, MUC4, MUC5AC, MUC16) are an important group of glycoproteins in pancreatic cancer and have been targeted for therapeutic treatment. Multiple efforts have been made to develop MUC peptide based vaccination for pancreatic cancer, unfortunately with no significant clinical effects[99]. New data suggests that it may be important to incorporate cancer associated glycoforms in the vaccine design so that the specific immunogenic epitopes expressed in tumors can be better mimicked[143-145]. Other mucin based targeted therapeutic approaches include radioimmunoconjugate of MUC1 antibodies and gene therapy designed to suppress MUC1 or mucin gene promoters[99].
The ECM is consisted of various biopolymers, and plays a pivotal role in cancer invasion and metastasis. Several anti-cancer therapeutic strategies targeting different ECM components have been considered, including inhibition of heparanase and proteases, anti-integrin therapy, and inhibition of GAGs of proteoglycans[146]. In addition, anti-cancer therapies targeting hyaluronan metabolic enzymes and hyaluronan-CD44 interactions have also been investigated[147,148]. In pancreatic cancer, recent studies demonstrated that depletion of stromal hyaluronan surrounding tumor improved drug delivery and significantly enhanced the efficacy of gemcitabine treatment[134,137,149].
Protein glycosylation is deeply involved in pancreatic tumorigenesis. Cancer associated changes in protein glycosylation and polysaccharides can profoundly affect cellular function and ECM organization, supporting tumor growth and metastasis, as well as influencing immuno-response and chemoresistance. The emerging technologies of glycoproteomics, glycomics and other chemical biology approaches provide powerful tools to interrogate the complex nature of protein glycosylation involved in pancreatic cancer. While significant efforts have been made, ranging from mechanistic investigation, to biomarker discovery and therapeutic development, many aspects of how glycosylation events orchestrate changes in cancer signaling pathways at the genomic, proteomic and metabolomic level to facilitate cancer progression remain to be elucidated. To analyze complex clinical samples and obtain an in-depth, comprehensive understanding of site specific glycosylation changes requires a concerted approach drawing from a variety of techniques. With the development of molecular techniques and bioinformatics, many of the current technical obstacles may be transient. Nonetheless, many strategies have been demonstrated to target protein glycosylation and polysaccharides for diagnostic and therapeutic gains in pancreatic cancer. These studies have laid foundation and will provide experimental guidance for future investigations.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: United States
Peer-review report classification
Grade A (Excellent): A
Grade B (Very good): 0
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Kucherlapati MH S- Editor: Qi Y L- Editor: A E- Editor: Zhang FF
| 1. | Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA Cancer J Clin. 2013;63:11-30. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8406] [Cited by in RCA: 8971] [Article Influence: 690.1] [Reference Citation Analysis (0)] |
| 2. | Vincent A, Herman J, Schulick R, Hruban RH, Goggins M. Pancreatic cancer. Lancet. 2011;378:607-620. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2129] [Cited by in RCA: 2115] [Article Influence: 151.1] [Reference Citation Analysis (3)] |
| 3. | Páez D, Labonte MJ, Lenz HJ. Pancreatic cancer: medical management (novel chemotherapeutics). Gastroenterol Clin North Am. 2012;41:189-209. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 27] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 4. | Siegel R, DeSantis C, Virgo K, Stein K, Mariotto A, Smith T, Cooper D, Gansler T, Lerro C, Fedewa S. Cancer treatment and survivorship statistics, 2012. CA Cancer J Clin. 2012;62:220-241. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1973] [Cited by in RCA: 2080] [Article Influence: 160.0] [Reference Citation Analysis (2)] |
| 5. | Bertozzi CR, Kiessling LL. Chemical glycobiology. Science. 2001;291:2357-2364. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1561] [Cited by in RCA: 1559] [Article Influence: 65.0] [Reference Citation Analysis (0)] |
| 6. | Khoury GA, Baliban RC, Floudas CA. Proteome-wide post-translational modification statistics: frequency analysis and curation of the swiss-prot database. Sci Rep. 2011;1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 556] [Cited by in RCA: 652] [Article Influence: 46.6] [Reference Citation Analysis (0)] |
| 7. | Haltiwanger RS, Lowe JB. Role of glycosylation in development. Annu Rev Biochem. 2004;73:491-537. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 596] [Cited by in RCA: 586] [Article Influence: 27.9] [Reference Citation Analysis (0)] |
| 8. | Helenius A, Aebi M. Intracellular functions of N-linked glycans. Science. 2001;291:2364-2369. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1802] [Cited by in RCA: 1789] [Article Influence: 74.5] [Reference Citation Analysis (0)] |
| 9. | Ohtsubo K, Marth JD. Glycosylation in cellular mechanisms of health and disease. Cell. 2006;126:855-867. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1977] [Cited by in RCA: 2210] [Article Influence: 116.3] [Reference Citation Analysis (0)] |
| 10. | Rudd PM, Elliott T, Cresswell P, Wilson IA, Dwek RA. Glycosylation and the immune system. Science. 2001;291:2370-2376. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1223] [Cited by in RCA: 1212] [Article Influence: 50.5] [Reference Citation Analysis (0)] |
| 11. | Woods RJ, Edge CJ, Dwek RA. Protein surface oligosaccharides and protein function. Nat Struct Biol. 1994;1:499-501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 72] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 12. | Bause E. Structural requirements of N-glycosylation of proteins. Studies with proline peptides as conformational probes. Biochem J. 1983;209:331-336. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 462] [Cited by in RCA: 493] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 13. | Jensen PH, Kolarich D, Packer NH. Mucin-type O-glycosylation--putting the pieces together. FEBS J. 2010;277:81-94. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 190] [Cited by in RCA: 189] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 14. | Hart GW, Housley MP, Slawson C. Cycling of O-linked beta-N-acetylglucosamine on nucleocytoplasmic proteins. Nature. 2007;446:1017-1022. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1042] [Cited by in RCA: 1146] [Article Influence: 63.7] [Reference Citation Analysis (0)] |
| 15. | Wei X, Li L. Comparative glycoproteomics: approaches and applications. Brief Funct Genomic Proteomic. 2009;8:104-113. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 42] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 16. | Floyd N, Vijayakrishnan B, Koeppe JR, Davis BG. Thiyl glycosylation of olefinic proteins: S-linked glycoconjugate synthesis. Angew Chem Int Ed Engl. 2009;48:7798-7802. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 175] [Cited by in RCA: 185] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 17. | Dennis JW, Granovsky M, Warren CE. Glycoprotein glycosylation and cancer progression. Biochim Biophys Acta. 1999;1473:21-34. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 517] [Cited by in RCA: 505] [Article Influence: 19.4] [Reference Citation Analysis (0)] |
| 18. | Kobata A. Altered glycosylation of surface glycoproteins in tumor cells and its clinical application. Pigment Cell Res. 1989;2:304-308. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 20] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 19. | Kobata A, Amano J. Altered glycosylation of proteins produced by malignant cells, and application for the diagnosis and immunotherapy of tumours. Immunol Cell Biol. 2005;83:429-439. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 193] [Cited by in RCA: 189] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 20. | Ono M, Hakomori S. Glycosylation defining cancer cell motility and invasiveness. Glycoconj J. 2004;20:71-78. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 108] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 21. | Pinho SS, Reis CA. Glycosylation in cancer: mechanisms and clinical implications. Nat Rev Cancer. 2015;15:540-555. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1664] [Cited by in RCA: 2189] [Article Influence: 218.9] [Reference Citation Analysis (0)] |
| 22. | Brooks SA, Carter TM, Royle L, Harvey DJ, Fry SA, Kinch C, Dwek RA, Rudd PM. Altered glycosylation of proteins in cancer: what is the potential for new anti-tumour strategies. Anticancer Agents Med Chem. 2008;8:2-21. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 82] [Cited by in RCA: 87] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 23. | Dwek RA, Butters TD, Platt FM, Zitzmann N. Targeting glycosylation as a therapeutic approach. Nat Rev Drug Discov. 2002;1:65-75. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 347] [Cited by in RCA: 340] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 24. | Vasconcelos-Dos-Santos A, Oliveira IA, Lucena MC, Mantuano NR, Whelan SA, Dias WB, Todeschini AR. Biosynthetic Machinery Involved in Aberrant Glycosylation: Promising Targets for Developing of Drugs Against Cancer. Front Oncol. 2015;5:138. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 87] [Cited by in RCA: 111] [Article Influence: 11.1] [Reference Citation Analysis (1)] |
| 25. | Silvestris N, Gnoni A, Brunetti AE, Vincenti L, Santini D, Tonini G, Merchionne F, Maiello E, Lorusso V, Nardulli P. Target therapies in pancreatic carcinoma. Curr Med Chem. 2014;21:948-965. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 37] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 26. | Troiani T, Martinelli E, Capasso A, Morgillo F, Orditura M, De Vita F, Ciardiello F. Targeting EGFR in pancreatic cancer treatment. Curr Drug Targets. 2012;13:802-810. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 106] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 27. | Akagi J, Takai E, Tamori Y, Nakagawa K, Ogawa M. CA19-9 epitope a possible marker for MUC-1/Y protein. Int J Oncol. 2001;18:1085-1091. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 11] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 28. | Yue T, Goldstein IJ, Hollingsworth MA, Kaul K, Brand RE, Haab BB. The prevalence and nature of glycan alterations on specific proteins in pancreatic cancer patients revealed using antibody-lectin sandwich arrays. Mol Cell Proteomics. 2009;8:1697-1707. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 110] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 29. | Yue T, Partyka K, Maupin KA, Hurley M, Andrews P, Kaul K, Moser AJ, Zeh H, Brand RE, Haab BB. Identification of blood-protein carriers of the CA 19-9 antigen and characterization of prevalence in pancreatic diseases. Proteomics. 2011;11:3665-3674. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 57] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 30. | Goonetilleke KS, Siriwardena AK. Systematic review of carbohydrate antigen (CA 19-9) as a biochemical marker in the diagnosis of pancreatic cancer. Eur J Surg Oncol. 2007;33:266-270. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 539] [Cited by in RCA: 611] [Article Influence: 32.2] [Reference Citation Analysis (1)] |
| 31. | Stowell SR, Ju T, Cummings RD. Protein glycosylation in cancer. Annu Rev Pathol. 2015;10:473-510. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 463] [Cited by in RCA: 637] [Article Influence: 63.7] [Reference Citation Analysis (0)] |
| 32. | Cecconi D, Palmieri M, Donadelli M. Proteomics in pancreatic cancer research. Proteomics. 2011;11:816-828. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 43] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 33. | Chen R, Pan S, Brentnall TA, Aebersold R. Proteomic profiling of pancreatic cancer for biomarker discovery. Mol Cell Proteomics. 2005;4:523-533. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 111] [Cited by in RCA: 107] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 34. | Gräntzdörffer I, Carl-McGrath S, Ebert MP, Röcken C. Proteomics of pancreatic cancer. Pancreas. 2008;36:329-336. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 32] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 35. | Omenn GS, Yocum AK, Menon R. Alternative splice variants, a new class of protein cancer biomarker candidates: findings in pancreatic cancer and breast cancer with systems biology implications. Dis Markers. 2010;28:241-251. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 43] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 36. | Pan S, Brentnall TA, Kelly K, Chen R. Tissue proteomics in pancreatic cancer study: discovery, emerging technologies, and challenges. Proteomics. 2013;13:710-721. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 34] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 37. | Pan S, Brentnall TA, Chen R. Proteomics analysis of bodily fluids in pancreatic cancer. Proteomics. 2015;15:2705-2715. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 47] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 38. | Sun C, Rosendahl AH, Ansari D, Andersson R. Proteome-based biomarkers in pancreatic cancer. World J Gastroenterol. 2011;17:4845-4852. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 29] [Cited by in RCA: 27] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 39. | Vimalachandran D, Costello E. Proteomic technologies and their application to pancreatic cancer. Expert Rev Proteomics. 2004;1:493-501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 18] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 40. | North SJ, Hitchen PG, Haslam SM, Dell A. Mass spectrometry in the analysis of N-linked and O-linked glycans. Curr Opin Struct Biol. 2009;19:498-506. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 191] [Cited by in RCA: 177] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 41. | Prescher JA, Bertozzi CR. Chemical technologies for probing glycans. Cell. 2006;126:851-854. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 172] [Cited by in RCA: 180] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 42. | Rudd PM, Guile GR, Küster B, Harvey DJ, Opdenakker G, Dwek RA. Oligosaccharide sequencing technology. Nature. 1997;388:205-207. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 116] [Cited by in RCA: 99] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 43. | Wuhrer M. Glycomics using mass spectrometry. Glycoconj J. 2013;30:11-22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 121] [Cited by in RCA: 118] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 44. | Ahn YH, Kim JY, Yoo JS. Quantitative mass spectrometric analysis of glycoproteins combined with enrichment methods. Mass Spectrom Rev. 2015;34:148-165. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 65] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 45. | Budnik BA, Lee RS, Steen JA. Global methods for protein glycosylation analysis by mass spectrometry. Biochim Biophys Acta. 2006;1764:1870-1880. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 61] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 46. | Lai ZW, Nice EC, Schilling O. Glycocapture-based proteomics for secretome analysis. Proteomics. 2013;13:512-525. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 21] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 47. | Lazar IM, Deng J, Ikenishi F, Lazar AC. Exploring the glycoproteomics landscape with advanced MS technologies. Electrophoresis. 2015;36:225-237. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 23] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 48. | Morelle W, Canis K, Chirat F, Faid V, Michalski JC. The use of mass spectrometry for the proteomic analysis of glycosylation. Proteomics. 2006;6:3993-4015. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 179] [Cited by in RCA: 162] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 49. | Nilsson J, Halim A, Grahn A, Larson G. Targeting the glycoproteome. Glycoconj J. 2013;30:119-136. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 33] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 50. | Pan S, Chen R, Aebersold R, Brentnall TA. Mass spectrometry based glycoproteomics--from a proteomics perspective. Mol Cell Proteomics. 2011;10:R110.003251. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 178] [Cited by in RCA: 204] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 51. | Wuhrer M, Catalina MI, Deelder AM, Hokke CH. Glycoproteomics based on tandem mass spectrometry of glycopeptides. J Chromatogr B Analyt Technol Biomed Life Sci. 2007;849:115-128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 356] [Cited by in RCA: 331] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 52. | Zhang Y, Yin H, Lu H. Recent progress in quantitative glycoproteomics. Glycoconj J. 2012;29:249-258. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 48] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 53. | Geng M, Zhang X, Bina M, Regnier F. Proteomics of glycoproteins based on affinity selection of glycopeptides from tryptic digests. J Chromatogr B Biomed Sci Appl. 2001;752:293-306. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 82] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 54. | Kaji H, Saito H, Yamauchi Y, Shinkawa T, Taoka M, Hirabayashi J, Kasai K, Takahashi N, Isobe T. Lectin affinity capture, isotope-coded tagging and mass spectrometry to identify N-linked glycoproteins. Nat Biotechnol. 2003;21:667-672. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 524] [Cited by in RCA: 503] [Article Influence: 22.9] [Reference Citation Analysis (0)] |
| 55. | Wang Y, Wu SL, Hancock WS. Approaches to the study of N-linked glycoproteins in human plasma using lectin affinity chromatography and nano-HPLC coupled to electrospray linear ion trap--Fourier transform mass spectrometry. Glycobiology. 2006;16:514-523. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 100] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 56. | Yang Z, Hancock WS. Monitoring glycosylation pattern changes of glycoproteins using multi-lectin affinity chromatography. J Chromatogr A. 2005;1070:57-64. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 88] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 57. | Liu T, Qian WJ, Gritsenko MA, Camp DG, Monroe ME, Moore RJ, Smith RD. Human plasma N-glycoproteome analysis by immunoaffinity subtraction, hydrazide chemistry, and mass spectrometry. J Proteome Res. 2005;4:2070-2080. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 349] [Cited by in RCA: 359] [Article Influence: 18.9] [Reference Citation Analysis (0)] |
| 58. | Pan S, Wang Y, Quinn JF, Peskind ER, Waichunas D, Wimberger JT, Jin J, Li JG, Zhu D, Pan C. Identification of glycoproteins in human cerebrospinal fluid with a complementary proteomic approach. J Proteome Res. 2006;5:2769-2779. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 81] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 59. | Zhang H, Li XJ, Martin DB, Aebersold R. Identification and quantification of N-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat Biotechnol. 2003;21:660-666. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1175] [Cited by in RCA: 1158] [Article Influence: 52.6] [Reference Citation Analysis (0)] |
| 60. | Zhang H, Aebersold R. Isolation of glycoproteins and identification of their N-linked glycosylation sites. Methods Mol Biol. 2006;328:177-185. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 15] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 61. | Sparbier K, Koch S, Kessler I, Wenzel T, Kostrzewa M. Selective isolation of glycoproteins and glycopeptides for MALDI-TOF MS detection supported by magnetic particles. J Biomol Tech. 2005;16:407-413. [PubMed] |
| 62. | Alvarez-Manilla G, Warren NL, Abney T, Atwood J, Azadi P, York WS, Pierce M, Orlando R. Tools for glycomics: relative quantitation of glycans by isotopic permethylation using 13CH3I. Glycobiology. 2007;17:677-687. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 158] [Cited by in RCA: 150] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 63. | Hägglund P, Bunkenborg J, Elortza F, Jensen ON, Roepstorff P. A new strategy for identification of N-glycosylated proteins and unambiguous assignment of their glycosylation sites using HILIC enrichment and partial deglycosylation. J Proteome Res. 2004;3:556-566. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 395] [Cited by in RCA: 368] [Article Influence: 17.5] [Reference Citation Analysis (0)] |
| 64. | Szajda SD, Waszkiewicz N, Chojnowska S, Zwierz K. Carbohydrate markers of pancreatic cancer. Biochem Soc Trans. 2011;39:340-343. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 18] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 65. | Hollingsworth MA, Swanson BJ. Mucins in cancer: protection and control of the cell surface. Nat Rev Cancer. 2004;4:45-60. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1288] [Cited by in RCA: 1361] [Article Influence: 64.8] [Reference Citation Analysis (0)] |
| 66. | Kaur S, Kumar S, Momi N, Sasson AR, Batra SK. Mucins in pancreatic cancer and its microenvironment. Nat Rev Gastroenterol Hepatol. 2013;10:607-620. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 172] [Cited by in RCA: 233] [Article Influence: 19.4] [Reference Citation Analysis (0)] |
| 67. | Nagata K, Horinouchi M, Saitou M, Higashi M, Nomoto M, Goto M, Yonezawa S. Mucin expression profile in pancreatic cancer and the precursor lesions. J Hepatobiliary Pancreat Surg. 2007;14:243-254. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 127] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 68. | Yue T, Maupin KA, Fallon B, Li L, Partyka K, Anderson MA, Brenner DE, Kaul K, Zeh H, Moser AJ. Enhanced discrimination of malignant from benign pancreatic disease by measuring the CA 19-9 antigen on specific protein carriers. PLoS One. 2011;6:e29180. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 52] [Cited by in RCA: 56] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 69. | Barrabés S, Pagès-Pons L, Radcliffe CM, Tabarés G, Fort E, Royle L, Harvey DJ, Moenner M, Dwek RA, Rudd PM. Glycosylation of serum ribonuclease 1 indicates a major endothelial origin and reveals an increase in core fucosylation in pancreatic cancer. Glycobiology. 2007;17:388-400. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 82] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 70. | Zhao J, Qiu W, Simeone DM, Lubman DM. N-linked glycosylation profiling of pancreatic cancer serum using capillary liquid phase separation coupled with mass spectrometric analysis. J Proteome Res. 2007;6:1126-1138. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 136] [Cited by in RCA: 136] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 71. | Sarrats A, Saldova R, Pla E, Fort E, Harvey DJ, Struwe WB, de Llorens R, Rudd PM, Peracaula R. Glycosylation of liver acute-phase proteins in pancreatic cancer and chronic pancreatitis. Proteomics Clin Appl. 2010;4:432-448. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 112] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 72. | Miyoshi E, Nakano M. Fucosylated haptoglobin is a novel marker for pancreatic cancer: detailed analyses of oligosaccharide structures. Proteomics. 2008;8:3257-3262. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 93] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 73. | Miyoshi E, Shinzaki S, Moriwaki K, Matsumoto H. Identification of fucosylated haptoglobin as a novel tumor marker for pancreatic cancer and its possible application for a clinical diagnostic test. Methods Enzymol. 2010;478:153-164. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 23] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 74. | Lin Z, Yin H, Lo A, Ruffin MT, Anderson MA, Simeone DM, Lubman DM. Label-free relative quantification of alpha-2-macroglobulin site-specific core-fucosylation in pancreatic cancer by LC-MS/MS. Electrophoresis. 2014;35:2108-2115. [PubMed] |
| 75. | Kontro H, Joenväärä S, Haglund C, Renkonen R. Comparison of sialylated N-glycopeptide levels in serum of pancreatic cancer patients, acute pancreatitis patients, and healthy controls. Proteomics. 2014;14:1713-1723. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 36] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 76. | Mann BF, Goetz JA, House MG, Schmidt CM, Novotny MV. Glycomic and proteomic profiling of pancreatic cyst fluids identifies hyperfucosylated lactosamines on the N-linked glycans of overexpressed glycoproteins. Mol Cell Proteomics. 2012;11:M111.015792. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 38] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 77. | Chaturvedi P, Singh AP, Chakraborty S, Chauhan SC, Bafna S, Meza JL, Singh PK, Hollingsworth MA, Mehta PP, Batra SK. MUC4 mucin interacts with and stabilizes the HER2 oncoprotein in human pancreatic cancer cells. Cancer Res. 2008;68:2065-2070. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 138] [Cited by in RCA: 139] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 78. | Remmers N, Bailey JM, Mohr AM, Hollingsworth MA. Molecular pathology of early pancreatic cancer. Cancer Biomark. 2010;9:421-440. [PubMed] |
| 79. | Simeone DM, Ji B, Banerjee M, Arumugam T, Li D, Anderson MA, Bamberger AM, Greenson J, Brand RE, Ramachandran V. CEACAM1, a novel serum biomarker for pancreatic cancer. Pancreas. 2007;34:436-443. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 108] [Cited by in RCA: 121] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 80. | Singh AP, Chaturvedi P, Batra SK. Emerging roles of MUC4 in cancer: a novel target for diagnosis and therapy. Cancer Res. 2007;67:433-436. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 110] [Cited by in RCA: 112] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 81. | Swanson BJ, McDermott KM, Singh PK, Eggers JP, Crocker PR, Hollingsworth MA. MUC1 is a counter-receptor for myelin-associated glycoprotein (Siglec-4a) and their interaction contributes to adhesion in pancreatic cancer perineural invasion. Cancer Res. 2007;67:10222-10229. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 88] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 82. | Contessa JN, Bhojani MS, Freeze HH, Rehemtulla A, Lawrence TS. Inhibition of N-linked glycosylation disrupts receptor tyrosine kinase signaling in tumor cells. Cancer Res. 2008;68:3803-3809. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 136] [Cited by in RCA: 168] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 83. | Purchio AF, Cooper JA, Brunner AM, Lioubin MN, Gentry LE, Kovacina KS, Roth RA, Marquardt H. Identification of mannose 6-phosphate in two asparagine-linked sugar chains of recombinant transforming growth factor-beta 1 precursor. J Biol Chem. 1988;263:14211-14215. [PubMed] |
| 84. | Sato C, Kim JH, Abe Y, Saito K, Yokoyama S, Kohda D. Characterization of the N-oligosaccharides attached to the atypical Asn-X-Cys sequence of recombinant human epidermal growth factor receptor. J Biochem. 2000;127:65-72. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 56] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 85. | Zheng M, Fang H, Hakomori S. Functional role of N-glycosylation in alpha 5 beta 1 integrin receptor. De-N-glycosylation induces dissociation or altered association of alpha 5 and beta 1 subunits and concomitant loss of fibronectin binding activity. J Biol Chem. 1994;269:12325-12331. [PubMed] |
| 86. | Wollscheid B, Bausch-Fluck D, Henderson C, O’Brien R, Bibel M, Schiess R, Aebersold R, Watts JD. Mass-spectrometric identification and relative quantification of N-linked cell surface glycoproteins. Nat Biotechnol. 2009;27:378-386. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 491] [Cited by in RCA: 452] [Article Influence: 28.3] [Reference Citation Analysis (0)] |
| 87. | Agard NJ, Bertozzi CR. Chemical approaches to perturb, profile, and perceive glycans. Acc Chem Res. 2009;42:788-797. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 144] [Cited by in RCA: 135] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 88. | Haun RS, Fan CY, Mackintosh SG, Zhao H, Tackett AJ. CD109 Overexpression in Pancreatic Cancer Identified by Cell-Surface Glycoprotein Capture. J Proteomics Bioinform. 2014;Suppl 10:S10003. [PubMed] |
| 89. | Haun RS, Quick CM, Siegel ER, Raju I, Mackintosh SG, Tackett AJ. Bioorthogonal labeling cell-surface proteins expressed in pancreatic cancer cells to identify potential diagnostic/therapeutic biomarkers. Cancer Biol Ther. 2015;16:1557-1565. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 23] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 90. | Zhu J, He J, Liu Y, Simeone DM, Lubman DM. Identification of glycoprotein markers for pancreatic cancer CD24+CD44+ stem-like cells using nano-LC-MS/MS and tissue microarray. J Proteome Res. 2012;11:2272-2281. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 64] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 91. | Almaraz RT, Tian Y, Bhattarcharya R, Tan E, Chen SH, Dallas MR, Chen L, Zhang Z, Zhang H, Konstantopoulos K. Metabolic flux increases glycoprotein sialylation: implications for cell adhesion and cancer metastasis. Mol Cell Proteomics. 2012;11:M112.017558. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 91] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 92. | Pan S, Chen R, Tamura Y, Crispin DA, Lai LA, May DH, McIntosh MW, Goodlett DR, Brentnall TA. Quantitative glycoproteomics analysis reveals changes in N-glycosylation level associated with pancreatic ductal adenocarcinoma. J Proteome Res. 2014;13:1293-1306. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 73] [Cited by in RCA: 85] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 93. | Foygel K, Wang H, Machtaler S, Lutz AM, Chen R, Pysz M, Lowe AW, Tian L, Carrigan T, Brentnall TA. Detection of pancreatic ductal adenocarcinoma in mice by ultrasound imaging of thymocyte differentiation antigen 1. Gastroenterology. 2013;145:885-894.e3. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 59] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 94. | Chen R, Brentnall TA, Pan S, Cooke K, Moyes KW, Lane Z, Crispin DA, Goodlett DR, Aebersold R, Bronner MP. Quantitative proteomics analysis reveals that proteins differentially expressed in chronic pancreatitis are also frequently involved in pancreatic cancer. Mol Cell Proteomics. 2007;6:1331-1342. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 113] [Cited by in RCA: 121] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 95. | Lowenfels AB, Maisonneuve P, Cavallini G, Ammann RW, Lankisch PG, Andersen JR, Dimagno EP, Andrén-Sandberg A, Domellöf L. Pancreatitis and the risk of pancreatic cancer. International Pancreatitis Study Group. N Engl J Med. 1993;328:1433-1437. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1255] [Cited by in RCA: 1139] [Article Influence: 35.6] [Reference Citation Analysis (0)] |
| 96. | Malka D, Hammel P, Maire F, Rufat P, Madeira I, Pessione F, Lévy P, Ruszniewski P. Risk of pancreatic adenocarcinoma in chronic pancreatitis. Gut. 2002;51:849-852. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 323] [Cited by in RCA: 308] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 97. | Pan S, Chen R, Stevens T, Bronner MP, May D, Tamura Y, McIntosh MW, Brentnall TA. Proteomics portrait of archival lesions of chronic pancreatitis. PLoS One. 2011;6:e27574. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 39] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 98. | Rosty C, Geradts J, Sato N, Wilentz RE, Roberts H, Sohn T, Cameron JL, Yeo CJ, Hruban RH, Goggins M. p16 Inactivation in pancreatic intraepithelial neoplasias (PanINs) arising in patients with chronic pancreatitis. Am J Surg Pathol. 2003;27:1495-1501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 70] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 99. | Torres MP, Chakraborty S, Souchek J, Batra SK. Mucin-based targeted pancreatic cancer therapy. Curr Pharm Des. 2012;18:2472-2481. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 75] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 100. | Carrara S, Cangi MG, Arcidiacono PG, Perri F, Petrone MC, Mezzi G, Boemo C, Talarico A, Cin ED, Grassini G. Mucin expression pattern in pancreatic diseases: findings from EUS-guided fine-needle aspiration biopsies. Am J Gastroenterol. 2011;106:1359-1363. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 47] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 101. | Higashi M, Yokoyama S, Yamamoto T, Goto Y, Kitazono I, Hiraki T, Taguchi H, Hashimoto S, Fukukura Y, Koriyama C. Mucin expression in endoscopic ultrasound-guided fine-needle aspiration specimens is a useful prognostic factor in pancreatic ductal adenocarcinoma. Pancreas. 2015;44:728-734. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 25] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 102. | Matsuyama M, Kondo F, Ishihara T, Yamaguchi T, Ito R, Tsuyuguchi T, Tawada K, Yokosuka O. Evaluation of pancreatic intraepithelial neoplasia and mucin expression in normal pancreata. J Hepatobiliary Pancreat Sci. 2012;19:242-248. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 14] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 103. | Yonezawa S, Higashi M, Yamada N, Yokoyama S, Goto M. Significance of mucin expression in pancreatobiliary neoplasms. J Hepatobiliary Pancreat Sci. 2010;17:108-124. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 89] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 104. | Park HU, Kim JW, Kim GE, Bae HI, Crawley SC, Yang SC, Gum JR, Batra SK, Rousseau K, Swallow DM. Aberrant expression of MUC3 and MUC4 membrane-associated mucins and sialyl Le(x) antigen in pancreatic intraepithelial neoplasia. Pancreas. 2003;26:e48-e54. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 74] [Cited by in RCA: 79] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 105. | Remmers N, Anderson JM, Linde EM, DiMaio DJ, Lazenby AJ, Wandall HH, Mandel U, Clausen H, Yu F, Hollingsworth MA. Aberrant expression of mucin core proteins and o-linked glycans associated with progression of pancreatic cancer. Clin Cancer Res. 2013;19:1981-1993. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 123] [Cited by in RCA: 136] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 106. | Chen SH, Dallas MR, Balzer EM, Konstantopoulos K. Mucin 16 is a functional selectin ligand on pancreatic cancer cells. FASEB J. 2012;26:1349-1359. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 57] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 107. | Wu YM, Nowack DD, Omenn GS, Haab BB. Mucin glycosylation is altered by pro-inflammatory signaling in pancreatic-cancer cells. J Proteome Res. 2009;8:1876-1886. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 68] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 108. | Jabbar KS, Verbeke C, Hyltander AG, Sjövall H, Hansson GC, Sadik R. Proteomic mucin profiling for the identification of cystic precursors of pancreatic cancer. J Natl Cancer Inst. 2014;106:djt439. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 42] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 109. | Binker MG, Binker-Cosen MJ, Binker-Cosen AA, Cosen-Binker LI. Microenvironmental factors and extracellular matrix degradation in pancreatic cancer. JOP. 2014;15:280-285. [PubMed] |
| 110. | Leeming DJ, Bay-Jensen AC, Vassiliadis E, Larsen MR, Henriksen K, Karsdal MA. Post-translational modifications of the extracellular matrix are key events in cancer progression: opportunities for biochemical marker development. Biomarkers. 2011;16:193-205. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 67] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 111. | Lu P, Weaver VM, Werb Z. The extracellular matrix: a dynamic niche in cancer progression. J Cell Biol. 2012;196:395-406. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2019] [Cited by in RCA: 2247] [Article Influence: 172.8] [Reference Citation Analysis (0)] |
| 112. | Malik R, Lelkes PI, Cukierman E. Biomechanical and biochemical remodeling of stromal extracellular matrix in cancer. Trends Biotechnol. 2015;33:230-236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 254] [Article Influence: 25.4] [Reference Citation Analysis (0)] |
| 113. | Multhaupt HA, Leitinger B, Gullberg D, Couchman JR. Extracellular matrix component signaling in cancer. Adv Drug Deliv Rev. 2016;97:28-40. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 121] [Cited by in RCA: 133] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 114. | Pickup MW, Mouw JK, Weaver VM. The extracellular matrix modulates the hallmarks of cancer. EMBO Rep. 2014;15:1243-1253. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1022] [Cited by in RCA: 1308] [Article Influence: 118.9] [Reference Citation Analysis (0)] |
| 115. | Willis AL, Sabeh F, Li XY, Weiss SJ. Extracellular matrix determinants and the regulation of cancer cell invasion stratagems. J Microsc. 2013;251:250-260. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 62] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 116. | Janik ME, Lityńska A, Vereecken P. Cell migration-the role of integrin glycosylation. Biochim Biophys Acta. 2010;1800:545-555. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 103] [Cited by in RCA: 115] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 117. | Grzesiak JJ, Ho JC, Moossa AR, Bouvet M. The integrin-extracellular matrix axis in pancreatic cancer. Pancreas. 2007;35:293-301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 88] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 118. | Camby I, Le Mercier M, Lefranc F, Kiss R. Galectin-1: a small protein with major functions. Glycobiology. 2006;16:137R-157R. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 595] [Cited by in RCA: 688] [Article Influence: 36.2] [Reference Citation Analysis (0)] |
| 119. | Gallagher WM, Currid CA, Whelan LC. Fibulins and cancer: friend or foe? Trends Mol Med. 2005;11:336-340. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 100] [Cited by in RCA: 109] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 120. | Rabinovich GA. Galectin-1 as a potential cancer target. Br J Cancer. 2005;92:1188-1192. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 169] [Cited by in RCA: 184] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 121. | Pan S, Chen R, Reimel BA, Crispin DA, Mirzaei H, Cooke K, Coleman JF, Lane Z, Bronner MP, Goodlett DR. Quantitative proteomics investigation of pancreatic intraepithelial neoplasia. Electrophoresis. 2009;30:1132-1144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 51] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 122. | Chen R, Pan S, Ottenhof NA, de Wilde RF, Wolfgang CL, Lane Z, Post J, Bronner MP, Willmann JK, Maitra A. Stromal galectin-1 expression is associated with long-term survival in resectable pancreatic ductal adenocarcinoma. Cancer Biol Ther. 2012;13:899-907. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 50] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 123. | Chen R, Dawson DW, Pan S, Ottenhof NA, de Wilde RF, Wolfgang CL, May DH, Crispin DA, Lai LA, Lay AR. Proteins associated with pancreatic cancer survival in patients with resectable pancreatic ductal adenocarcinoma. Lab Invest. 2015;95:43-55. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 46] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 124. | Kudo Y, Siriwardena BS, Hatano H, Ogawa I, Takata T. Periostin: novel diagnostic and therapeutic target for cancer. Histol Histopathol. 2007;22:1167-1174. [PubMed] |
| 125. | Coulson-Thomas YM, Gesteira TF, Norton AL, Kao WW, Nader HB, Coulson-Thomas VJ. The role of proteoglycans in the reactive stroma on tumor growth and progression. Histol Histopathol. 2015;30:33-41. [PubMed] |
| 126. | Chen R, Yi EC, Donohoe S, Pan S, Eng J, Cooke K, Crispin DA, Lane Z, Goodlett DR, Bronner MP. Pancreatic cancer proteome: the proteins that underlie invasion, metastasis, and immunologic escape. Gastroenterology. 2005;129:1187-1197. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 142] [Cited by in RCA: 133] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 127. | Chen WB, Lenschow W, Tiede K, Fischer JW, Kalthoff H, Ungefroren H. Smad4/DPC4-dependent regulation of biglycan gene expression by transforming growth factor-beta in pancreatic tumor cells. J Biol Chem. 2002;277:36118-36128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 68] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 128. | Köninger J, Giese T, di Mola FF, Wente MN, Esposito I, Bachem MG, Giese NA, Büchler MW, Friess H. Pancreatic tumor cells influence the composition of the extracellular matrix. Biochem Biophys Res Commun. 2004;322:943-949. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 73] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 129. | Köninger J, Giese NA, di Mola FF, Berberat P, Giese T, Esposito I, Bachem MG, Büchler MW, Friess H. Overexpressed decorin in pancreatic cancer: potential tumor growth inhibition and attenuation of chemotherapeutic action. Clin Cancer Res. 2004;10:4776-4783. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 75] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 130. | Weber CK, Sommer G, Michl P, Fensterer H, Weimer M, Gansauge F, Leder G, Adler G, Gress TM. Biglycan is overexpressed in pancreatic cancer and induces G1-arrest in pancreatic cancer cell lines. Gastroenterology. 2001;121:657-667. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 100] [Cited by in RCA: 108] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 131. | Fraser JR, Laurent TC, Laurent UB. Hyaluronan: its nature, distribution, functions and turnover. J Intern Med. 1997;242:27-33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1386] [Cited by in RCA: 1400] [Article Influence: 50.0] [Reference Citation Analysis (0)] |
| 132. | Misra S, Hascall VC, Markwald RR, Ghatak S. Interactions between Hyaluronan and Its Receptors (CD44, RHAMM) Regulate the Activities of Inflammation and Cancer. Front Immunol. 2015;6:201. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 413] [Cited by in RCA: 594] [Article Influence: 59.4] [Reference Citation Analysis (0)] |
| 133. | Turley EA, Noble PW, Bourguignon LY. Signaling properties of hyaluronan receptors. J Biol Chem. 2002;277:4589-4592. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 773] [Cited by in RCA: 774] [Article Influence: 33.7] [Reference Citation Analysis (0)] |
| 134. | Provenzano PP, Hingorani SR. Hyaluronan, fluid pressure, and stromal resistance in pancreas cancer. Br J Cancer. 2013;108:1-8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 214] [Cited by in RCA: 255] [Article Influence: 21.3] [Reference Citation Analysis (0)] |
| 135. | Itano N, Zhuo L, Kimata K. Impact of the hyaluronan-rich tumor microenvironment on cancer initiation and progression. Cancer Sci. 2008;99:1720-1725. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 116] [Cited by in RCA: 113] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 136. | Nikitovic D, Tzardi M, Berdiaki A, Tsatsakis A, Tzanakakis GN. Cancer microenvironment and inflammation: role of hyaluronan. Front Immunol. 2015;6:169. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 69] [Cited by in RCA: 91] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 137. | Provenzano PP, Cuevas C, Chang AE, Goel VK, Von Hoff DD, Hingorani SR. Enzymatic targeting of the stroma ablates physical barriers to treatment of pancreatic ductal adenocarcinoma. Cancer Cell. 2012;21:418-429. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1408] [Cited by in RCA: 1640] [Article Influence: 126.2] [Reference Citation Analysis (0)] |
| 138. | Chanmee T, Ontong P, Kimata K, Itano N. Key Roles of Hyaluronan and Its CD44 Receptor in the Stemness and Survival of Cancer Stem Cells. Front Oncol. 2015;5:180. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 109] [Cited by in RCA: 148] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 139. | Ma Z, Vocadlo DJ, Vosseller K. Hyper-O-GlcNAcylation is anti-apoptotic and maintains constitutive NF-κB activity in pancreatic cancer cells. J Biol Chem. 2013;288:15121-15130. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 166] [Cited by in RCA: 208] [Article Influence: 17.3] [Reference Citation Analysis (0)] |
| 140. | Westwood JA, Murray WK, Trivett M, Haynes NM, Solomon B, Mileshkin L, Ball D, Michael M, Burman A, Mayura-Guru P. The Lewis-Y carbohydrate antigen is expressed by many human tumors and can serve as a target for genetically redirected T cells despite the presence of soluble antigen in serum. J Immunother. 2009;32:292-301. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 48] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 141. | Sabbatini PJ, Kudryashov V, Ragupathi G, Danishefsky SJ, Livingston PO, Bornmann W, Spassova M, Zatorski A, Spriggs D, Aghajanian C. Immunization of ovarian cancer patients with a synthetic Lewis(y)-protein conjugate vaccine: a phase 1 trial. Int J Cancer. 2000;87:79-85. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 142. | Saleh MN, Sugarman S, Murray J, Ostroff JB, Healey D, Jones D, Daniel CR, LeBherz D, Brewer H, Onetto N. Phase I trial of the anti-Lewis Y drug immunoconjugate BR96-doxorubicin in patients with lewis Y-expressing epithelial tumors. J Clin Oncol. 2000;18:2282-2292. [PubMed] |
| 143. | Dziadek S, Kunz H. Synthesis of tumor-associated glycopeptide antigens for the development of tumor-selective vaccines. Chem Rec. 2004;3:308-321. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 44] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 144. | Sørensen AL, Reis CA, Tarp MA, Mandel U, Ramachandran K, Sankaranarayanan V, Schwientek T, Graham R, Taylor-Papadimitriou J, Hollingsworth MA. Chemoenzymatically synthesized multimeric Tn/STn MUC1 glycopeptides elicit cancer-specific anti-MUC1 antibody responses and override tolerance. Glycobiology. 2006;16:96-107. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 198] [Cited by in RCA: 218] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 145. | Tarp MA, Clausen H. Mucin-type O-glycosylation and its potential use in drug and vaccine development. Biochim Biophys Acta. 2008;1780:546-563. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 245] [Cited by in RCA: 231] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 146. | Harisi R, Jeney A. Extracellular matrix as target for antitumor therapy. Onco Targets Ther. 2015;8:1387-1398. [PubMed] |
| 147. | Karbownik MS, Nowak JZ. Hyaluronan: towards novel anti-cancer therapeutics. Pharmacol Rep. 2013;65:1056-1074. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 107] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 148. | Misra S, Heldin P, Hascall VC, Karamanos NK, Skandalis SS, Markwald RR, Ghatak S. Hyaluronan-CD44 interactions as potential targets for cancer therapy. FEBS J. 2011;278:1429-1443. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 401] [Cited by in RCA: 380] [Article Influence: 27.1] [Reference Citation Analysis (0)] |
| 149. | Hingorani SR, Harris WP, Beck JT, Berdov BA, Wagner SA, Pshevlotsky EM, Tjulandin SA, Gladkov OA, Holcombe RF, Korn R. Phase Ib Study of PEGylated Recombinant Human Hyaluronidase and Gemcitabine in Patients with Advanced Pancreatic Cancer. Clin Cancer Res. 2016;22:2848-2854. [PubMed] |