Published online Nov 14, 2016. doi: 10.3748/wjg.v22.i42.9257
Peer-review started: August 14, 2016
First decision: September 12, 2016
Revised: October 7, 2016
Accepted: October 30, 2016
Article in press: October 31, 2016
Published online: November 14, 2016
Processing time: 91 Days and 10.7 Hours
The intestinal microbiome is a reservoir of microbial antigens and activated immune cells. The aims of this review were to describe the role of the intestinal microbiome in generating innate and adaptive immune responses, indicate how these responses contribute to the development of systemic immune-mediated diseases, and encourage investigations that improve the understanding and management of autoimmune hepatitis. Alterations in the composition of the intestinal microflora (dysbiosis) can disrupt intestinal and systemic immune tolerances for commensal bacteria. Toll-like receptors within the intestine can recognize microbe-associated molecular patterns and shape subsets of T helper lymphocytes that may cross-react with host antigens (molecular mimicry). Activated gut-derived lymphocytes can migrate to lymph nodes, and gut-derived microbial antigens can translocate to extra-intestinal sites. Inflammasomes can form within hepatocytes and hepatic stellate cells, and they can drive the pro-inflammatory, immune-mediated, and fibrotic responses. Diet, designer probiotics, vitamin supplements, re-colonization methods, antibiotics, drugs that decrease intestinal permeability, and molecular interventions that block signaling pathways may emerge as adjunctive regimens that complement conventional immunosuppressive management. In conclusion, investigations of the intestinal microbiome are warranted in autoimmune hepatitis and promise to clarify pathogenic mechanisms and suggest alternative management strategies.
Core tip: The intestinal microbiome is a reservoir of microbial antigens and activated immune cells that have been implicated in the pathogenesis of diverse systemic immune-mediated diseases. Dysbiosis, increased intestinal permeability, and molecular mimicry between microbial and self-antigens may initiate or sustain autoimmune hepatitis. Multiple drug, molecular, dietary, and probiotic interventions can modify the intestinal microbiome and attenuate the immune response. The role of the intestinal microbiome in autoimmune hepatitis warrants rigorous study, and new therapies may emerge that strengthen current treatment regimens.
- Citation: Czaja AJ. Factoring the intestinal microbiome into the pathogenesis of autoimmune hepatitis. World J Gastroenterol 2016; 22(42): 9257-9278
- URL: https://www.wjgnet.com/1007-9327/full/v22/i42/9257.htm
- DOI: https://dx.doi.org/10.3748/wjg.v22.i42.9257
Albert J Czaja, MD is Professor Emeritus of Medicine at the Mayo Clinic College of Medicine, Rochester, Minnesota, United States (Figure 1). He graduated from Dartmouth College in 1965, received a Bachelor of Medical Science at the Dartmouth Medical School in 1966, and MD from Harvard Medical School in 1968. He was trained in internal medicine at the Philadelphia General Hospital, University of Pennsylvania Division, from 1968-1972, and he was a major in the United States Army Medical Corps from 1972-1975 at the United States Army Institute of Surgical Research at Fort Sam Houston, Texas. Dr. Czaja was then an NIH Research Fellow in hepatology at the Mayo Clinic under the mentorship of W.H.J. Summerskill, MD from 1975-1977. He joined the staff of the Mayo Clinic Division of Gastroenterology and Hepatology in 1977, and he was appointed Professor of medicine in 1986. His research has focused on chronic hepatitis, especially autoimmune hepatitis, and he has contributed to the understanding of its diagnosis, treatment, prognosis, genetic predispositions, pathogenic mechanisms, and consequences. He has collaborated and published with investigators from 15 countries, and he has mentored physicians who have become leaders in their academic institutions and professional organizations. Dr. Czaja is a Fellow of the American College of Physicians, American College of Gastroenterology, American Gastroenterological Association, and American Association for the Study of Liver Diseases. He received the Meritorious Service Medal of the United States Army Medical Corps in 1976; the Fiterman award for distinguished clinical investigation in hepatology by the American Gastroenterological Association in 1997; the Henry S. Plummer Distinguished Physician Award by the Mayo Clinic Department of Medicine in 2006; the Distinguished Clinician Award of the American Gastroenterological Association in 2007; the Distinguished Clinician Educator Award (1st recipient) of the American Association for the Study of Liver Diseases; the Gold Medal of the Canadian Association for the Study of the Liver and the Canadian Liver Foundation for the advancement of hepatology in autoimmune liver diseases and the education of young clinicians in 2013; and the Mayo Clinic Distinguished Alumnus Award in 2016. Dr. Czaja was a founding member of the International Autoimmune Hepatitis Group in 1992, and he continues to describe investigational pathways that promise to improve the treatment strategies of autoimmune liver disease. He has written over 540 articles, including 87 book chapters.
Autoimmune hepatitis is a chronic immune-mediated inflammatory liver disease of uncertain cause[1,2]. Population-based epidemiological studies have indicated that it is a rare chronic liver disease with an annual incidence of 0.67-1.9 cases per 100000 persons and a point prevalence of 4-42.9 cases per 100000 persons[3-8]. Prevalence varies widely by geographical region (Singapore, 4 per 100000[4]; Sweden, 10.7 per 100000[9]; southern Israel, 11 per 100000[8]; Spain, 11.6 per 100000[6]; Norway, 16.9 per 100000[3]; Netherlands, 18.3 per 100000[10]; Denmark, 23.9 per 100000[11]; New Zealand, 24.5 per 100000[7]; and Alaska, 42.9 per 100000[5]), and the incidence has been increasing in Denmark[11] and the Netherlands[10]. Rigorous epidemiological studies have not been performed in the adult population of the United States, but studies in the children of Utah have indicated an overall incidence of 0.4 cases per 100000 and a prevalence of 3 cases per 100000[12]. The wide variability in the annual incidence and point prevalence of autoimmune hepatitis in different geographical regions and ethnic groups suggests that genetic and environmental factors contribute to its occurrence[2,13,14].
Genetic factors within[15-17] and outside[18-22] the major histocompatibility complex (MHC) have been implicated as susceptibility factors for autoimmune hepatitis. Cytochrome P450 2D6 (CYP2D6) and formiminotransferase cylcodeaminase have been proposed as key antigenic targets in some patients[23-25], and homologies between peptide sequences in CYP2D6 and hepatitis C virus (HCV), herpes simplex virus (HSV), and cytomegalovirus (CMV) have suggested that molecular mimicry between foreign and self-antigens initiates and sustains the disease[2,23,26-28].
The principal target antigen in most white North American and northern European adults with autoimmune hepatitis is unknown, and as yet unrecognized self-antigens or foreign antigens that resemble self-antigens may trigger the disease or increase susceptibility to it, possibly by skewing components of the innate and adaptive immune responses toward a pro-inflammatory, autoreactive profile[2,29]. The commensal bacteria of the intestine and their metabolic by-products constitute a reservoir of foreign antigens that can interact with mucosal immune cells and influence systemic immune responses[30-34]. The human microbiome has already been implicated in the occurrence of multiple systemic immune-mediated diseases, including type 1 diabetes[35-37], rheumatoid arthritis[38-41], multiple sclerosis[42], and inflammatory bowel disease[43-45], and its role in the inflammatory liver diseases is also being scrutinized[46-48].
Non-alcoholic steatohepatitis (NASH) may progress because of an influx of microbial products in the portal circulation, activation of toll-like receptors (TLRs) 4 and 9, and subsequent release of pro-inflammatory cytokines, including tumor necrosis factor-alfa (TNF-α) and interleukin (IL)-1β[49-51]. Primary sclerosing cholangitis (PSC) expresses high levels of TLR4 and TRL9 in biliary epithelial cells (BEC), produces IL-1β, IL-8, and interferon (IFN)-γ in response to lipopolysaccharide (LPS), and commonly manifests atypical perinuclear anti-neutrophil cytoplasmic antibodies (pANCA)[52-54]. These antibodies are directed against β-tubulin which cross-reacts with an antigen (FtsZ) present in all intestinal bacteria[53]. Germ-free mice develop histologically more severe PSC than conventionally housed animals, and these findings suggest that commensal bacteria have a protective role against PSC[55].
Similarly, primary biliary cholangitis (PBC) expresses TLR4 in BEC and periportal hepatocytes, produces pro-inflammatory cytokines (IL-1β, IL-6, IL-8, and TNF-α) in response to intestinal bacterial components [LPS, flagellin, and cytosine-phosphorothioate-guanine oligonucleotide (CpG)], manifests BEC-destructive LPS-stimulated natural killer cells, and produces antimitochondrial antibodies that target an antigen (pyruvate dehydrogenase complex-E2) which shares sequence homologies with intestinal Escherichia coli[56-62].
Autoimmune hepatitis may also be influenced by the intestinal microbiome. Concurrent features of PBC or PSC occur in 7%-18% of patients (“overlap syndromes”)[63,64]; atypical pANCA are present in 49%-92% of individuals with autoimmune hepatitis[65-68]; and alterations in the composition of the intestinal microbiota (dysbiosis) have been found in experimental autoimmune hepatitis[69]. The structural proteins binding intestinal epithelial cells (zona occludens 1 and occludin) are reduced in patients with autoimmune hepatitis compared to healthy volunteers; plasma LPS levels are increased; and the numbers of intestinal anaerobes (Bifidobacterium and Lactobacillus) are decreased[70]. These findings support the concept that autoimmune hepatitis is associated with dysbiosis, increased permeability of the gastrointestinal mucosal barrier, and translocation of gut-derived microbial products into the systemic circulation.
The goals of this review are to describe the role of the intestinal microbiome in generating innate and adaptive immune responses, indicate how these responses may contribute to the development or maintenance of systemic autoimmune responses, and encourage investigations in autoimmune hepatitis that might improve understanding of its pathogenesis and results of its management.
Abstracts cited in PubMed were identified using the search words “intestinal microbiome”, “intestinal microbiome and autoimmunity”, and “intestinal microbiome and autoimmune hepatitis”. Key aspects of the abstracts judged pertinent to the review were noted, and full-length articles were selected from the abstracts. A secondary bibliography was developed from the references cited in the selected full-length articles, and additional PubMed searches were performed to expand the concepts developed in these articles. The discovery process involving abstract review and the acquisition of full-length articles was repeated, and a tertiary bibliography was developed after reviewing these selected articles. The number of abstracts cited by PubMed and reviewed for pertinence to this topic during the primary, secondary and tertiary searches exceeded 3800. Those judged most pertinent to the topic exceeded 200, and the number of full-length articles reviewed was 66.
The human intestinal tract contains 10-11 trillion bacteria comprising 500-1500 different species[47,71-76]. Actinobacteria, Firmicutes, Proteobacteria, and Bacteroidetes are the four phyla that predominate[77-79], and a phylogenetic core of microbial species has been defined that is composed of 66 Operational Taxonomic Units (OTUs) present in most individuals[80]. Luminal microbiota have greater diversity, and they are more tightly clustered than mucosal microbiota[81]. Firmicutes and Actinobacteria are more abundant in the luminal populations, and Proteobacteria are more abundant in the mucosal populations[81].
The composition of the microbiome is influenced by diverse environmental factors, including community sanitation levels and vaccination programs, and by host-related variables, including method of obstetrical delivery, age, genetic predisposition, dietary habits, personal hygiene, and antibiotic exposures. Changes in the intestinal microbiome tend to be slow from late childhood through adulthood with marked changes occurring mainly with advanced age[74,82-85]. The microbiome becomes less diverse and more variable over short intervals with aging, and the species of Bacteroides, Clostridium and Escherichia coli constitute a greater proportion of the microflora in individuals aged ≥ 65 years[82-84].
The intestinal microbiome varies in diverse ethnic groups[75,86-89], and this diversity may reflect genetic factors, demographic issues (age, gender, socioeconomic status), lifestyle features (alcohol use, smoking, adiposity), and long-term diet[90-92]. Disparities in the intestinal microbiota have been recognized between ethnic groups in the same country (rural versus urbanized)[88,89,93,94] and between countries (Africa versus Europe, cross-Europe, and cross-Asia)[87,88,91], and socioeconomic variations at individual and neighborhood levels have been associated with many of these differences[88,95]. The nature of the long-term diet may be the critical element affected by the socioeconomic status[95].
Amongst the diversity within the intestinal microbiota, common functional and phylogenetic elements have also been described[89,96-99]. These common elements may be indispensable for the well-being of the individual as they can produce short-chain fatty acids, synthesize vitamins, and aid in digestion, metabolism and immune defense[89]. The functional and phylogenetic core components have been shared across heterogeneous healthy human populations, and they tend to co-exist[89].
The intestinal microbiome is essential for development of the intestinal immune responses which in turn maintain tolerance of the microflora[41,100,101]. Germ-free mice have fewer CD4+ T lymphocytes in the lamina propria of the intestine, hypoplastic Peyer’s patches, less immunoglobulin A production, and disorganized zones of T and B lymphocytes in the spleen and lymph nodes compared to wild-type mice[102,103]. These immune deficiencies are corrected by the introduction of Bacteroides fragilis[104]. Colonization also induces the production by IL-10-secreting, regulatory T cells (Tregs), possibly in response to the secretion of polysaccharide A by the bacteria and direct activation of TLR2 on the Foxp3+ Tregs[105-108]. The introduction of Clostridium species induces similar changes[109,110].
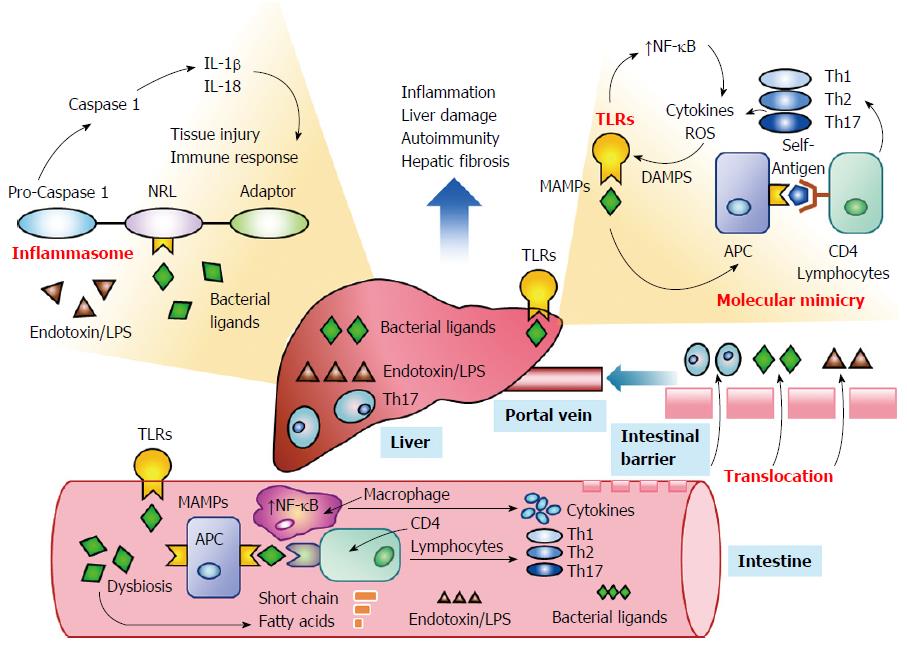
Emerging evidence suggests that the intestinal microbiome can influence systemic immune responses by activating TLRs[34,47,48] and promoting the formation of inflammasomes within the liver[51,111-113] (Figure 2). Changes in the composition of the intestinal microbiome induced by antibiotics, genetic factors, or the disease (dysbiosis) may sustain or enhance the innate and adaptive immune responses by overcoming or circumventing normal tolerogenic responses to the commensal bacteria[32,114-116]. Bacterial components may act as antigens that stimulate the systemic immune response[30,31,34,47,48] or that prime immune cells within the intestine that subsequently access peripheral lymphoid tissue[39,117,118]. Molecular mimicry between microbial and host-derived antigens and promiscuous targeting by antigen-sensitized lymphocytes may then initiate or strengthen the autoreactive response in genetically-predisposed individuals[33,57,115]. Investigations in cell cultures, animal models, and patients with diverse systemic immune-mediated diseases have justified these hypotheses and warranted their consideration in autoimmune hepatitis[46-48].
TLRs are the key receptors within the intestine that recognize microbe-associated molecular patterns, pathogen-associated molecular patterns, and damage-associated molecular patterns[34,47,48,119,120] (Table 1). They are instrumental in generating an innate immune response to pathogens and cellular distress signals, and they can shape subsets of T helper (Th) lymphocytes that recognize microbial components and have the potential to cross-react with host antigens[41,121,122]. Ten TLRs have been described in humans[123], and each responds preferentially to specific ligands which may be viral and bacterial proteins or endogenous ligands in the absence of infection[34,120]. All stimulated TLRs except TLR3 activate a signaling pathway that is dependent on the myeloid differentiation factor 88 (MyD88)[124,125]. Signaling through the MyD88 pathway in turn activates nuclear factor-kappa B (NF-κB) and promotes the transcription of pro-inflammatory cytokines (TNF-α, IL-1β, and IL-6)[123,125,126].
| Key requirement | Features | Mechanisms |
| Activation of TLRs | Intestinal receptors responsive to MAMPs and DAMPS[34,47,48,120] | Increases pro-inflammatory cytokines[126] |
| Signaling dependent on MyD88[124,125] | Upregulates class II MHC[127] | |
| Activates NF-κB[123,125,126] | Increases co-stimulatory molecules[41,127] | |
| Favors T lymphocyte activation[41] | Promotes pathogen-specific responses[142] | |
| Modulates actions of Tregs[141,142] | LPS activates TLR4[123,134] | |
| Present in hepatocytes, HSCs, Kupffer cells, sinusoidal epithelial cells, BEC[48] | Sequences in bacteria activate TLR9[135] | |
| TLR4 in HSCs promote fibrosis[46,144] | ||
| Implicated in other liver diseases[58,148] | ||
| Stimulation of inflammasomes | Protein complexes that release pro-inflammatory IL-1β and IL-18[111-113] | Upregulated in hepatocytes by LPS[113] |
| NLRs sense microbial products[156] | Activates pro-caspase 1[156] | |
| Upregulated in Kupffer cells, hepatocytes, and sinusoidal epithelial cells[113] | Promotes hepatic fibrosis[50] | |
| Activation by highly diverse ligands[112] | Shapes innate and adaptive immunity[112,160] | |
| Implicated in NAFLD[51] | ||
| Activation separate from TLRs[112,155] | ||
| Emergence of dysbiosis | Microflora differ from commensals[116] | Can activate TLRs and NLRs[116,173] |
| Dysbiosis varies in specific diseases[116] | Genetic factors may affect composition[177] | |
| Less bacterial diversity common[170] | Gender-related compositional differences[179] | |
| Antibiotics most frequent basis[165,175] | May affect gender-related autoimmunity[180] | |
| Uncertain cause or effect of disease[116] | Present in AIH and experimental NASH[47,69] | |
| Molecular mimicry | Microbial and self-homologies[33,185] | pANCA react with bacterial antigen[53] |
| Cross-reacting antibodies[53,57,184] | AMA cross-reacts with Escherichia coli[56,57] | |
| Promiscuous activity of effectors[186] | Increasingly distant homologues targeted[187] | |
| Epitope spread[187] | ||
| Breech of intestinal mucosal barrier | Gut-derived products enter system[195] | Gut-derived lymphocytes in lymph nodes[118] |
| Translocation prime basis[46,195] | Microbial components in peripheral blood[195] | |
| Active transport also possible[230] | Activates TLRs and NLRs[123,130] | |
| Implicated in NASH and diabetes[197,198] |
TLRs can also influence the adaptive immune response[41] (Figure 2). TLRs expressed by dendritic cells and macrophages can upregulate class II molecules of the MHC and enhance antigen presentation to CD4+ helper T lymphocytes[127] (Table 1). TLRs can also increase the expression of the co-stimulatory molecules, CD80, CD86 and CD40, on the antigen presenting cells and thereby favor T lymphocyte activation and differentiation[41,127]. Kupffer cells express all TLRs except TLR5[128], and they are the primary cells within the liver that respond to TLR ligands. The production of pro-inflammatory cytokines, chemokines, and reactive oxygen species by the Kupffer cells promotes liver inflammation[129] and the innate and adaptive immune responses[41,127]. Hepatocytes, BEC, hepatic stellate cells (HSCs), and sinusoidal epithelial cells also express TLRs, but only HSCs express TLR1 through TLR9[48,130].
The cytokine profile shapes the subsets of T lymphocytes that constitute the immune-mediated response, and it is influenced by the particular TLRs that are activated by ligands within the microenvironment (Table 1). Activation of TLR4 and TLR9 promotes the release of IL-12 and favors a type 1 cytokine pathway that is pro-inflammatory[131]. TLR4 also induces the secretion of IL-23 and promotes the expansion and survival of pro-inflammatory Th 17 lymphocytes[132]. In contrast, activation of TLR2 favors the production of IL-10 and IL-13 which promotes an anti-inflammatory type 2 cytokine response[133].
LPS from gram-negative bacteria is the principal ligand activating TLR4[123,134], and un-methylated CpG sequences in bacterial and viral genomes activate TLR9[135] (Table 1). The viral proteins of HCV, CMV and HSV are key ligands that activate TLR2[34,136,137]. TLR2, TLR5, TLR7, and TLR8 are expressed by CD4+ T lymphocytes, and ligands that activate these TLRs (viral proteins[136], flagellin[138], and single-stranded ribonucleic acid[139]) can directly activate memory lymphocytes and stimulate their proliferation[140]. Naturally occurring Tregs express TLR2, TLR5 and TLR8, and they can also be activated directly by viral and bacterial components[141]. TLRs can also block the suppressive effect of Tregs by the recognition of microbial products that induce secretion of IL-6[142]. Pathogen-specific adaptive immune responses can be favored, and defense mechanisms can be strengthened. Microbial elements can thereby modulate the innate and adaptive immune responses through the TLRs and affect immune homeostasis indirectly by modulating the cytokine profile or directly by affecting immune cell proliferation.
TLR4 is a crucial signaling pathway by which HSCs increase the extracellular matrix[46] (Table 1). The production of chemokines and adhesion molecules is mediated by activated TLR4 in HSCs[46], and the chemo-attraction of inflammatory and immune cells to the liver stimulates the fibrotic process[143]. TLR4 signaling also promotes activation of transforming growth factor-beta (TGF-β) by down-regulating production of an endogenous inhibitor of the TGF-β receptor[144]. Furthermore, TLR4 signaling may down-regulate microRNA molecules that suppress the transcription of collagen[46,145]. A polymorphism of the TLR4 gene may impair the response of TLR4 to LPS. The LPS-induced signaling pathway that activates NF-κB may thereby be disrupted and the production of the pro-inflammatory cytokines, TNF-α and interferon-beta, may be reduced[146]. In this fashion, a genetic variation may affect the response of TLR4 to microbial ligands and the propensity for progressive hepatic fibrosis.
The signaling pathway involving TLR4, MyD88, and NF-κB has been implicated in the progression of multiple liver diseases[54,58,144,147,148]. Concentrations of gut-derived endotoxins have been increased in animal models of hepatic fibrosis[149,150] and in the systemic and portal circulation of patients with cirrhosis[151,152]. Other TLRs may respond to different microbial ligands and influence the subsets of lymphocytes that orchestrate the autoreactive response. TLR signaling pathways have not been evaluated in autoimmune hepatitis.
Inflammasomes are protein complexes that form within the cytoplasm of diverse cells, including macrophages, hepatocytes, and HSCs, in response to stimuli associated with cellular stress, damage or infection[112,153,154] (Table 1). By releasing pro-inflammatory cytokines IL-1β and IL-18, they drive the inflammatory response to tissue injury and influence cell death, inflammatory activity, and fibrosis[111,113] (Figure 2). TLRs and inflammasomes have separate routes of activation[112,155], but cooperation between them is pivotal in promoting communication between the intestinal microbiota and the systemic immune response[34]. Factors that increase the expression of inflammasomes, such as saturated fatty acids and bacterial endotoxin, may increase activation of TLR4 and promote hepatic fibrosis[50].
Inflammasomes consist of a sensor protein that is within the family of non-obese diabetes (NOD)-like receptors (NLRs), an adaptor molecule (apoptosis-associated speck-like CARD-domain containing protein), and pro-caspase 1[156] (Table 1). The inflammasomes can sense microbial products[157] and metabolic stress[112], activate pro-caspase 1[158,159], trigger the release of pro-inflammatory cytokines[153], and shape the innate[112] and adaptive immune responses[160]. The expression of NLRP3 is upregulated in hepatocytes after stimulation with LPS, and Kupffer cells and sinusoidal epithelial cells also express high levels of NLRP1 and NLRP3[113].
The structural diversity of the ligands that activate NLRP3 is greater than the structural motifs that activate the TLRs, and the inflammasomes may be responsive to a broader range of activation signals than the TLRs[161] (Table 1). Together the TLRs and NLRs provide receptors for signaling pathways that can respond to diverse endogenous and exogenous danger signals, including microbial components, and they each can generate pro-inflammatory responses that sustain and enhance the innate and adaptive immune responses to liver injury. TLRs may also have a counter-regulatory effect on the inflammasomes[34]. Chronic stimulation of the TLRs by LPS induces the production of IL-10 and reduces the activation of NLRP3[162]. Furthermore, activation of TLR2 or TLR4 can increase the autophagy of hepatocytes, the degradation of NLRP3, and the suppression of IL-1β production[163]. Inflammasomes have not been characterized in autoimmune hepatitis, and their interactions with TLRs have not been defined in this disease.
Systemic inflammatory and immune-mediated diseases have been associated with intestinal microbiomes that distinguish them from normal or other disease-specific populations[116] (Table 1). The intestinal microbiota have differed in patients with rheumatoid arthritis compared to patients with fibromyalgia[164]. Decreased diversity in the microbiome has been associated with an increased risk of type 1 diabetes[165], the occurrence of atopic diseases, including asthma[166-169], and the presence of Crohn’s disease[170]. Patients with multiple sclerosis have reduced numbers of Clostridia and Bacteroides species compared to normal individuals[42], and patients with type 1 diabetes have more colonies of Bacteroides[37,171]. Compositional shifts in the intestinal microbiota, particularly the relative frequencies of certain bacterial taxa, have been associated with the phenotype and genotype of inflammatory bowel disease[172].
These findings have suggested that alterations in the composition of the intestinal microflora (dysbiosis) may disrupt intestinal and systemic immune tolerances and contribute to immune-mediated diseases[114,116]. Depletion of the commensal bacteria may allow intestinal populations of pathogenic or immunogenic organisms to proliferate and generate ligands that activate TLRs and NLRs[116,173] (Figure 2). The major uncertainty has been whether the dysbiosis has been a cause or an effect of the disease.
Antibiotics are the principal agents promoting dysbiosis, and their use has been implicated in creating the dysbiosis associated with the occurrence of atopic diseases[167,168,174], asthma[166], type 1 diabetes[165], and celiac disease[175]. Twin studies have also indicated that genetic factors can shape the intestinal microbiome[176,177], and immune-mediated diseases with genetic predispositions have manifested dysbiosis[30,47,116]. Importantly, the contribution of the 8.1 ancestral haplotype, which includes the DRB1 alleles commonly associated with systemic autoimmune diseases including autoimmune hepatitis, is probably small[178], and dysbiosis rather than genetic factors has been implicated in the occurrence of experimental NASH[51].
The composition of the intestinal microflora may also influence the gender bias for autoimmune disease[179-181] (Table 1). Colonization by commensal microbes early in the life of NOD mice raises serum testosterone levels and protects male mice from developing type 1 diabetes[179]. Furthermore, the transfer of the intestinal microbiota from mature male NOD mice to immature female NOD mice alters the intestinal microbiome of the females and protects them from developing diabetes[179]. Blockade of the androgen receptor attenuates the microbiome-specific changes in the female mice and supports the concept that the commensal bacteria of the intestine can affect the propensity for autoimmune disease in genetically-susceptible animals by altering sex hormone levels or receptor sensitivities[179].
Gender may also influence the composition of the intestinal microbiota and in turn the propensity to develop autoimmune disease[180] (Table 1). The intestinal microbiota differ in male and female NOD mice, and this difference disappears after male castration. Furthermore, the greater frequency of female NOD mice to develop type 1 diabetes compared to male NOD mice is lost in germ-free animals[180]. These findings suggest that the intestinal microbiome can influence sex hormone levels[179] and also be influenced by them[180], possibly in a self-amplification loop.
The increased female propensity for autoimmune disease may relate to estrogenic effects that modulate the autoreactive response directly by affecting the pro- and anti-inflammatory cytokine pathways of lymphocyte differentiation[182] and indirectly by altering the intestinal microbiome to favor the translocation of sensitizing microbial antigens[180]. Imbalances between blood estrogen and progesterone levels have affected the immune response during and immediately after pregnancy[182], and the treatment of peripheral blood mononuclear cells with 17-β estradiol has increased their response to immunogens and the expression of TLR8[183]. The relationship between sex hormone levels and the intestinal microbiome during pregnancy, menses, and menopause remains uncertain and important to clarify.
Multiple factors can promote dysbiosis, and a pathogenic or circumstantial relationship with autoimmune disease has not been established. Nevertheless, the association of dysbiosis with diverse systemic immune-mediated diseases[116], its recognition in autoimmune hepatitis[69,70], its possible genetic associations[176,177], and its gender bias[179-181] suggest that dysbiosis may constitute an important antigenic or hormonal reservoir that can promote the autoreactive response in diverse systemic immune diseases, including autoimmune hepatitis.
Epitopes can be shared between microbial components and self-antigens, and this molecular mimicry can result in cross-reacting antibodies or the activation of T lymphocytes in genetically-predisposed individuals[33,184,185] (Figure 2). The reactive T lymphocytes can in turn exhibit promiscuous activity[186] and target epitopes that are distant homologues to the initial antigenic trigger (epitope spread)[187] (Table 1). Bacterial components have generated autoantibodies found in systemic autoimmune diseases, such as systemic lupus erythematosus (SLE)[188] and the antiphospholipid syndrome[189], and they have been implicated in the progression of SLE[190] and the exacerbation of Sjogren’s syndrome, possibly through the activation of memory lymphocytes[33,188,191]. The atypical pANCA found in PSC and autoimmune hepatitis target an antigen (β-tubulin) that cross-reacts with a bacterial antigen[53], and the antibodies to pyruvate dehydrogenase complex-E2 found in PBC cross-react with intestinal Escherichia coli[56,57]. The principal mechanism by which the intestinal microbiota may sustain or extend the autoreactive response is molecular mimicry, and experimental animal models of autoimmune hepatitis should evaluate this hypothesis.
Interactions between the intestinal microenvironment and the systemic immune response imply that the natural barrier between the intestinal and systemic domains can be breeched (Table 2). There is ample evidence to justify this supposition, but the actual mechanisms are uncertain. Reactive T lymphocytes expressing intestinal receptors can be found in the pancreatic islets and lymph nodes of patients and mice with type 1 diabetes[117,118,192,193], and lymphocytes originating in the gut mucosa have been implicated in the autoreactive response in experimental autoimmune encephalomyelitis[194]. Furthermore, microbial components have been detected in the plasma of patients with cirrhosis[151,152] and portal circulation of animals with non-alcoholic fatty liver disease[51].
| Microbial Effect | Features | Mechanisms |
| Translocation | Migration of gut-derived products[195,224] | Gut-derived SCFA affect tight junctions[200] |
| Tight junctions weakened[218] | Butyrate strengthens intestinal barrier[203] | |
| Increased intestinal permeability[195,218] | Induces mucin synthesis[201,203] | |
| Paracellular migration[37,224] | Reduces bacterial translocation[204] | |
| Consequences[192] | Increases peripheral Tregs[205] | |
| LPS and CpG delivered to liver[123,130,195] | Inhibits NF-κB and inflammation[207] | |
| Activated immune cells translocate[118,193] | Lactate strengthens intestinal barrier[37] | |
| Translocated microbial antigens activate peripheral immune cells[185] | Fermented to butyrate[215,216] | |
| TLRs and NLRs activated[123,130] | Low butyrate- and lactate- producing bacteria associated with weak barrier[217,218] | |
| Increased mucosal permeability | Intestinal epithelial cells bound together by junctional complex of proteins[222,223] | TLRs affect molecular mediators[225,226] |
| Occludin main component[222] | Signaling pathways disrupted[223] | |
| Zona occludens couples cytoskeleton[222] | Junctional binding proteins dissociated[224] | |
| Cingulin contacts cells[222] | Paracellular migration routes formed[37,224] | |
| Actin and myosin anchor cells[222] | E. coli and C. difficile key effectors[37] | |
| Intermediate filaments bind cells[222] | ||
| Signaling pathways seal junction[223] | ||
| Protein kinase C modulates occludin[222] | ||
| Active transport | Bacterial antigens actively transported across intestinal barrier[230] | M cells in Peyer’s patches capable of active transport[230] |
The migration of gut-derived bacteria and bacterial products from the intestinal lumen to the liver, mesenteric lymph nodes, and other extra-intestinal sites may occur by translocation[195] (Figure 2). Translocation implies that intestinal permeability has been increased, possibly because tight junctions within the intestinal mucosa have been weakened or the intestinal barrier has been overwhelmed by bacterial overgrowth[46,195-197]. The translocated bacterial products, including LPS and unmethylated CpG, can then be delivered to the liver via the portal vein and activate TLRs and NLRs[123,130]. Dysbiosis and a “leaky gut” have been implicated in the development of NASH, diabetes, and the metabolic syndrome[197,198].
Bacteria produce short chain fatty acids (acetic acid, butyric acid, and propionic acid) which can affect the tight junctions within the intestinal mucosa[197,199,200] (Table 2). Butyrate, which is the conjugate base of butyric acid, induces mucin synthesis in the intestinal mucosa, strengthens tight junctions, and reduces bacterial transport across the stressed epithelium[201-204]. Butyrate may also have an anti-inflammatory effect by promoting the extra-thymic differentiation of peripheral Tregs[205,206] and inhibiting NF-κB and the transcription of pro-inflammatory cytokines[207]. Other short chain fatty acids (propionate) and bacterial by-products (succinate and acetate) do not induce the production of mucin and may increase gut permeability[37,202].
Sodium butyrate acts in part by modulating the beta-catenin-dependent Wnt signaling pathway within cells[208]. This pathway affects the transcription of genes that influence cell proliferation and differentiation. In colon carcinoma cell lines, the levels of beta-catenin transcriptional complexes within the cell influences its physiological response to butyrate. High levels of transcriptional complexes result in apoptosis of the cell and low levels result in the reversible limitation of cell growth after exposure to butyrate[208]. The ability of butyrate to modulate cell proliferation and apoptosis may in turn influence cell viability and function, and these effects may help maintain the integrity of the gastrointestinal mucosal barrier.
Butyrate can also modulate cell responses to stress of the endoplasmic reticulum by promoting the apoptosis of the cell or its preservation through autophagy[209-211]. Butyrate enhances the expression of peroxisome proliferator-activated receptor-gamma and the activation of caspases (especially caspase 3) that induce apoptosis in colorectal cell lines[212]. It is also one of the short chain fatty acids, including propionate, that can induce autophagy in distressed cells and preserve their survival by generating energy and retarding the intrinsic (mitochondrial) pathway of apoptosis[213,214]. Gut-derived short chain fatty acids, such as butyrate and propionate, may be important moderators of intestinal mucosal cell proliferation and function, and they may contribute to the prevention of systemic autoimmune responses and progressive colorectal cancer[208].
Lactate, which is a bacterial byproduct of carbohydrate fermentation, also reduces intestinal permeability[203] (Table 2). Lactate is fermented mainly to butyrate by intestinal microflora[215]. The acetyl-coenzyme A pathway is the major route of butyrate production from lactate, and the intestinal microflora have considerable variability in lactate consumption[215]. Furthermore, lactate-utilizing bacteria exhibit variable production of butyrate depending on the availability of other substrates[216]. Patients with type 1 diabetes have a lower proportion of butyrate- and lactate-producing bacteria in their intestinal microbiome than case-control subjects, and the dysbiosis favoring increased intestinal permeability may contribute to the development of type 1 diabetes[217,218].
Intestinal epithelial cells are bound together by structural proteins[37,219-221] that are organized into a tripartite junctional complex consisting of a tight junction, adherens junction, and desmosome[222]. Occludin is the only known transmembrane protein with a domain in the paracellular space, and it is the principal component of the tight junction. Zona occludens 1 and 2 and cingulin are non-transmembrane proteins found in tight junctions at sites of cell-to-cell contact[222]. They are bound to occludin, and they probably couple cells to the cytoskeleton[222,223]. Actin and myosin filaments anchor cells together by calcium-dependent adhesion molecules (E-cadherins) in an adherens junction, and intermediate filaments are anchored to desmosomes and help bind cells[222]. Multiple cellular signaling pathways affect the assembly and sealing of the junctions[223], and they are cell-type specific with protein kinase C modulating occludin and zona occludens 1[222].
Escherichia coli and Clostridia difficile can dissociate the binding proteins and increase intestinal permeability by opening a paracellular route[37,224] (Table 2). The TLRs on the intestinal epithelial cells can modulate the integrity of the intestinal barrier, possibly by influencing the expression of molecular mediators that can affect the structure or function of the binding proteins[225,226]. Activation of TLR2 increases the phosphorylation of isoforms of protein kinase C, and this action has been associated with enhanced expression of zona occludens and the sealing of tight junctions[225]. Conversely, activation of TLR4 reduces the expression of phosphorylated occludin and increases intercellular permeability[226]. Bacterial ligands derived from different microbial species may influence intestinal permeability through TLR signaling and the translocation of microbial products through a porous intestinal barrier may contribute to a systemic autoreactive response[227-229].
Another mechanism by which the microbiome may influence the systemic immune response is by the active transport of bacterial antigens across the mucosal barrier by M cells within Peyer’s patches[230]. Whereas immune cells can be activated in the intestine and migrate to the liver or peripheral lymph tissue by translocation, they may also be activated in the systemic circulation by translocated or actively transported bacterial components that are presented by antigen presenting cells and recognized as foreign antigens by circulating naïve CD4+ T helper lymphocytes[39,117,118,193].
The intestinal microbiome has been implicated in the pathogenesis of diverse inflammatory liver diseases, including NASH, PSC and PBC[48,51,54,59,62], and the pathogenic pathways of autoimmune hepatitis have been incompletely defined[2]. The principal target antigen remains unclear in most patients with autoimmune hepatitis, and conventional corticosteroid therapies have been unable to consistently induce sustained treatment-free remissions[231,232]. Progressive hepatic fibrosis occurs in 25% of patients[233], and emerging drug therapies and molecular interventions can suppress the immune response but not eliminate the disease[234].
The intestinal microbiome is a reservoir of antigens and activated immune cells that could initiate, exacerbate, or perpetuate autoimmune hepatitis[30-34]; dysbiosis has been demonstrated in experimental and human autoimmune hepatitis[69,70]; atypical pANCA, manifested in most patients with autoimmune hepatitis, cross-react with an antigen found in intestinal bacteria[53]; and the permeability of the intestinal mucosal barrier has been increased in patients with the disease[70]. Investigations of the microbiome in autoimmune hepatitis may discover new antigens and suggest new therapies that might eliminate the primary antigen or the supplemental antigens that sustain or advance the disease.
Traditional stool culture techniques are limited in assessing the intestinal microbiome mainly because anaerobic organisms are difficult to culture[235] and some microbial species may elude detection by conventional protocols[235,236]. A common method for studying the diversity of the intestinal microflora has been to sequence the 16S ribosomal ribonucleic acid (rRNA) gene[235,237]. The 16S rRNA gene is present in all prokaryotic cells; it has highly variable regions interspersed with highly conserved regions; and its sequences are unique to the major groups of prokaryotic organisms[237]. These signature sequences can be used to reconstruct the phylogeny of the intestinal microbiome[236].
Primers are designed that are complementary to the universally conserved regions that flank the variable regions, and the bacterial species and their proportions in the microbiome are determined[237,238]. The variable regions are amplified by polymerase chain reaction (PCR), and the PCR products are purified for sequencing[81]. Sequencing results are compared to established annotated datasets[235]. The sequencing protocol misidentifies or omits a critical residue in only 1% of procedures[237]. Ambiguities resulting from the sequencing determinations are the most common errors, and they may reflect inadequacies in the available datasets[235,237]. Only known bacterial sequences or sequences closely homologous to known bacterial sequences can be analyzed[235].
Databases are available and they continue to evolve for analyses of 16S rRNA gene sequences[236,239]. Quantitative Insights Into Microbial Ecology (QIIME)[240] and Mothur, a comprehensive software package that integrates several algorithms from pre-existing software[241], are open-source tools that can be used to describe and compare microbial communities. Short 16S rRNA sequences can be organized into OTUs, and a motif-based hierarchical method can analyze massive metagenomic datasets with high accuracy[242]. The Human Microbiome Project provides a catalogue of bacterial species, and it will help define the nature of the intestinal microbiome, the factors that affect its composition, distribution and evolution, and the relationship of the intestinal microflora to human health and disease[236,243,244].
Further advances in the techniques used to reconstruct the composition of the human intestinal microbiome include microarray technology, fingerprinting techniques such as determination of the terminal restriction fragment length polymorphisms, and next-generation sequencing (NGS)[79,245-247]. Microarray hybridization of deoxyribonucleic acid provides a high throughput platform that consists of several thousand probes that can detect nucleic acid sequences simultaneously[246-248]. Unknown microbial sequences and uncharacterized microbial populations are undetected by the microarray techniques, and uncertainties about the existence and importance of undiscovered microbial populations are the major limitations of this method[247,248].
The Human Gut Chip has 4441 probes that includes 2442 probes specific for known microbes and 1919 probes that are explorative for unknown microbes[247]. Probes with overlapping similarities are becoming more sensitive to microbial species that are less abundant[249], and probes with explorative designs are being coupled to probes with microbial specificity in an effort to identify microorganisms with uncharacterized sequences[247,250]. Essential for the design of useful explorative probes is the correct anticipation of genetic variations within the microbial community and the construction of probes with high sensitivity and specificity[249].
Currently, intestinal ecosystems are being studied mainly by 16S rRNA sequencing[236]. This technique is useful in identifying the microbial species that constitute the intestinal microbiome, determining the evolution and transition of the microbial community (phylogeny) and quantitating the microbial diversity[236]. The challenge is to define the functions of the microbiome, and whole genome sequencing (WGS) may provide these insights[236,251]. Array-based NGS can analyze the whole genome, exons, and regions of interest, and it may emerge as the lens by which to understand the function of the microflora[79].
The intestinal microbiome can be manipulated by dietary adjustments[37,90,252], probiotic preparations[197,253,254], supplements of vitamin A and retinoic acid[255,256], antibiotics[169,257], intestinal re-colonization[33,107,109,110], pharmacological agents that decrease intestinal permeability[258-260], molecular interventions that block TLR signaling and the production of pro-inflammatory cytokines[261], molecular interventions (polysaccharide A) that stimulate anti-inflammatory responses[105,262], and short chain fatty acids that modulate signaling pathways that affect gene expression, intestinal barrier integrity, and inflammatory responses[200] (Table 3).
| Treatment Consideration | Nature | Findings |
| Dietary adjustments | Animal protein, saturated fats[90] | Bacteroides, Firmicutes (including Clostridia), and Prevotella favored by different dietary regimens[37,90] |
| High carbohydrate diets[90] | ||
| Low fat high fiber diet[90] | ||
| Probiotic preparations | Bifidobacterium bifidum[253] | Expands Tregs in cell culture[278] |
| Lactobacillus strains[254,263,266] | Prevents diabetes in NOD mice[263] | |
| Lactobacillus rhamnosus[276] | Improves liver tests in rat model[266] | |
| Anaerostipes caccae[277] | Increases tight junction proteins[276] | |
| Consumes lactate and produces butyrate[277] | ||
| Vitamin A and retinoic acid | Retinoic acid supplement[255] | Restores Lactobacilli in lupus model[255] |
| Dietary vitamin A[256] | Regulates cytokines in lupus model[256] | |
| Induces IL-10-producing Tregs[279] | ||
| Antibiotics | Tetracycline, minocycline[257] | Reduces activity in RA[257] |
| Vancomycin, metronidazole[269] | Improves tests and pruritus in PSC[269] | |
| Re-colonization | Bacteroides fragilis[107] | Induces Tregs in colitis model[107,109,110] |
| Fecal transplantation | Clostridia species[109,110] | |
| Intestinal barrier protectors | Gelatin tannate[258-260] | Enhances mucus barrier[258,259] |
| Reduces activity in murine colitis[259] | ||
| Alters composition of microbiota[259] | ||
| Limits inflammatory effects of LPS[260] | ||
| Inhibits IL-8 and TNF-α in LPS cells[260] | ||
| TLR inhibitors | Oligodeoxynucleotides blocking TLR7 signaling[261] | Improves tests and reduces activity in murine model of lupus nephritis[261] |
| Improves autoimmune lung injury[261] | ||
| Molecular interventions | Polysaccharide A[105,262] | Induces IL-10 producing Tregs[105,262] |
| Protects against EAE in mice[105] | ||
| Short chain fatty acids | Acetate, propionate, butyrate[200] | Modulates gut signaling pathways[200] |
| Inhibits histone deacetylases[200,264] | ||
| Regulates gene expression[200] | ||
| Enhances gut integrity[200] |
Antibiotics (tetracycline and minocycline) have reduced disease activity in rheumatoid arthritis, especially in seropositive patients with disease of short duration[257]. Probiotic supplements containing Bifidobacterium bifidum have promoted the expansion of Tregs in cell cultures[253], and probiotics enriched with strains of Lactobacillus alone or in combination with retinoic acid have prevented the development of type 1 diabetes in NOD mice[254,263]. Gelatin tannate has been used in a murine model of acute colitis to protect the mucosal barrier, alter the composition of the microbiota, and decrease inflammatory activity[259]. Gelatin tannate has also been evaluated in LPS-stimulated cell cultures, and it has inhibited the expression of the intercellular adhesion molecule-1 and reduced the production of IL-8 and TNF-α in a dose-dependent fashion[260]. Oligodeoxynucleotides designed to block TLR7 signaling have improved tests and reduced activity in a murine model of lupus nephritis and lung injury[261]; polysaccharide A has induced IL-10 producing Tregs in experimental autoimmune encephalitis[105,262]; and short chain fatty acids have modulated intestinal signaling pathways, inhibited histone deacetylases, regulated gene expression, and increased the integrity of the intestinal barrier[200,264].
Manipulations of the intestinal microbiota have also shown promise in animal models and patients with liver disease. In rats with carbon tetrachloride-induced cirrhosis, antibiotic therapy and probiotic supplements have decreased systemic endotoxin levels and improved liver tests[265], and in rats with ischemic/re-perfusion liver injury, probiotic supplements with Lactobacillus have reduced the production of pro-inflammatory, pro-fibrotic cytokines and improved liver tests[266] (Table 3). The intestinal microbiota have been implicated in the pathogenesis of PSC[267,268], and a small randomized clinical trial has indicated that treatment with vancomycin or metronidazole can improve serum alkaline phosphatase and bilirubin levels and decrease pruritus[269]. Clarification of the role of the intestinal microbiome in autoimmune hepatitis is necessary to direct investigational strategies that would help develop ancillary interventions to improve the outcome of this disease.
Manipulations of the intestinal microbiota, especially with antibiotics, may have adverse consequences which must be defined in animal and human studies and counter-balanced against potential benefits (Table 3). The intestinal microbiome performs important digestive and detoxification functions, produces nutrients and short chain fatty acids that can affect intestinal integrity, defends against invading pathogens, and influences the innate and adaptive immune responses within and outside the intestine[30,270,271]. Dysbiosis associated with antibiotics, especially in early age, may perturb immune tolerance for the microflora and predispose to other immune-mediated diseases (asthma, celiac disease, and type 1 diabetes)[165,166,175,272]. Antibiotic manipulations may also favor the emergence of drug-resistant pathogenic or immunogenic microflora[169]. The optimal nature and duration of the manipulations that might impact on the intestinal microbiome are uncertain, and the durability of the responses are unclear. Much work needs to be done to establish microbiome manipulation as a way forward in autoimmune hepatitis, but observations already made in diverse systemic immune-mediated diseases and the unmet needs in the management of autoimmune hepatitis justify rigorous evaluation of this possibility.
Dysbiosis has already been described in experimental and human autoimmune hepatitis[69,70]; antibodies reactive to antigens homologous to bacterial antigens (atypical pANCA) have been commonly present in patients with the disease[53,65]; and increased permeability of the intestinal mucosal barrier has been demonstrated[70]. The intestinal microbiome is an available source of immune stimulatory antigens, products, and immune cells that already have been implicated in multiple systemic immune-mediated (rheumatoid arthritis, diabetes, multiple sclerosis, inflammatory bowel disease)[35,38,40-43] and chronic liver diseases (NASH, PBC)[51,54]. The role of the intestinal microbiome in the occurrence and behavior of autoimmune hepatitis warrants rigorous evaluation.
The sequencing of the 16S rRNA gene can be used to characterize the intestinal microbial population and determine disease-specific dysbioses[236]. Microarrays comprised of thousands of microbial probes can be applied to enhance comprehension of the “pathological” components of the intestinal microbiome[247,249]. WGS and NGS of genomic regions specific for the disease can define the disease-related metagenome[79], and associations between gut-derived micro-organisms and the immune-mediated mechanisms of autoimmune hepatitis can then be evaluated in experimental models.
Adjunctive forms of therapy may emerge to complement current immunosuppressive regimens. Individual bacterial species within the microbiome may be manipulated by diet, designer probiotics, re-colonization methods, antibiotics, selected vitamin supplements, and pharmacological agents that decrease intestinal permeability[236]. Molecular interventions that block TLR signaling or modulate signaling pathways may also evolve to reduce pro-inflammatory cytokine production, limit unfavorable gene expression, and strengthen the integrity of the intestinal barrier[200,261].
Major obstacles to the performance of these studies are the limited number of publicly available tools to analyze the microbial metagenome, especially in translational settings[239], the multiplicity of environmental factors (diet, antibiotics, sanitation) that can affect variations of the microbiome among communities and between individuals[273,274], and uncertainties about the roles of luminal and mucosal microbiota in directing the immune response[81,275].
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: United States
Peer-review report classification
Grade A (Excellent): A, A
Grade B (Very good): 0
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Boros M, Huerta-Franco M S- Editor: Qi Y L- Editor: A E- Editor: Zhang FF
| 1. | Manns MP, Czaja AJ, Gorham JD, Krawitt EL, Mieli-Vergani G, Vergani D, Vierling JM. Diagnosis and management of autoimmune hepatitis. Hepatology. 2010;51:2193-2213. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1039] [Cited by in RCA: 1010] [Article Influence: 67.3] [Reference Citation Analysis (0)] |
| 2. | Czaja AJ. Transitioning from Idiopathic to Explainable Autoimmune Hepatitis. Dig Dis Sci. 2015;60:2881-2900. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 42] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 3. | Boberg KM, Aadland E, Jahnsen J, Raknerud N, Stiris M, Bell H. Incidence and prevalence of primary biliary cirrhosis, primary sclerosing cholangitis, and autoimmune hepatitis in a Norwegian population. Scand J Gastroenterol. 1998;33:99-103. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 362] [Cited by in RCA: 330] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 4. | Lee YM, Teo EK, Ng TM, Khor C, Fock KM. Autoimmune hepatitis in Singapore: a rare syndrome affecting middle-aged women. J Gastroenterol Hepatol. 2001;16:1384-1389. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 47] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 5. | Hurlburt KJ, McMahon BJ, Deubner H, Hsu-Trawinski B, Williams JL, Kowdley KV. Prevalence of autoimmune liver disease in Alaska Natives. Am J Gastroenterol. 2002;97:2402-2407. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 174] [Cited by in RCA: 172] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 6. | Primo J, Merino C, Fernández J, Molés JR, Llorca P, Hinojosa J. [Incidence and prevalence of autoimmune hepatitis in the area of the Hospital de Sagunto (Spain)]. Gastroenterol Hepatol. 2004;27:239-243. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 40] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 7. | Ngu JH, Bechly K, Chapman BA, Burt MJ, Barclay ML, Gearry RB, Stedman CA. Population-based epidemiology study of autoimmune hepatitis: a disease of older women? J Gastroenterol Hepatol. 2010;25:1681-1686. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 123] [Cited by in RCA: 135] [Article Influence: 9.0] [Reference Citation Analysis (1)] |
| 8. | Delgado JS, Vodonos A, Malnick S, Kriger O, Wilkof-Segev R, Delgado B, Novack V, Rosenthal A, Menachem Y, Melzer E. Autoimmune hepatitis in southern Israel: a 15-year multicenter study. J Dig Dis. 2013;14:611-618. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 42] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 9. | Werner M, Prytz H, Ohlsson B, Almer S, Björnsson E, Bergquist A, Wallerstedt S, Sandberg-Gertzén H, Hultcrantz R, Sangfelt P. Epidemiology and the initial presentation of autoimmune hepatitis in Sweden: a nationwide study. Scand J Gastroenterol. 2008;43:1232-1240. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 157] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 10. | van Gerven NM, Verwer BJ, Witte BI, van Erpecum KJ, van Buuren HR, Maijers I, Visscher AP, Verschuren EC, van Hoek B, Coenraad MJ. Epidemiology and clinical characteristics of autoimmune hepatitis in the Netherlands. Scand J Gastroenterol. 2014;49:1245-1254. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 113] [Cited by in RCA: 125] [Article Influence: 11.4] [Reference Citation Analysis (0)] |
| 11. | Grønbæk L, Vilstrup H, Jepsen P. Autoimmune hepatitis in Denmark: incidence, prevalence, prognosis, and causes of death. A nationwide registry-based cohort study. J Hepatol. 2014;60:612-617. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 246] [Article Influence: 22.4] [Reference Citation Analysis (0)] |
| 12. | Deneau M, Jensen MK, Holmen J, Williams MS, Book LS, Guthery SL. Primary sclerosing cholangitis, autoimmune hepatitis, and overlap in Utah children: epidemiology and natural history. Hepatology. 2013;58:1392-1400. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 161] [Cited by in RCA: 154] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 13. | Czaja AJ. Genetic factors affecting the occurrence, clinical phenotype, and outcome of autoimmune hepatitis. Clin Gastroenterol Hepatol. 2008;6:379-388. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 79] [Cited by in RCA: 70] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 14. | Czaja AJ. Autoimmune hepatitis in diverse ethnic populations and geographical regions. Expert Rev Gastroenterol Hepatol. 2013;7:365-385. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 48] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 15. | Strettell MD, Donaldson PT, Thomson LJ, Santrach PJ, Moore SB, Czaja AJ, Williams R. Allelic basis for HLA-encoded susceptibility to type 1 autoimmune hepatitis. Gastroenterology. 1997;112:2028-2035. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 206] [Cited by in RCA: 191] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 16. | Djilali-Saiah I, Fakhfakh A, Louafi H, Caillat-Zucman S, Debray D, Alvarez F. HLA class II influences humoral autoimmunity in patients with type 2 autoimmune hepatitis. J Hepatol. 2006;45:844-850. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 68] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 17. | van Gerven NM, de Boer YS, Zwiers A, Verwer BJ, Drenth JP, van Hoek B, van Erpecum KJ, Beuers U, van Buuren HR, den Ouden JW. HLA-DRB1*03: 01 and HLA-DRB1*04: 01 modify the presentation and outcome in autoimmune hepatitis type-1. Genes Immun. 2015;16:247-252. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 81] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 18. | Cookson S, Constantini PK, Clare M, Underhill JA, Bernal W, Czaja AJ, Donaldson PT. Frequency and nature of cytokine gene polymorphisms in type 1 autoimmune hepatitis. Hepatology. 1999;30:851-856. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 127] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 19. | Vogel A, Strassburg CP, Manns MP. Genetic association of vitamin D receptor polymorphisms with primary biliary cirrhosis and autoimmune hepatitis. Hepatology. 2002;35:126-131. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 191] [Cited by in RCA: 181] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 20. | Paladino N, Flores AC, Fainboim H, Schroder T, Cuarterolo M, Lezama C, Ballerga EG, Levi D, Tanno H, Costanzo G. The most severe forms of type I autoimmune hepatitis are associated with genetically determined levels of TGF-beta1. Clin Immunol. 2010;134:305-312. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 22] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 21. | Migita K, Nakamura M, Abiru S, Jiuchi Y, Nagaoka S, Komori A, Hashimoto S, Bekki S, Yamasaki K, Komatsu T. Association of STAT4 polymorphisms with susceptibility to type-1 autoimmune hepatitis in the Japanese population. PLoS One. 2013;8:e71382. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 22] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 22. | de Boer YS, van Gerven NM, Zwiers A, Verwer BJ, van Hoek B, van Erpecum KJ, Beuers U, van Buuren HR, Drenth JP, den Ouden JW. Genome-wide association study identifies variants associated with autoimmune hepatitis type 1. Gastroenterology. 2014;147:443-452.e5. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 219] [Cited by in RCA: 225] [Article Influence: 20.5] [Reference Citation Analysis (0)] |
| 23. | Manns MP, Griffin KJ, Sullivan KF, Johnson EF. LKM-1 autoantibodies recognize a short linear sequence in P450IID6, a cytochrome P-450 monooxygenase. J Clin Invest. 1991;88:1370-1378. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 346] [Cited by in RCA: 304] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 24. | Lapierre P, Hajoui O, Homberg JC, Alvarez F. Formiminotransferase cyclodeaminase is an organ-specific autoantigen recognized by sera of patients with autoimmune hepatitis. Gastroenterology. 1999;116:643-649. [PubMed] |
| 25. | Muratori L, Sztul E, Muratori P, Gao Y, Ripalti A, Ponti C, Lenzi M, Landini MP, Bianchi FB. Distinct epitopes on formiminotransferase cyclodeaminase induce autoimmune liver cytosol antibody type 1. Hepatology. 2001;34:494-501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 42] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 26. | Ma Y, Thomas MG, Okamoto M, Bogdanos DP, Nagl S, Kerkar N, Lopes AR, Muratori L, Lenzi M, Bianchi FB. Key residues of a major cytochrome P4502D6 epitope are located on the surface of the molecule. J Immunol. 2002;169:277-285. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 55] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 27. | Kerkar N, Choudhuri K, Ma Y, Mahmoud A, Bogdanos DP, Muratori L, Bianchi F, Williams R, Mieli-Vergani G, Vergani D. Cytochrome P4502D6(193-212): a new immunodominant epitope and target of virus/self cross-reactivity in liver kidney microsomal autoantibody type 1-positive liver disease. J Immunol. 2003;170:1481-1489. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 157] [Cited by in RCA: 139] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 28. | Czaja AJ. Autoimmune hepatitis. Part A: pathogenesis. Expert Rev Gastroenterol Hepatol. 2007;1:113-128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 58] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 29. | Montano-Loza AJ, Czaja AJ. Cell mediators of autoimmune hepatitis and their therapeutic implications. Dig Dis Sci. 2015;60:1528-1542. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 30] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 30. | Wu HJ, Wu E. The role of gut microbiota in immune homeostasis and autoimmunity. Gut Microbes. 2012;3:4-14. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 622] [Cited by in RCA: 902] [Article Influence: 69.4] [Reference Citation Analysis (0)] |
| 31. | Geuking MB, Köller Y, Rupp S, McCoy KD. The interplay between the gut microbiota and the immune system. Gut Microbes. 2014;5:411-418. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 134] [Cited by in RCA: 148] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 32. | Van Praet JT, Donovan E, Vanassche I, Drennan MB, Windels F, Dendooven A, Allais L, Cuvelier CA, van de Loo F, Norris PS. Commensal microbiota influence systemic autoimmune responses. EMBO J. 2015;34:466-474. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 83] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 33. | Sánchez B, Hevia A, González S, Margolles A. Interaction of Intestinal Microorganisms with the Human Host in the Framework of Autoimmune Diseases. Front Immunol. 2015;6:594. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 26] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 34. | Ignacio A, Morales CI, Câmara NO, Almeida RR. Innate Sensing of the Gut Microbiota: Modulation of Inflammatory and Autoimmune Diseases. Front Immunol. 2016;7:54. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 52] [Cited by in RCA: 60] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 35. | Wen L, Ley RE, Volchkov PY, Stranges PB, Avanesyan L, Stonebraker AC, Hu C, Wong FS, Szot GL, Bluestone JA. Innate immunity and intestinal microbiota in the development of Type 1 diabetes. Nature. 2008;455:1109-1113. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1623] [Cited by in RCA: 1498] [Article Influence: 88.1] [Reference Citation Analysis (1)] |
| 36. | Li Y, Liu Y, Chu CQ. Th17 Cells in Type 1 Diabetes: Role in the Pathogenesis and Regulation by Gut Microbiome. Mediators Inflamm. 2015;2015:638470. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 42] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 37. | Mejía-León ME, Barca AM. Diet, Microbiota and Immune System in Type 1 Diabetes Development and Evolution. Nutrients. 2015;7:9171-9184. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 70] [Cited by in RCA: 71] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 38. | Abdollahi-Roodsaz S, Joosten LA, Koenders MI, Devesa I, Roelofs MF, Radstake TR, Heuvelmans-Jacobs M, Akira S, Nicklin MJ, Ribeiro-Dias F. Stimulation of TLR2 and TLR4 differentially skews the balance of T cells in a mouse model of arthritis. J Clin Invest. 2008;118:205-216. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 365] [Cited by in RCA: 410] [Article Influence: 24.1] [Reference Citation Analysis (0)] |
| 39. | Wu HJ, Ivanov II, Darce J, Hattori K, Shima T, Umesaki Y, Littman DR, Benoist C, Mathis D. Gut-residing segmented filamentous bacteria drive autoimmune arthritis via T helper 17 cells. Immunity. 2010;32:815-827. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1330] [Cited by in RCA: 1239] [Article Influence: 82.6] [Reference Citation Analysis (0)] |
| 40. | Abdollahi-Roodsaz S, Koenders MI, Walgreen B, Bolscher J, Helsen MM, van den Bersselaar LA, van Lent PL, van de Loo FA, van den Berg WB. Toll-like receptor 2 controls acute immune complex-driven arthritis in mice by regulating the inhibitory Fcγ receptor IIB. Arthritis Rheum. 2013;65:2583-2593. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 12] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 41. | Rogier R, Koenders MI, Abdollahi-Roodsaz S. Toll-like receptor mediated modulation of T cell response by commensal intestinal microbiota as a trigger for autoimmune arthritis. J Immunol Res. 2015;2015:527696. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 50] [Cited by in RCA: 71] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 42. | Miyake S, Kim S, Suda W, Oshima K, Nakamura M, Matsuoka T, Chihara N, Tomita A, Sato W, Kim SW. Dysbiosis in the Gut Microbiota of Patients with Multiple Sclerosis, with a Striking Depletion of Species Belonging to Clostridia XIVa and IV Clusters. PLoS One. 2015;10:e0137429. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 463] [Cited by in RCA: 570] [Article Influence: 57.0] [Reference Citation Analysis (0)] |
| 43. | Sokol H, Seksik P, Rigottier-Gois L, Lay C, Lepage P, Podglajen I, Marteau P, Doré J. Specificities of the fecal microbiota in inflammatory bowel disease. Inflamm Bowel Dis. 2006;12:106-111. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 334] [Cited by in RCA: 323] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 44. | Garrett WS, Gallini CA, Yatsunenko T, Michaud M, DuBois A, Delaney ML, Punit S, Karlsson M, Bry L, Glickman JN. Enterobacteriaceae act in concert with the gut microbiota to induce spontaneous and maternally transmitted colitis. Cell Host Microbe. 2010;8:292-300. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 687] [Cited by in RCA: 643] [Article Influence: 42.9] [Reference Citation Analysis (0)] |
| 45. | Elinav E, Strowig T, Kau AL, Henao-Mejia J, Thaiss CA, Booth CJ, Peaper DR, Bertin J, Eisenbarth SC, Gordon JI. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell. 2011;145:745-757. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1659] [Cited by in RCA: 1578] [Article Influence: 112.7] [Reference Citation Analysis (0)] |
| 46. | Seki E, Schnabl B. Role of innate immunity and the microbiota in liver fibrosis: crosstalk between the liver and gut. J Physiol. 2012;590:447-458. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 270] [Cited by in RCA: 337] [Article Influence: 24.1] [Reference Citation Analysis (0)] |
| 47. | Henao-Mejia J, Elinav E, Thaiss CA, Licona-Limon P, Flavell RA. Role of the intestinal microbiome in liver disease. J Autoimmun. 2013;46:66-73. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 116] [Cited by in RCA: 129] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 48. | Miyake Y, Yamamoto K. Role of gut microbiota in liver diseases. Hepatol Res. 2013;43:139-146. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 56] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 49. | Miura K, Kodama Y, Inokuchi S, Schnabl B, Aoyama T, Ohnishi H, Olefsky JM, Brenner DA, Seki E. Toll-like receptor 9 promotes steatohepatitis by induction of interleukin-1beta in mice. Gastroenterology. 2010;139:323-324.e7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 527] [Cited by in RCA: 623] [Article Influence: 41.5] [Reference Citation Analysis (0)] |
| 50. | Csak T, Ganz M, Pespisa J, Kodys K, Dolganiuc A, Szabo G. Fatty acid and endotoxin activate inflammasomes in mouse hepatocytes that release danger signals to stimulate immune cells. Hepatology. 2011;54:133-144. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 442] [Cited by in RCA: 533] [Article Influence: 38.1] [Reference Citation Analysis (0)] |
| 51. | Henao-Mejia J, Elinav E, Jin C, Hao L, Mehal WZ, Strowig T, Thaiss CA, Kau AL, Eisenbarth SC, Jurczak MJ. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature. 2012;482:179-185. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1620] [Cited by in RCA: 1880] [Article Influence: 144.6] [Reference Citation Analysis (0)] |
| 52. | Karrar A, Broomé U, Södergren T, Jaksch M, Bergquist A, Björnstedt M, Sumitran-Holgersson S. Biliary epithelial cell antibodies link adaptive and innate immune responses in primary sclerosing cholangitis. Gastroenterology. 2007;132:1504-1514. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 92] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 53. | Terjung B, Söhne J, Lechtenberg B, Gottwein J, Muennich M, Herzog V, Mähler M, Sauerbruch T, Spengler U. p-ANCAs in autoimmune liver disorders recognise human beta-tubulin isotype 5 and cross-react with microbial protein FtsZ. Gut. 2010;59:808-816. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 107] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 54. | Mueller T, Beutler C, Picó AH, Shibolet O, Pratt DS, Pascher A, Neuhaus P, Wiedenmann B, Berg T, Podolsky DK. Enhanced innate immune responsiveness and intolerance to intestinal endotoxins in human biliary epithelial cells contributes to chronic cholangitis. Liver Int. 2011;31:1574-1588. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 63] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 55. | Tabibian JH, O’Hara SP, Trussoni CE, Tietz PS, Splinter PL, Mounajjed T, Hagey LR, LaRusso NF. Absence of the intestinal microbiota exacerbates hepatobiliary disease in a murine model of primary sclerosing cholangitis. Hepatology. 2016;63:185-196. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 182] [Cited by in RCA: 165] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 56. | Hopf U, Möller B, Stemerowicz R, Lobeck H, Rodloff A, Freudenberg M, Galanos C, Huhn D. Relation between Escherichia coli R(rough)-forms in gut, lipid A in liver, and primary biliary cirrhosis. Lancet. 1989;2:1419-1422. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 103] [Cited by in RCA: 89] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 57. | Bogdanos DP, Baum H, Grasso A, Okamoto M, Butler P, Ma Y, Rigopoulou E, Montalto P, Davies ET, Burroughs AK. Microbial mimics are major targets of crossreactivity with human pyruvate dehydrogenase in primary biliary cirrhosis. J Hepatol. 2004;40:31-39. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 103] [Cited by in RCA: 108] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 58. | Wang AP, Migita K, Ito M, Takii Y, Daikoku M, Yokoyama T, Komori A, Nakamura M, Yatsuhashi H, Ishibashi H. Hepatic expression of toll-like receptor 4 in primary biliary cirrhosis. J Autoimmun. 2005;25:85-91. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 75] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 59. | Mao TK, Lian ZX, Selmi C, Ichiki Y, Ashwood P, Ansari AA, Coppel RL, Shimoda S, Ishibashi H, Gershwin ME. Altered monocyte responses to defined TLR ligands in patients with primary biliary cirrhosis. Hepatology. 2005;42:802-808. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 137] [Cited by in RCA: 143] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 60. | Kikuchi K, Lian ZX, Yang GX, Ansari AA, Ikehara S, Kaplan M, Miyakawa H, Coppel RL, Gershwin ME. Bacterial CpG induces hyper-IgM production in CD27(+) memory B cells in primary biliary cirrhosis. Gastroenterology. 2005;128:304-312. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 116] [Cited by in RCA: 108] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 61. | Honda Y, Yamagiwa S, Matsuda Y, Takamura M, Ichida T, Aoyagi Y. Altered expression of TLR homolog RP105 on monocytes hypersensitive to LPS in patients with primary biliary cirrhosis. J Hepatol. 2007;47:404-411. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 28] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 62. | Shimoda S, Harada K, Niiro H, Shirabe K, Taketomi A, Maehara Y, Tsuneyama K, Nakanuma Y, Leung P, Ansari AA. Interaction between Toll-like receptors and natural killer cells in the destruction of bile ducts in primary biliary cirrhosis. Hepatology. 2011;53:1270-1281. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 108] [Cited by in RCA: 103] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 63. | Czaja AJ. Frequency and nature of the variant syndromes of autoimmune liver disease. Hepatology. 1998;28:360-365. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 237] [Cited by in RCA: 211] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 64. | Czaja AJ. The overlap syndromes of autoimmune hepatitis. Dig Dis Sci. 2013;58:326-343. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 26] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 65. | Targan SR, Landers C, Vidrich A, Czaja AJ. High-titer antineutrophil cytoplasmic antibodies in type-1 autoimmune hepatitis. Gastroenterology. 1995;108:1159-1166. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 129] [Cited by in RCA: 117] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 66. | Bansi D, Chapman R, Fleming K. Antineutrophil cytoplasmic antibodies in chronic liver diseases: prevalence, titre, specificity and IgG subclass. J Hepatol. 1996;24:581-586. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 45] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 67. | Zauli D, Ghetti S, Grassi A, Descovich C, Cassani F, Ballardini G, Muratori L, Bianchi FB. Anti-neutrophil cytoplasmic antibodies in type 1 and 2 autoimmune hepatitis. Hepatology. 1997;25:1105-1107. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 92] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 68. | Roozendaal C, de Jong MA, van den Berg AP, van Wijk RT, Limburg PC, Kallenberg CG. Clinical significance of anti-neutrophil cytoplasmic antibodies (ANCA) in autoimmune liver diseases. J Hepatol. 2000;32:734-741. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 66] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 69. | Yuksel M, Wang Y, Tai N, Peng J, Guo J, Beland K, Lapierre P, David C, Alvarez F, Colle I. A novel “humanized mouse” model for autoimmune hepatitis and the association of gut microbiota with liver inflammation. Hepatology. 2015;62:1536-1550. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 92] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 70. | Lin R, Zhou L, Zhang J, Wang B. Abnormal intestinal permeability and microbiota in patients with autoimmune hepatitis. Int J Clin Exp Pathol. 2015;8:5153-5160. [PubMed] |
| 71. | Zoetendal EG, Vaughan EE, de Vos WM. A microbial world within us. Mol Microbiol. 2006;59:1639-1650. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 275] [Cited by in RCA: 256] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 72. | Booijink CC, Zoetendal EG, Kleerebezem M, de Vos WM. Microbial communities in the human small intestine: coupling diversity to metagenomics. Future Microbiol. 2007;2:285-295. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 97] [Cited by in RCA: 101] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 73. | Zoetendal EG, Rajilic-Stojanovic M, de Vos WM. High-throughput diversity and functionality analysis of the gastrointestinal tract microbiota. Gut. 2008;57:1605-1615. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 454] [Cited by in RCA: 456] [Article Influence: 26.8] [Reference Citation Analysis (0)] |
| 74. | Lozupone CA, Stombaugh JI, Gordon JI, Jansson JK, Knight R. Diversity, stability and resilience of the human gut microbiota. Nature. 2012;489:220-230. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3051] [Cited by in RCA: 3647] [Article Influence: 280.5] [Reference Citation Analysis (0)] |
| 75. | Hollister EB, Gao C, Versalovic J. Compositional and functional features of the gastrointestinal microbiome and their effects on human health. Gastroenterology. 2014;146:1449-1458. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 300] [Cited by in RCA: 326] [Article Influence: 29.6] [Reference Citation Analysis (0)] |
| 76. | Rajilić-Stojanović M, Smidt H, de Vos WM. Diversity of the human gastrointestinal tract microbiota revisited. Environ Microbiol. 2007;9:2125-2136. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 395] [Cited by in RCA: 382] [Article Influence: 21.2] [Reference Citation Analysis (0)] |
| 77. | Bäckhed F, Ley RE, Sonnenburg JL, Peterson DA, Gordon JI. Host-bacterial mutualism in the human intestine. Science. 2005;307:1915-1920. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3394] [Cited by in RCA: 3545] [Article Influence: 177.3] [Reference Citation Analysis (5)] |
| 78. | Gill SR, Pop M, Deboy RT, Eckburg PB, Turnbaugh PJ, Samuel BS, Gordon JI, Relman DA, Fraser-Liggett CM, Nelson KE. Metagenomic analysis of the human distal gut microbiome. Science. 2006;312:1355-1359. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3709] [Cited by in RCA: 3192] [Article Influence: 168.0] [Reference Citation Analysis (0)] |
| 79. | Ma Y, Shi N, Li M, Chen F, Niu H. Applications of Next-generation Sequencing in Systemic Autoimmune Diseases. Genomics Proteomics Bioinformatics. 2015;13:242-249. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 25] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 80. | Tap J, Mondot S, Levenez F, Pelletier E, Caron C, Furet JP, Ugarte E, Muñoz-Tamayo R, Paslier DL, Nalin R. Towards the human intestinal microbiota phylogenetic core. Environ Microbiol. 2009;11:2574-2584. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 627] [Cited by in RCA: 635] [Article Influence: 39.7] [Reference Citation Analysis (0)] |
| 81. | Ringel Y, Maharshak N, Ringel-Kulka T, Wolber EA, Sartor RB, Carroll IM. High throughput sequencing reveals distinct microbial populations within the mucosal and luminal niches in healthy individuals. Gut Microbes. 2015;6:173-181. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 157] [Article Influence: 15.7] [Reference Citation Analysis (0)] |
| 82. | Mariat D, Firmesse O, Levenez F, Guimarăes V, Sokol H, Doré J, Corthier G, Furet JP. The Firmicutes/Bacteroidetes ratio of the human microbiota changes with age. BMC Microbiol. 2009;9:123. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 997] [Cited by in RCA: 1246] [Article Influence: 77.9] [Reference Citation Analysis (0)] |
| 83. | Claesson MJ, Cusack S, O’Sullivan O, Greene-Diniz R, de Weerd H, Flannery E, Marchesi JR, Falush D, Dinan T, Fitzgerald G. Composition, variability, and temporal stability of the intestinal microbiota of the elderly. Proc Natl Acad Sci USA. 2011;108 Suppl 1:4586-4591. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1162] [Cited by in RCA: 1244] [Article Influence: 88.9] [Reference Citation Analysis (2)] |
| 84. | Jalanka-Tuovinen J, Salonen A, Nikkilä J, Immonen O, Kekkonen R, Lahti L, Palva A, de Vos WM. Intestinal microbiota in healthy adults: temporal analysis reveals individual and common core and relation to intestinal symptoms. PLoS One. 2011;6:e23035. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 237] [Cited by in RCA: 259] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 85. | Power SE, O’Toole PW, Stanton C, Ross RP, Fitzgerald GF. Intestinal microbiota, diet and health. Br J Nutr. 2014;111:387-402. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 286] [Cited by in RCA: 329] [Article Influence: 27.4] [Reference Citation Analysis (0)] |
| 86. | Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486:207-214. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9292] [Cited by in RCA: 8068] [Article Influence: 620.6] [Reference Citation Analysis (2)] |
| 87. | Lin A, Bik EM, Costello EK, Dethlefsen L, Haque R, Relman DA, Singh U. Distinct distal gut microbiome diversity and composition in healthy children from Bangladesh and the United States. PLoS One. 2013;8:e53838. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 220] [Cited by in RCA: 239] [Article Influence: 19.9] [Reference Citation Analysis (0)] |
| 88. | Chong CW, Ahmad AF, Lim YA, Teh CS, Yap IK, Lee SC, Chin YT, Loke P, Chua KH. Effect of ethnicity and socioeconomic variation to the gut microbiota composition among pre-adolescent in Malaysia. Sci Rep. 2015;5:13338. [PubMed] |
| 89. | Zhang J, Guo Z, Xue Z, Sun Z, Zhang M, Wang L, Wang G, Wang F, Xu J, Cao H. A phylo-functional core of gut microbiota in healthy young Chinese cohorts across lifestyles, geography and ethnicities. ISME J. 2015;9:1979-1990. [PubMed] |
| 90. | Wu GD, Chen J, Hoffmann C, Bittinger K, Chen YY, Keilbaugh SA, Bewtra M, Knights D, Walters WA, Knight R. Linking long-term dietary patterns with gut microbial enterotypes. Science. 2011;334:105-108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4098] [Cited by in RCA: 4544] [Article Influence: 324.6] [Reference Citation Analysis (1)] |
| 91. | De Filippo C, Cavalieri D, Di Paola M, Ramazzotti M, Poullet JB, Massart S, Collini S, Pieraccini G, Lionetti P. Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci USA. 2010;107:14691-14696. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3584] [Cited by in RCA: 4033] [Article Influence: 268.9] [Reference Citation Analysis (0)] |
| 92. | Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, Contreras M, Magris M, Hidalgo G, Baldassano RN, Anokhin AP. Human gut microbiome viewed across age and geography. Nature. 2012;486:222-227. [PubMed] |
| 93. | Rampelli S, Schnorr SL, Consolandi C, Turroni S, Severgnini M, Peano C, Brigidi P, Crittenden AN, Henry AG, Candela M. Metagenome Sequencing of the Hadza Hunter-Gatherer Gut Microbiota. Curr Biol. 2015;25:1682-1693. [PubMed] |
| 94. | Segata N. Gut Microbiome: Westernization and the Disappearance of Intestinal Diversity. Curr Biol. 2015;25:R611-R613. [PubMed] |
| 95. | Miller GE, Engen PA, Gillevet PM, Shaikh M, Sikaroodi M, Forsyth CB, Mutlu E, Keshavarzian A. Lower Neighborhood Socioeconomic Status Associated with Reduced Diversity of the Colonic Microbiota in Healthy Adults. PLoS One. 2016;11:e0148952. [PubMed] |
| 96. | Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010;464:59-65. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9101] [Cited by in RCA: 7840] [Article Influence: 522.7] [Reference Citation Analysis (4)] |
| 97. | Turnbaugh PJ, Hamady M, Yatsunenko T, Cantarel BL, Duncan A, Ley RE, Sogin ML, Jones WJ, Roe BA, Affourtit JP. A core gut microbiome in obese and lean twins. Nature. 2009;457:480-484. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6397] [Cited by in RCA: 5667] [Article Influence: 354.2] [Reference Citation Analysis (1)] |
| 98. | Huse SM, Ye Y, Zhou Y, Fodor AA. A core human microbiome as viewed through 16S rRNA sequence clusters. PLoS One. 2012;7:e34242. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 367] [Cited by in RCA: 388] [Article Influence: 29.8] [Reference Citation Analysis (0)] |
| 99. | Zupancic ML, Cantarel BL, Liu Z, Drabek EF, Ryan KA, Cirimotich S, Jones C, Knight R, Walters WA, Knights D. Analysis of the gut microbiota in the old order Amish and its relation to the metabolic syndrome. PLoS One. 2012;7:e43052. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 177] [Cited by in RCA: 157] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 100. | Turnbaugh PJ, Ridaura VK, Faith JJ, Rey FE, Knight R, Gordon JI. The effect of diet on the human gut microbiome: a metagenomic analysis in humanized gnotobiotic mice. Sci Transl Med. 2009;1:6ra14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2376] [Cited by in RCA: 2165] [Article Influence: 135.3] [Reference Citation Analysis (0)] |
| 101. | Belkaid Y, Hand TW. Role of the microbiota in immunity and inflammation. Cell. 2014;157:121-141. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2488] [Cited by in RCA: 3430] [Article Influence: 311.8] [Reference Citation Analysis (1)] |
| 102. | Macpherson AJ, Harris NL. Interactions between commensal intestinal bacteria and the immune system. Nat Rev Immunol. 2004;4:478-485. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1099] [Cited by in RCA: 1142] [Article Influence: 54.4] [Reference Citation Analysis (0)] |
| 103. | Macpherson AJ, Geuking MB, McCoy KD. Immune responses that adapt the intestinal mucosa to commensal intestinal bacteria. Immunology. 2005;115:153-162. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 207] [Cited by in RCA: 178] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 104. | Mazmanian SK, Liu CH, Tzianabos AO, Kasper DL. An immunomodulatory molecule of symbiotic bacteria directs maturation of the host immune system. Cell. 2005;122:107-118. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1932] [Cited by in RCA: 2094] [Article Influence: 104.7] [Reference Citation Analysis (0)] |
| 105. | Ochoa-Repáraz J, Mielcarz DW, Wang Y, Begum-Haque S, Dasgupta S, Kasper DL, Kasper LH. A polysaccharide from the human commensal Bacteroides fragilis protects against CNS demyelinating disease. Mucosal Immunol. 2010;3:487-495. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 364] [Cited by in RCA: 401] [Article Influence: 26.7] [Reference Citation Analysis (0)] |
| 106. | Ochoa-Repáraz J, Mielcarz DW, Ditrio LE, Burroughs AR, Begum-Haque S, Dasgupta S, Kasper DL, Kasper LH. Central nervous system demyelinating disease protection by the human commensal Bacteroides fragilis depends on polysaccharide A expression. J Immunol. 2010;185:4101-4108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 267] [Cited by in RCA: 298] [Article Influence: 19.9] [Reference Citation Analysis (0)] |
| 107. | Round JL, Mazmanian SK. Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc Natl Acad Sci USA. 2010;107:12204-12209. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1484] [Cited by in RCA: 1676] [Article Influence: 111.7] [Reference Citation Analysis (0)] |
| 108. | Round JL, Lee SM, Li J, Tran G, Jabri B, Chatila TA, Mazmanian SK. The Toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science. 2011;332:974-977. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1311] [Cited by in RCA: 1212] [Article Influence: 86.6] [Reference Citation Analysis (0)] |
| 109. | Atarashi K, Tanoue T, Shima T, Imaoka A, Kuwahara T, Momose Y, Cheng G, Yamasaki S, Saito T, Ohba Y. Induction of colonic regulatory T cells by indigenous Clostridium species. Science. 2011;331:337-341. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2568] [Cited by in RCA: 2880] [Article Influence: 192.0] [Reference Citation Analysis (0)] |
| 110. | Atarashi K, Tanoue T, Oshima K, Suda W, Nagano Y, Nishikawa H, Fukuda S, Saito T, Narushima S, Hase K. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature. 2013;500:232-236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1875] [Cited by in RCA: 2140] [Article Influence: 178.3] [Reference Citation Analysis (2)] |
| 111. | Watanabe A, Sohail MA, Gomes DA, Hashmi A, Nagata J, Sutterwala FS, Mahmood S, Jhandier MN, Shi Y, Flavell RA. Inflammasome-mediated regulation of hepatic stellate cells. Am J Physiol Gastrointest Liver Physiol. 2009;296:G1248-G1257. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 166] [Cited by in RCA: 191] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 112. | Strowig T, Henao-Mejia J, Elinav E, Flavell R. Inflammasomes in health and disease. Nature. 2012;481:278-286. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1530] [Cited by in RCA: 1783] [Article Influence: 137.2] [Reference Citation Analysis (1)] |
| 113. | Boaru SG, Borkham-Kamphorst E, Tihaa L, Haas U, Weiskirchen R. Expression analysis of inflammasomes in experimental models of inflammatory and fibrotic liver disease. J Inflamm (Lond). 2012;9:49. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 88] [Cited by in RCA: 116] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 114. | Cerf-Bensussan N, Gaboriau-Routhiau V. The immune system and the gut microbiota: friends or foes? Nat Rev Immunol. 2010;10:735-744. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 453] [Cited by in RCA: 489] [Article Influence: 32.6] [Reference Citation Analysis (0)] |
| 115. | Singh R, Bullard J, Kalra M, Assefa S, Kaul AK, Vonfeldt K, Strom SC, Conrad RS, Sharp HL, Kaul R. Status of bacterial colonization, Toll-like receptor expression and nuclear factor-kappa B activation in normal and diseased human livers. Clin Immunol. 2011;138:41-49. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 42] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 116. | Frank DN, Zhu W, Sartor RB, Li E. Investigating the biological and clinical significance of human dysbioses. Trends Microbiol. 2011;19:427-434. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 137] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 117. | Hänninen A, Salmi M, Simell O, Jalkanen S. Mucosa-associated (beta 7-integrinhigh) lymphocytes accumulate early in the pancreas of NOD mice and show aberrant recirculation behavior. Diabetes. 1996;45:1173-1180. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 65] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 118. | Paronen J, Klemetti P, Kantele JM, Savilahti E, Perheentupa J, Akerblom HK, Vaarala O. Glutamate decarboxylase-reactive peripheral blood lymphocytes from patients with IDDM express gut-specific homing receptor alpha4beta7-integrin. Diabetes. 1997;46:583-588. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 55] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 119. | Kumar H, Kawai T, Akira S. Pathogen recognition by the innate immune system. Int Rev Immunol. 2011;30:16-34. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1380] [Cited by in RCA: 1545] [Article Influence: 110.4] [Reference Citation Analysis (0)] |
| 120. | Yu L, Wang L, Chen S. Endogenous toll-like receptor ligands and their biological significance. J Cell Mol Med. 2010;14:2592-2603. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 351] [Cited by in RCA: 422] [Article Influence: 30.1] [Reference Citation Analysis (0)] |
| 121. | Kuhn KA, Pedraza I, Demoruelle MK. Mucosal immune responses to microbiota in the development of autoimmune disease. Rheum Dis Clin North Am. 2014;40:711-725. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 34] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 122. | Frosali S, Pagliari D, Gambassi G, Landolfi R, Pandolfi F, Cianci R. How the Intricate Interaction among Toll-Like Receptors, Microbiota, and Intestinal Immunity Can Influence Gastrointestinal Pathology. J Immunol Res. 2015;2015:489821. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 126] [Cited by in RCA: 174] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 123. | Takeuchi O, Akira S. Pattern recognition receptors and inflammation. Cell. 2010;140:805-820. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5434] [Cited by in RCA: 6551] [Article Influence: 436.7] [Reference Citation Analysis (0)] |
| 124. | Bonnert TP, Garka KE, Parnet P, Sonoda G, Testa JR, Sims JE. The cloning and characterization of human MyD88: a member of an IL-1 receptor related family. FEBS Lett. 1997;402:81-84. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 108] [Cited by in RCA: 115] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 125. | Yamamoto M, Takeda K. Current views of toll-like receptor signaling pathways. Gastroenterol Res Pract. 2010;2010:240365. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 130] [Cited by in RCA: 156] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 126. | Tseng PH, Matsuzawa A, Zhang W, Mino T, Vignali DA, Karin M. Different modes of ubiquitination of the adaptor TRAF3 selectively activate the expression of type I interferons and proinflammatory cytokines. Nat Immunol. 2010;11:70-75. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 286] [Cited by in RCA: 326] [Article Influence: 20.4] [Reference Citation Analysis (0)] |
| 127. | Pasare C, Medzhitov R. Toll-like receptors and acquired immunity. Semin Immunol. 2004;16:23-26. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 142] [Cited by in RCA: 146] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 128. | Wu J, Meng Z, Jiang M, Zhang E, Trippler M, Broering R, Bucchi A, Krux F, Dittmer U, Yang D. Toll-like receptor-induced innate immune responses in non-parenchymal liver cells are cell type-specific. Immunology. 2010;129:363-374. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 142] [Cited by in RCA: 177] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 129. | Seki E, Tsutsui H, Nakano H, Tsuji N, Hoshino K, Adachi O, Adachi K, Futatsugi S, Kuida K, Takeuchi O. Lipopolysaccharide-induced IL-18 secretion from murine Kupffer cells independently of myeloid differentiation factor 88 that is critically involved in induction of production of IL-12 and IL-1beta. J Immunol. 2001;166:2651-2657. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 178] [Cited by in RCA: 187] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 130. | Seki E, Brenner DA. Toll-like receptors and adaptor molecules in liver disease: update. Hepatology. 2008;48:322-335. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 515] [Cited by in RCA: 552] [Article Influence: 32.5] [Reference Citation Analysis (0)] |
| 131. | Agrawal S, Agrawal A, Doughty B, Gerwitz A, Blenis J, Van Dyke T, Pulendran B. Cutting edge: different Toll-like receptor agonists instruct dendritic cells to induce distinct Th responses via differential modulation of extracellular signal-regulated kinase-mitogen-activated protein kinase and c-Fos. J Immunol. 2003;171:4984-4989. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 572] [Cited by in RCA: 582] [Article Influence: 27.7] [Reference Citation Analysis (0)] |
| 132. | Kattah MG, Wong MT, Yocum MD, Utz PJ. Cytokines secreted in response to Toll-like receptor ligand stimulation modulate differentiation of human Th17 cells. Arthritis Rheum. 2008;58:1619-1629. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 45] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 133. | Dillon S, Agrawal A, Van Dyke T, Landreth G, McCauley L, Koh A, Maliszewski C, Akira S, Pulendran B. A Toll-like receptor 2 ligand stimulates Th2 responses in vivo, via induction of extracellular signal-regulated kinase mitogen-activated protein kinase and c-Fos in dendritic cells. J Immunol. 2004;172:4733-4743. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 361] [Cited by in RCA: 362] [Article Influence: 17.2] [Reference Citation Analysis (0)] |
| 134. | Chow JC, Young DW, Golenbock DT, Christ WJ, Gusovsky F. Toll-like receptor-4 mediates lipopolysaccharide-induced signal transduction. J Biol Chem. 1999;274:10689-10692. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1455] [Cited by in RCA: 1488] [Article Influence: 57.2] [Reference Citation Analysis (0)] |
| 135. | Bauer S, Kirschning CJ, Häcker H, Redecke V, Hausmann S, Akira S, Wagner H, Lipford GB. Human TLR9 confers responsiveness to bacterial DNA via species-specific CpG motif recognition. Proc Natl Acad Sci USA. 2001;98:9237-9242. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1132] [Cited by in RCA: 1153] [Article Influence: 48.0] [Reference Citation Analysis (0)] |
| 136. | Dolganiuc A, Oak S, Kodys K, Golenbock DT, Finberg RW, Kurt-Jones E, Szabo G. Hepatitis C core and nonstructural 3 proteins trigger toll-like receptor 2-mediated pathways and inflammatory activation. Gastroenterology. 2004;127:1513-1524. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 238] [Cited by in RCA: 235] [Article Influence: 11.2] [Reference Citation Analysis (0)] |
| 137. | Wang B, Trippler M, Pei R, Lu M, Broering R, Gerken G, Schlaak JF. Toll-like receptor activated human and murine hepatic stellate cells are potent regulators of hepatitis C virus replication. J Hepatol. 2009;51:1037-1045. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 73] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 138. | Yoon SI, Kurnasov O, Natarajan V, Hong M, Gudkov AV, Osterman AL, Wilson IA. Structural basis of TLR5-flagellin recognition and signaling. Science. 2012;335:859-864. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 369] [Cited by in RCA: 434] [Article Influence: 33.4] [Reference Citation Analysis (0)] |
| 139. | Diebold SS, Kaisho T, Hemmi H, Akira S, Reis e Sousa C. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science. 2004;303:1529-1531. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2469] [Cited by in RCA: 2553] [Article Influence: 121.6] [Reference Citation Analysis (0)] |
| 140. | Caron G, Duluc D, Frémaux I, Jeannin P, David C, Gascan H, Delneste Y. Direct stimulation of human T cells via TLR5 and TLR7/8: flagellin and R-848 up-regulate proliferation and IFN-gamma production by memory CD4+ T cells. J Immunol. 2005;175:1551-1557. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 321] [Cited by in RCA: 341] [Article Influence: 17.1] [Reference Citation Analysis (0)] |
| 141. | Kabelitz D. Expression and function of Toll-like receptors in T lymphocytes. Curr Opin Immunol. 2007;19:39-45. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 272] [Cited by in RCA: 278] [Article Influence: 15.4] [Reference Citation Analysis (0)] |
| 142. | Pasare C, Medzhitov R. Toll pathway-dependent blockade of CD4+CD25+ T cell-mediated suppression by dendritic cells. Science. 2003;299:1033-1036. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1585] [Cited by in RCA: 1578] [Article Influence: 71.7] [Reference Citation Analysis (0)] |
| 143. | Seki E, De Minicis S, Gwak GY, Kluwe J, Inokuchi S, Bursill CA, Llovet JM, Brenner DA, Schwabe RF. CCR1 and CCR5 promote hepatic fibrosis in mice. J Clin Invest. 2009;119:1858-1870. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 244] [Article Influence: 15.3] [Reference Citation Analysis (0)] |
| 144. | Seki E, De Minicis S, Osterreicher CH, Kluwe J, Osawa Y, Brenner DA, Schwabe RF. TLR4 enhances TGF-beta signaling and hepatic fibrosis. Nat Med. 2007;13:1324-1332. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1361] [Cited by in RCA: 1556] [Article Influence: 86.4] [Reference Citation Analysis (1)] |
| 145. | Roderburg C, Urban GW, Bettermann K, Vucur M, Zimmermann H, Schmidt S, Janssen J, Koppe C, Knolle P, Castoldi M. Micro-RNA profiling reveals a role for miR-29 in human and murine liver fibrosis. Hepatology. 2011;53:209-218. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 656] [Cited by in RCA: 661] [Article Influence: 47.2] [Reference Citation Analysis (0)] |
| 146. | Figueroa L, Xiong Y, Song C, Piao W, Vogel SN, Medvedev AE. The Asp299Gly polymorphism alters TLR4 signaling by interfering with recruitment of MyD88 and TRIF. J Immunol. 2012;188:4506-4515. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 105] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 147. | Thuy S, Ladurner R, Volynets V, Wagner S, Strahl S, Königsrainer A, Maier KP, Bischoff SC, Bergheim I. Nonalcoholic fatty liver disease in humans is associated with increased plasma endotoxin and plasminogen activator inhibitor 1 concentrations and with fructose intake. J Nutr. 2008;138:1452-1455. [PubMed] |
| 148. | Gao B, Seki E, Brenner DA, Friedman S, Cohen JI, Nagy L, Szabo G, Zakhari S. Innate immunity in alcoholic liver disease. Am J Physiol Gastrointest Liver Physiol. 2011;300:G516-G525. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 176] [Cited by in RCA: 184] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 149. | Nolan JP, Leibowitz AI. Endotoxin and the liver. III. Modification of acute carbon tetrachloride injury by polymyxin b--an antiendotoxin. Gastroenterology. 1978;75:445-449. [PubMed] |
| 150. | Grinko I, Geerts A, Wisse E. Experimental biliary fibrosis correlates with increased numbers of fat-storing and Kupffer cells, and portal endotoxemia. J Hepatol. 1995;23:449-458. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 51] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 151. | Lin RS, Lee FY, Lee SD, Tsai YT, Lin HC, Lu RH, Hsu WC, Huang CC, Wang SS, Lo KJ. Endotoxemia in patients with chronic liver diseases: relationship to severity of liver diseases, presence of esophageal varices, and hyperdynamic circulation. J Hepatol. 1995;22:165-172. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 258] [Cited by in RCA: 268] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 152. | Chan CC, Hwang SJ, Lee FY, Wang SS, Chang FY, Li CP, Chu CJ, Lu RH, Lee SD. Prognostic value of plasma endotoxin levels in patients with cirrhosis. Scand J Gastroenterol. 1997;32:942-946. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 83] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 153. | Schroder K, Tschopp J. The inflammasomes. Cell. 2010;140:821-832. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3880] [Cited by in RCA: 4547] [Article Influence: 303.1] [Reference Citation Analysis (0)] |
| 154. | Davis BK, Wen H, Ting JP. The inflammasome NLRs in immunity, inflammation, and associated diseases. Annu Rev Immunol. 2011;29:707-735. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1286] [Cited by in RCA: 1282] [Article Influence: 91.6] [Reference Citation Analysis (0)] |
| 155. | Franchi L, Kamada N, Nakamura Y, Burberry A, Kuffa P, Suzuki S, Shaw MH, Kim YG, Núñez G. NLRC4-driven production of IL-1β discriminates between pathogenic and commensal bacteria and promotes host intestinal defense. Nat Immunol. 2012;13:449-456. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 285] [Cited by in RCA: 317] [Article Influence: 24.4] [Reference Citation Analysis (0)] |
| 156. | Henao-Mejia J, Elinav E, Strowig T, Flavell RA. Inflammasomes: far beyond inflammation. Nat Immunol. 2012;13:321-324. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 140] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 157. | Medzhitov R, Janeway CA. Innate immunity: the virtues of a nonclonal system of recognition. Cell. 1997;91:295-298. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1714] [Cited by in RCA: 1630] [Article Influence: 58.2] [Reference Citation Analysis (0)] |
| 158. | Keller M, Rüegg A, Werner S, Beer HD. Active caspase-1 is a regulator of unconventional protein secretion. Cell. 2008;132:818-831. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 660] [Cited by in RCA: 694] [Article Influence: 40.8] [Reference Citation Analysis (0)] |
| 159. | Yu HB, Finlay BB. The caspase-1 inflammasome: a pilot of innate immune responses. Cell Host Microbe. 2008;4:198-208. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 165] [Cited by in RCA: 171] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 160. | Martinon F, Mayor A, Tschopp J. The inflammasomes: guardians of the body. Annu Rev Immunol. 2009;27:229-265. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1734] [Cited by in RCA: 1848] [Article Influence: 115.5] [Reference Citation Analysis (0)] |
| 161. | Jin MS, Lee JO. Structures of the toll-like receptor family and its ligand complexes. Immunity. 2008;29:182-191. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 373] [Cited by in RCA: 408] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 162. | Gurung P, Li B, Subbarao Malireddi RK, Lamkanfi M, Geiger TL, Kanneganti TD. Chronic TLR Stimulation Controls NLRP3 Inflammasome Activation through IL-10 Mediated Regulation of NLRP3 Expression and Caspase-8 Activation. Sci Rep. 2015;5:14488. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 94] [Cited by in RCA: 127] [Article Influence: 12.7] [Reference Citation Analysis (0)] |
| 163. | Chuang SY, Yang CH, Chou CC, Chiang YP, Chuang TH, Hsu LC. TLR-induced PAI-2 expression suppresses IL-1β processing via increasing autophagy and NLRP3 degradation. Proc Natl Acad Sci USA. 2013;110:16079-16084. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 126] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 164. | Vaahtovuo J, Munukka E, Korkeamäki M, Luukkainen R, Toivanen P. Fecal microbiota in early rheumatoid arthritis. J Rheumatol. 2008;35:1500-1505. [PubMed] |
| 165. | Boursi B, Mamtani R, Haynes K, Yang YX. The effect of past antibiotic exposure on diabetes risk. Eur J Endocrinol. 2015;172:639-648. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 107] [Cited by in RCA: 111] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 166. | Kozyrskyj AL, Ernst P, Becker AB. Increased risk of childhood asthma from antibiotic use in early life. Chest. 2007;131:1753-1759. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 181] [Cited by in RCA: 189] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 167. | Bisgaard H, Li N, Bonnelykke K, Chawes BL, Skov T, Paludan-Müller G, Stokholm J, Smith B, Krogfelt KA. Reduced diversity of the intestinal microbiota during infancy is associated with increased risk of allergic disease at school age. J Allergy Clin Immunol. 2011;128:646-652.e1-5. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 492] [Cited by in RCA: 535] [Article Influence: 38.2] [Reference Citation Analysis (0)] |
| 168. | Abrahamsson TR, Jakobsson HE, Andersson AF, Björkstén B, Engstrand L, Jenmalm MC. Low diversity of the gut microbiota in infants with atopic eczema. J Allergy Clin Immunol. 2012;129:434-440, 440.e1-2. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 518] [Cited by in RCA: 584] [Article Influence: 41.7] [Reference Citation Analysis (0)] |
| 169. | Francino MP. Antibiotics and the Human Gut Microbiome: Dysbioses and Accumulation of Resistances. Front Microbiol. 2015;6:1543. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 340] [Cited by in RCA: 516] [Article Influence: 57.3] [Reference Citation Analysis (0)] |
| 170. | Manichanh C, Rigottier-Gois L, Bonnaud E, Gloux K, Pelletier E, Frangeul L, Nalin R, Jarrin C, Chardon P, Marteau P. Reduced diversity of faecal microbiota in Crohn’s disease revealed by a metagenomic approach. Gut. 2006;55:205-211. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1633] [Cited by in RCA: 1677] [Article Influence: 88.3] [Reference Citation Analysis (0)] |
| 171. | Kostic AD, Gevers D, Siljander H, Vatanen T, Hyötyläinen T, Hämäläinen AM, Peet A, Tillmann V, Pöhö P, Mattila I. The dynamics of the human infant gut microbiome in development and in progression toward type 1 diabetes. Cell Host Microbe. 2015;17:260-273. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 691] [Cited by in RCA: 854] [Article Influence: 85.4] [Reference Citation Analysis (0)] |
| 172. | Frank DN, Robertson CE, Hamm CM, Kpadeh Z, Zhang T, Chen H, Zhu W, Sartor RB, Boedeker EC, Harpaz N. Disease phenotype and genotype are associated with shifts in intestinal-associated microbiota in inflammatory bowel diseases. Inflamm Bowel Dis. 2011;17:179-184. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 499] [Cited by in RCA: 465] [Article Influence: 33.2] [Reference Citation Analysis (0)] |
| 173. | Sartor RB. Genetics and environmental interactions shape the intestinal microbiome to promote inflammatory bowel disease versus mucosal homeostasis. Gastroenterology. 2010;139:1816-1819. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 132] [Cited by in RCA: 141] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 174. | Foliaki S, Pearce N, Björkstén B, Mallol J, Montefort S, von Mutius E. Antibiotic use in infancy and symptoms of asthma, rhinoconjunctivitis, and eczema in children 6 and 7 years old: International Study of Asthma and Allergies in Childhood Phase III. J Allergy Clin Immunol. 2009;124:982-989. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 99] [Cited by in RCA: 103] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 175. | Mårild K, Ye W, Lebwohl B, Green PH, Blaser MJ, Card T, Ludvigsson JF. Antibiotic exposure and the development of coeliac disease: a nationwide case-control study. BMC Gastroenterol. 2013;13:109. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 112] [Cited by in RCA: 129] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 176. | Spor A, Koren O, Ley R. Unravelling the effects of the environment and host genotype on the gut microbiome. Nat Rev Microbiol. 2011;9:279-290. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1009] [Cited by in RCA: 1059] [Article Influence: 75.6] [Reference Citation Analysis (0)] |
| 177. | Goodrich JK, Waters JL, Poole AC, Sutter JL, Koren O, Blekhman R, Beaumont M, Van Treuren W, Knight R, Bell JT. Human genetics shape the gut microbiome. Cell. 2014;159:789-799. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1911] [Cited by in RCA: 2150] [Article Influence: 215.0] [Reference Citation Analysis (0)] |
| 178. | Hov JR, Zhong H, Qin B, Anmarkrud JA, Holm K, Franke A, Lie BA, Karlsen TH. The Influence of the Autoimmunity-Associated Ancestral HLA Haplotype AH8.1 on the Human Gut Microbiota: A Cross-Sectional Study. PLoS One. 2015;10:e0133804. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 26] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 179. | Markle JG, Frank DN, Mortin-Toth S, Robertson CE, Feazel LM, Rolle-Kampczyk U, von Bergen M, McCoy KD, Macpherson AJ, Danska JS. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science. 2013;339:1084-1088. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1230] [Cited by in RCA: 1469] [Article Influence: 122.4] [Reference Citation Analysis (0)] |
| 180. | Yurkovetskiy L, Burrows M, Khan AA, Graham L, Volchkov P, Becker L, Antonopoulos D, Umesaki Y, Chervonsky AV. Gender bias in autoimmunity is influenced by microbiota. Immunity. 2013;39:400-412. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 565] [Cited by in RCA: 691] [Article Influence: 57.6] [Reference Citation Analysis (0)] |
| 181. | Markle JG, Frank DN, Adeli K, von Bergen M, Danska JS. Microbiome manipulation modifies sex-specific risk for autoimmunity. Gut Microbes. 2014;5:485-493. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 60] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 182. | Whitacre CC, Reingold SC, O’Looney PA. A gender gap in autoimmunity. Science. 1999;283:1277-1278. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 613] [Cited by in RCA: 589] [Article Influence: 22.7] [Reference Citation Analysis (0)] |
| 183. | Young NA, Wu LC, Burd CJ, Friedman AK, Kaffenberger BH, Rajaram MV, Schlesinger LS, James H, Shupnik MA, Jarjour WN. Estrogen modulation of endosome-associated toll-like receptor 8: an IFNα-independent mechanism of sex-bias in systemic lupus erythematosus. Clin Immunol. 2014;151:66-77. [PubMed] |
| 184. | Bogdanos DP, Choudhuri K, Vergani D. Molecular mimicry and autoimmune liver disease: virtuous intentions, malign consequences. Liver. 2001;21:225-232. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 126] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 185. | Christen U, von Herrath MG. Induction, acceleration or prevention of autoimmunity by molecular mimicry. Mol Immunol. 2004;40:1113-1120. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 68] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 186. | Doherty DG, Penzotti JE, Koelle DM, Kwok WW, Lybrand TP, Masewicz S, Nepom GT. Structural basis of specificity and degeneracy of T cell recognition: pluriallelic restriction of T cell responses to a peptide antigen involves both specific and promiscuous interactions between the T cell receptor, peptide, and HLA-DR. J Immunol. 1998;161:3527-3535. [PubMed] |
| 187. | Hintermann E, Holdener M, Bayer M, Loges S, Pfeilschifter JM, Granier C, Manns MP, Christen U. Epitope spreading of the anti-CYP2D6 antibody response in patients with autoimmune hepatitis and in the CYP2D6 mouse model. J Autoimmun. 2011;37:242-253. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 51] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 188. | Zhang W, Reichlin M. A possible link between infection with burkholderia bacteria and systemic lupus erythematosus based on epitope mimicry. Clin Dev Immunol. 2008;2008:683489. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 31] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 189. | Ruff WE, Vieira SM, Kriegel MA. The role of the gut microbiota in the pathogenesis of antiphospholipid syndrome. Curr Rheumatol Rep. 2015;17:472. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 32] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 190. | Deng GM, Tsokos GC. Cholera toxin B accelerates disease progression in lupus-prone mice by promoting lipid raft aggregation. J Immunol. 2008;181:4019-4026. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 96] [Cited by in RCA: 81] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 191. | Szymula A, Rosenthal J, Szczerba BM, Bagavant H, Fu SM, Deshmukh US. T cell epitope mimicry between Sjögren’s syndrome Antigen A (SSA)/Ro60 and oral, gut, skin and vaginal bacteria. Clin Immunol. 2014;152:1-9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 113] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 192. | Sorini C, Falcone M. Shaping the (auto)immune response in the gut: the role of intestinal immune regulation in the prevention of type 1 diabetes. Am J Clin Exp Immunol. 2013;2:156-171. [PubMed] |
| 193. | Turley SJ, Lee JW, Dutton-Swain N, Mathis D, Benoist C. Endocrine self and gut non-self intersect in the pancreatic lymph nodes. Proc Natl Acad Sci USA. 2005;102:17729-17733. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 124] [Cited by in RCA: 140] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 194. | Berer K, Mues M, Koutrolos M, Rasbi ZA, Boziki M, Johner C, Wekerle H, Krishnamoorthy G. Commensal microbiota and myelin autoantigen cooperate to trigger autoimmune demyelination. Nature. 2011;479:538-541. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 867] [Cited by in RCA: 936] [Article Influence: 66.9] [Reference Citation Analysis (0)] |
| 195. | Wiest R, Garcia-Tsao G. Bacterial translocation (BT) in cirrhosis. Hepatology. 2005;41:422-433. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 492] [Cited by in RCA: 505] [Article Influence: 25.3] [Reference Citation Analysis (0)] |
| 196. | Son G, Kremer M, Hines IN. Contribution of gut bacteria to liver pathobiology. Gastroenterol Res Pract. 2010;2010. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 76] [Cited by in RCA: 107] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 197. | Konturek PC, Haziri D, Brzozowski T, Hess T, Heyman S, Kwiecien S, Konturek SJ, Koziel J. Emerging role of fecal microbiota therapy in the treatment of gastrointestinal and extra-gastrointestinal diseases. J Physiol Pharmacol. 2015;66:483-491. [PubMed] |
| 198. | Mehal WZ. The Gordian Knot of dysbiosis, obesity and NAFLD. Nat Rev Gastroenterol Hepatol. 2013;10:637-644. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 105] [Cited by in RCA: 118] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 199. | Al-Lahham SH, Peppelenbosch MP, Roelofsen H, Vonk RJ, Venema K. Biological effects of propionic acid in humans; metabolism, potential applications and underlying mechanisms. Biochim Biophys Acta. 2010;1801:1175-1183. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 347] [Cited by in RCA: 385] [Article Influence: 25.7] [Reference Citation Analysis (0)] |
| 200. | Tan J, McKenzie C, Potamitis M, Thorburn AN, Mackay CR, Macia L. The role of short-chain fatty acids in health and disease. Adv Immunol. 2014;121:91-119. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1076] [Cited by in RCA: 1616] [Article Influence: 146.9] [Reference Citation Analysis (0)] |
| 201. | Finnie IA, Dwarakanath AD, Taylor BA, Rhodes JM. Colonic mucin synthesis is increased by sodium butyrate. Gut. 1995;36:93-99. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 149] [Cited by in RCA: 173] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 202. | Burger-van Paassen N, Vincent A, Puiman PJ, van der Sluis M, Bouma J, Boehm G, van Goudoever JB, van Seuningen I, Renes IB. The regulation of intestinal mucin MUC2 expression by short-chain fatty acids: implications for epithelial protection. Biochem J. 2009;420:211-219. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 349] [Cited by in RCA: 436] [Article Influence: 27.3] [Reference Citation Analysis (0)] |
| 203. | Peng L, Li ZR, Green RS, Holzman IR, Lin J. Butyrate enhances the intestinal barrier by facilitating tight junction assembly via activation of AMP-activated protein kinase in Caco-2 cell monolayers. J Nutr. 2009;139:1619-1625. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1443] [Cited by in RCA: 1346] [Article Influence: 84.1] [Reference Citation Analysis (0)] |
| 204. | Lewis K, Lutgendorff F, Phan V, Söderholm JD, Sherman PM, McKay DM. Enhanced translocation of bacteria across metabolically stressed epithelia is reduced by butyrate. Inflamm Bowel Dis. 2010;16:1138-1148. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 194] [Cited by in RCA: 250] [Article Influence: 16.7] [Reference Citation Analysis (0)] |
| 205. | Arpaia N, Campbell C, Fan X, Dikiy S, van der Veeken J, deRoos P, Liu H, Cross JR, Pfeffer K, Coffer PJ. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature. 2013;504:451-455. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2516] [Cited by in RCA: 3434] [Article Influence: 286.2] [Reference Citation Analysis (0)] |
| 206. | Singh N, Gurav A, Sivaprakasam S, Brady E, Padia R, Shi H, Thangaraju M, Prasad PD, Manicassamy S, Munn DH. Activation of Gpr109a, receptor for niacin and the commensal metabolite butyrate, suppresses colonic inflammation and carcinogenesis. Immunity. 2014;40:128-139. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1198] [Cited by in RCA: 1694] [Article Influence: 154.0] [Reference Citation Analysis (0)] |
| 207. | Segain JP, Raingeard de la Blétière D, Bourreille A, Leray V, Gervois N, Rosales C, Ferrier L, Bonnet C, Blottière HM, Galmiche JP. Butyrate inhibits inflammatory responses through NFkappaB inhibition: implications for Crohn’s disease. Gut. 2000;47:397-403. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 886] [Cited by in RCA: 974] [Article Influence: 39.0] [Reference Citation Analysis (0)] |
| 208. | Bordonaro M, Lazarova DL, Augenlicht LH, Sartorelli AC. Cell type- and promoter-dependent modulation of the Wnt signaling pathway by sodium butyrate. Int J Cancer. 2002;97:42-51. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 28] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 209. | Rabinowitz JD, White E. Autophagy and metabolism. Science. 2010;330:1344-1348. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1632] [Cited by in RCA: 1584] [Article Influence: 105.6] [Reference Citation Analysis (0)] |
| 210. | Zhang J, Yi M, Zha L, Chen S, Li Z, Li C, Gong M, Deng H, Chu X, Chen J. Sodium Butyrate Induces Endoplasmic Reticulum Stress and Autophagy in Colorectal Cells: Implications for Apoptosis. PLoS One. 2016;11:e0147218. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 75] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 211. | Madrigal-Matute J, Cuervo AM. Regulation of Liver Metabolism by Autophagy. Gastroenterology. 2016;150:328-339. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 242] [Cited by in RCA: 247] [Article Influence: 27.4] [Reference Citation Analysis (0)] |
| 212. | Schwab M, Reynders V, Ulrich S, Zahn N, Stein J, Schröder O. PPARgamma is a key target of butyrate-induced caspase-3 activation in the colorectal cancer cell line Caco-2. Apoptosis. 2006;11:1801-1811. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 48] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 213. | Tang Y, Chen Y, Jiang H, Nie D. Short-chain fatty acids induced autophagy serves as an adaptive strategy for retarding mitochondria-mediated apoptotic cell death. Cell Death Differ. 2011;18:602-618. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 131] [Cited by in RCA: 151] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 214. | Czaja AJ. Targeting apoptosis in autoimmune hepatitis. Dig Dis Sci. 2014;59:2890-2904. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 42] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 215. | Bourriaud C, Robins RJ, Martin L, Kozlowski F, Tenailleau E, Cherbut C, Michel C. Lactate is mainly fermented to butyrate by human intestinal microfloras but inter-individual variation is evident. J Appl Microbiol. 2005;99:201-212. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 236] [Cited by in RCA: 245] [Article Influence: 12.3] [Reference Citation Analysis (0)] |
| 216. | Duncan SH, Louis P, Flint HJ. Lactate-utilizing bacteria, isolated from human feces, that produce butyrate as a major fermentation product. Appl Environ Microbiol. 2004;70:5810-5817. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 676] [Cited by in RCA: 786] [Article Influence: 37.4] [Reference Citation Analysis (0)] |
| 217. | Brown CT, Davis-Richardson AG, Giongo A, Gano KA, Crabb DB, Mukherjee N, Casella G, Drew JC, Ilonen J, Knip M. Gut microbiome metagenomics analysis suggests a functional model for the development of autoimmunity for type 1 diabetes. PLoS One. 2011;6:e25792. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 496] [Cited by in RCA: 590] [Article Influence: 42.1] [Reference Citation Analysis (0)] |
| 218. | Visser J, Rozing J, Sapone A, Lammers K, Fasano A. Tight junctions, intestinal permeability, and autoimmunity: celiac disease and type 1 diabetes paradigms. Ann N Y Acad Sci. 2009;1165:195-205. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 187] [Cited by in RCA: 175] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 219. | Vaarala O, Atkinson MA, Neu J. The “perfect storm” for type 1 diabetes: the complex interplay between intestinal microbiota, gut permeability, and mucosal immunity. Diabetes. 2008;57:2555-2562. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 371] [Cited by in RCA: 366] [Article Influence: 21.5] [Reference Citation Analysis (0)] |
| 220. | Fanning AS, Jameson BJ, Jesaitis LA, Anderson JM. The tight junction protein ZO-1 establishes a link between the transmembrane protein occludin and the actin cytoskeleton. J Biol Chem. 1998;273:29745-29753. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1010] [Cited by in RCA: 1061] [Article Influence: 39.3] [Reference Citation Analysis (0)] |
| 221. | Watts T, Berti I, Sapone A, Gerarduzzi T, Not T, Zielke R, Fasano A. Role of the intestinal tight junction modulator zonulin in the pathogenesis of type I diabetes in BB diabetic-prone rats. Proc Natl Acad Sci USA. 2005;102:2916-2921. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 234] [Cited by in RCA: 226] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 222. | Denker BM, Nigam SK. Molecular structure and assembly of the tight junction. Am J Physiol. 1998;274:F1-F9. [PubMed] |
| 223. | Mitic LL, Anderson JM. Molecular architecture of tight junctions. Annu Rev Physiol. 1998;60:121-142. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 550] [Cited by in RCA: 519] [Article Influence: 19.2] [Reference Citation Analysis (0)] |
| 224. | Sharma R, Young C, Neu J. Molecular modulation of intestinal epithelial barrier: contribution of microbiota. J Biomed Biotechnol. 2010;2010:305879. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 95] [Cited by in RCA: 131] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 225. | Cario E, Gerken G, Podolsky DK. Toll-like receptor 2 enhances ZO-1-associated intestinal epithelial barrier integrity via protein kinase C. Gastroenterology. 2004;127:224-238. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 362] [Cited by in RCA: 382] [Article Influence: 18.2] [Reference Citation Analysis (0)] |
| 226. | Li X, Wang C, Nie J, Lv D, Wang T, Xu Y. Toll-like receptor 4 increases intestinal permeability through up-regulation of membrane PKC activity in alcoholic steatohepatitis. Alcohol. 2013;47:459-465. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 37] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 227. | Fasano A. Zonulin and its regulation of intestinal barrier function: the biological door to inflammation, autoimmunity, and cancer. Physiol Rev. 2011;91:151-175. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 500] [Cited by in RCA: 635] [Article Influence: 45.4] [Reference Citation Analysis (0)] |
| 228. | Fasano A. Leaky gut and autoimmune diseases. Clin Rev Allergy Immunol. 2012;42:71-78. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 181] [Cited by in RCA: 216] [Article Influence: 16.6] [Reference Citation Analysis (0)] |
| 229. | Clarke TB, Davis KM, Lysenko ES, Zhou AY, Yu Y, Weiser JN. Recognition of peptidoglycan from the microbiota by Nod1 enhances systemic innate immunity. Nat Med. 2010;16:228-231. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 793] [Cited by in RCA: 881] [Article Influence: 58.7] [Reference Citation Analysis (0)] |
| 230. | Hase K, Kawano K, Nochi T, Pontes GS, Fukuda S, Ebisawa M, Kadokura K, Tobe T, Fujimura Y, Kawano S. Uptake through glycoprotein 2 of FimH(+) bacteria by M cells initiates mucosal immune response. Nature. 2009;462:226-230. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 450] [Cited by in RCA: 490] [Article Influence: 32.7] [Reference Citation Analysis (0)] |
| 231. | van Gerven NM, Verwer BJ, Witte BI, van Hoek B, Coenraad MJ, van Erpecum KJ, Beuers U, van Buuren HR, de Man RA, Drenth JP. Relapse is almost universal after withdrawal of immunosuppressive medication in patients with autoimmune hepatitis in remission. J Hepatol. 2013;58:141-147. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 137] [Cited by in RCA: 164] [Article Influence: 13.7] [Reference Citation Analysis (0)] |
| 232. | Czaja AJ. Review article: permanent drug withdrawal is desirable and achievable for autoimmune hepatitis. Aliment Pharmacol Ther. 2014;39:1043-1058. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 26] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 233. | Czaja AJ, Carpenter HA. Progressive fibrosis during corticosteroid therapy of autoimmune hepatitis. Hepatology. 2004;39:1631-1638. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 88] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 234. | Czaja AJ. Current and prospective pharmacotherapy for autoimmune hepatitis. Expert Opin Pharmacother. 2014;15:1715-1736. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 29] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 235. | Chan ER, Hester J, Kalady M, Xiao H, Li X, Serre D. A novel method for determining microflora composition using dynamic phylogenetic analysis of 16S ribosomal RNA deep sequencing data. Genomics. 2011;98:253-259. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 9] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 236. | Mandal RS, Saha S, Das S. Metagenomic surveys of gut microbiota. Genomics Proteomics Bioinformatics. 2015;13:148-158. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 59] [Cited by in RCA: 65] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 237. | Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci USA. 1985;82:6955-6959. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2037] [Cited by in RCA: 1798] [Article Influence: 45.0] [Reference Citation Analysis (0)] |
| 238. | Wang Q, Garrity GM, Tiedje JM, Cole JR. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261-5267. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12842] [Cited by in RCA: 13189] [Article Influence: 732.7] [Reference Citation Analysis (0)] |
| 239. | Riehle K, Coarfa C, Jackson A, Ma J, Tandon A, Paithankar S, Raghuraman S, Mistretta TA, Saulnier D, Raza S. The Genboree Microbiome Toolset and the analysis of 16S rRNA microbial sequences. BMC Bioinformatics. 2012;13 Suppl 13:S11. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 36] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 240. | Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335-336. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29318] [Cited by in RCA: 23743] [Article Influence: 1582.9] [Reference Citation Analysis (0)] |
| 241. | Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537-7541. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14372] [Cited by in RCA: 13803] [Article Influence: 862.7] [Reference Citation Analysis (0)] |
| 242. | Wei ZG, Zhang SW. MtHc: a motif-based hierarchical method for clustering massive 16S rRNA sequences into OTUs. Mol Biosyst. 2015;11:1907-1913. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 17] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 243. | Turnbaugh PJ, Ley RE, Hamady M, Fraser-Liggett CM, Knight R, Gordon JI. The human microbiome project. Nature. 2007;449:804-810. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4573] [Cited by in RCA: 3725] [Article Influence: 206.9] [Reference Citation Analysis (0)] |
| 244. | Jumpstart Consortium Human Microbiome Project Data Generation Working Group. Evaluation of 16S rDNA-based community profiling for human microbiome research. PLoS One. 2012;7:e39315. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 202] [Cited by in RCA: 206] [Article Influence: 15.8] [Reference Citation Analysis (0)] |
| 245. | Brugère JF, Mihajlovski A, Missaoui M, Peyret P. Tools for stools: the challenge of assessing human intestinal microbiota using molecular diagnostics. Expert Rev Mol Diagn. 2009;9:353-365. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 23] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 246. | Fraher MH, O’Toole PW, Quigley EM. Techniques used to characterize the gut microbiota: a guide for the clinician. Nat Rev Gastroenterol Hepatol. 2012;9:312-322. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 226] [Cited by in RCA: 235] [Article Influence: 18.1] [Reference Citation Analysis (0)] |
| 247. | Tottey W, Denonfoux J, Jaziri F, Parisot N, Missaoui M, Hill D, Borrel G, Peyretaillade E, Alric M, Harris HM. The human gut chip “HuGChip”, an explorative phylogenetic microarray for determining gut microbiome diversity at family level. PLoS One. 2013;8:e62544. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 34] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 248. | DeSantis TZ, Brodie EL, Moberg JP, Zubieta IX, Piceno YM, Andersen GL. High-density universal 16S rRNA microarray analysis reveals broader diversity than typical clone library when sampling the environment. Microb Ecol. 2007;53:371-383. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 377] [Cited by in RCA: 281] [Article Influence: 15.6] [Reference Citation Analysis (0)] |
| 249. | Rajilić-Stojanović M, Heilig HG, Molenaar D, Kajander K, Surakka A, Smidt H, de Vos WM. Development and application of the human intestinal tract chip, a phylogenetic microarray: analysis of universally conserved phylotypes in the abundant microbiota of young and elderly adults. Environ Microbiol. 2009;11:1736-1751. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 366] [Cited by in RCA: 347] [Article Influence: 21.7] [Reference Citation Analysis (0)] |
| 250. | Dugat-Bony E, Peyretaillade E, Parisot N, Biderre-Petit C, Jaziri F, Hill D, Rimour S, Peyret P. Detecting unknown sequences with DNA microarrays: explorative probe design strategies. Environ Microbiol. 2012;14:356-371. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 28] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 251. | Eisen JA. Environmental shotgun sequencing: its potential and challenges for studying the hidden world of microbes. PLoS Biol. 2007;5:e82. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 186] [Cited by in RCA: 152] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 252. | Vieira SM, Pagovich OE, Kriegel MA. Diet, microbiota and autoimmune diseases. Lupus. 2014;23:518-526. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 122] [Cited by in RCA: 129] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 253. | López P, González-Rodríguez I, Sánchez B, Ruas-Madiedo P, Suárez A, Margolles A, Gueimonde M. Interaction of Bifidobacterium bifidum LMG13195 with HT29 cells influences regulatory-T-cell-associated chemokine receptor expression. Appl Environ Microbiol. 2012;78:2850-2857. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 46] [Cited by in RCA: 53] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 254. | Dolpady J, Sorini C, Di Pietro C, Cosorich I, Ferrarese R, Saita D, Clementi M, Canducci F, Falcone M. Oral Probiotic VSL#3 Prevents Autoimmune Diabetes by Modulating Microbiota and Promoting Indoleamine 2,3-Dioxygenase-Enriched Tolerogenic Intestinal Environment. J Diabetes Res. 2016;2016:7569431. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 99] [Cited by in RCA: 105] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 255. | Zhang H, Liao X, Sparks JB, Luo XM. Dynamics of gut microbiota in autoimmune lupus. Appl Environ Microbiol. 2014;80:7551-7560. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 182] [Cited by in RCA: 223] [Article Influence: 20.3] [Reference Citation Analysis (0)] |
| 256. | Hsieh CC, Lin BF. Dietary factors regulate cytokines in murine models of systemic lupus erythematosus. Autoimmun Rev. 2011;11:22-27. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 34] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 257. | Stone M, Fortin PR, Pacheco-Tena C, Inman RD. Should tetracycline treatment be used more extensively for rheumatoid arthritis? Metaanalysis demonstrates clinical benefit with reduction in disease activity. J Rheumatol. 2003;30:2112-2122. [PubMed] |
| 258. | Lopetuso LR, Scaldaferri F, Bruno G, Petito V, Franceschi F, Gasbarrini A. The therapeutic management of gut barrier leaking: the emerging role for mucosal barrier protectors. Eur Rev Med Pharmacol Sci. 2015;19:1068-1076. [PubMed] |
| 259. | Scaldaferri F, Lopetuso LR, Petito V, Cufino V, Bilotta M, Arena V, Stigliano E, Maulucci G, Papi M, Emiliana CM. Gelatin tannate ameliorates acute colitis in mice by reinforcing mucus layer and modulating gut microbiota composition: Emerging role for ‘gut barrier protectors’ in IBD? United European Gastroenterol J. 2014;2:113-122. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 32] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 260. | Frasca G, Cardile V, Puglia C, Bonina C, Bonina F. Gelatin tannate reduces the proinflammatory effects of lipopolysaccharide in human intestinal epithelial cells. Clin Exp Gastroenterol. 2012;5:61-67. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 14] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 261. | Pawar RD, Ramanjaneyulu A, Kulkarni OP, Lech M, Segerer S, Anders HJ. Inhibition of Toll-like receptor-7 (TLR-7) or TLR-7 plus TLR-9 attenuates glomerulonephritis and lung injury in experimental lupus. J Am Soc Nephrol. 2007;18:1721-1731. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 176] [Cited by in RCA: 178] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 262. | Telesford KM, Yan W, Ochoa-Reparaz J, Pant A, Kircher C, Christy MA, Begum-Haque S, Kasper DL, Kasper LH. A commensal symbiotic factor derived from Bacteroides fragilis promotes human CD39(+)Foxp3(+) T cells and Treg function. Gut Microbes. 2015;6:234-242. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 189] [Article Influence: 18.9] [Reference Citation Analysis (0)] |
| 263. | Calcinaro F, Dionisi S, Marinaro M, Candeloro P, Bonato V, Marzotti S, Corneli RB, Ferretti E, Gulino A, Grasso F. Oral probiotic administration induces interleukin-10 production and prevents spontaneous autoimmune diabetes in the non-obese diabetic mouse. Diabetologia. 2005;48:1565-1575. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 247] [Cited by in RCA: 230] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 264. | Latham T, Mackay L, Sproul D, Karim M, Culley J, Harrison DJ, Hayward L, Langridge-Smith P, Gilbert N, Ramsahoye BH. Lactate, a product of glycolytic metabolism, inhibits histone deacetylase activity and promotes changes in gene expression. Nucleic Acids Res. 2012;40:4794-4803. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 199] [Cited by in RCA: 254] [Article Influence: 19.5] [Reference Citation Analysis (0)] |
| 265. | Zhang W, Gu Y, Chen Y, Deng H, Chen L, Chen S, Zhang G, Gao Z. Intestinal flora imbalance results in altered bacterial translocation and liver function in rats with experimental cirrhosis. Eur J Gastroenterol Hepatol. 2010;22:1481-1486. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 36] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 266. | Nardone G, Compare D, Liguori E, Di Mauro V, Rocco A, Barone M, Napoli A, Lapi D, Iovene MR, Colantuoni A. Protective effects of Lactobacillus paracasei F19 in a rat model of oxidative and metabolic hepatic injury. Am J Physiol Gastrointest Liver Physiol. 2010;299:G669-G676. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 75] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 267. | Elfaki DA, Lindor KD. Antibiotics for the treatment of primary sclerosing cholangitis. Am J Ther. 2011;18:261-265. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 20] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 268. | Tabibian JH, Talwalkar JA, Lindor KD. Role of the microbiota and antibiotics in primary sclerosing cholangitis. Biomed Res Int. 2013;2013:389537. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 76] [Cited by in RCA: 70] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 269. | Tabibian JH, Weeding E, Jorgensen RA, Petz JL, Keach JC, Talwalkar JA, Lindor KD. Randomised clinical trial: vancomycin or metronidazole in patients with primary sclerosing cholangitis - a pilot study. Aliment Pharmacol Ther. 2013;37:604-612. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 217] [Cited by in RCA: 199] [Article Influence: 16.6] [Reference Citation Analysis (0)] |
| 270. | Hooper LV, Littman DR, Macpherson AJ. Interactions between the microbiota and the immune system. Science. 2012;336:1268-1273. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3003] [Cited by in RCA: 2988] [Article Influence: 229.8] [Reference Citation Analysis (0)] |
| 271. | Nikoopour E, Singh B. Reciprocity in microbiome and immune system interactions and its implications in disease and health. Inflamm Allergy Drug Targets. 2014;13:94-104. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 17] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 272. | Candon S, Perez-Arroyo A, Marquet C, Valette F, Foray AP, Pelletier B, Milani C, Ventura M, Bach JF, Chatenoud L. Antibiotics in early life alter the gut microbiome and increase disease incidence in a spontaneous mouse model of autoimmune insulin-dependent diabetes. PLoS One. 2015;10:e0125448. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 149] [Cited by in RCA: 165] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 273. | Wu GD, Lewis JD, Hoffmann C, Chen YY, Knight R, Bittinger K, Hwang J, Chen J, Berkowsky R, Nessel L. Sampling and pyrosequencing methods for characterizing bacterial communities in the human gut using 16S sequence tags. BMC Microbiol. 2010;10:206. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 276] [Cited by in RCA: 300] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 274. | Eckburg PB, Bik EM, Bernstein CN, Purdom E, Dethlefsen L, Sargent M, Gill SR, Nelson KE, Relman DA. Diversity of the human intestinal microbial flora. Science. 2005;308:1635-1638. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5700] [Cited by in RCA: 5591] [Article Influence: 279.6] [Reference Citation Analysis (2)] |
| 275. | Van den Abbeele P, Van de Wiele T, Verstraete W, Possemiers S. The host selects mucosal and luminal associations of coevolved gut microorganisms: a novel concept. FEMS Microbiol Rev. 2011;35:681-704. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 169] [Cited by in RCA: 180] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 276. | Laval L, Martin R, Natividad JN, Chain F, Miquel S, Desclée de Maredsous C, Capronnier S, Sokol H, Verdu EF, van Hylckama Vlieg JE. Lactobacillus rhamnosus CNCM I-3690 and the commensal bacterium Faecalibacterium prausnitzii A2-165 exhibit similar protective effects to induced barrier hyper-permeability in mice. Gut Microbes. 2015;6:1-9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 153] [Article Influence: 15.3] [Reference Citation Analysis (0)] |
| 277. | Sato T, Matsumoto K, Okumura T, Yokoi W, Naito E, Yoshida Y, Nomoto K, Ito M, Sawada H. Isolation of lactate-utilizing butyrate-producing bacteria from human feces and in vivo administration of Anaerostipes caccae strain L2 and galacto-oligosaccharides in a rat model. FEMS Microbiol Ecol. 2008;66:528-536. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 63] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 278. | Casanova MJ, Chaparro M, Domènech E, Barreiro-de Acosta M, Bermejo F, Iglesias E, Gomollón F, Rodrigo L, Calvet X, Esteve M. Safety of thiopurines and anti-TNF-α drugs during pregnancy in patients with inflammatory bowel disease. Am J Gastroenterol. 2013;108:433-440. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 166] [Cited by in RCA: 161] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 279. | Bakdash G, Vogelpoel LT, van Capel TM, Kapsenberg ML, de Jong EC. Retinoic acid primes human dendritic cells to induce gut-homing, IL-10-producing regulatory T cells. Mucosal Immunol. 2015;8:265-278. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 163] [Cited by in RCA: 182] [Article Influence: 18.2] [Reference Citation Analysis (0)] |