Published online May 21, 2014. doi: 10.3748/wjg.v20.i19.5897
Revised: October 2, 2013
Accepted: December 5, 2013
Published online: May 21, 2014
Processing time: 299 Days and 0.3 Hours
AIM: To assess hepatitis C virus (HCV) genotype patterns among high-risk Iranian groups, using real-time RT-PCR.
METHODS: In this study, we evaluated the distribution of different HCV genotypes among injection drug users and other high-risk groups over a 4-year period (from 2009 to 2012) using real-time polymerase chain reaction (PCR). Sera from 888 HCV-infected patients residing in southern and southwest Iran were genotyped using real-time PCR with common primers and specific probes. These patients were grouped into distinct exposure categories. Illicit drug users constituted the primary group and were further evaluated for HCV genotype distribution and parameters such as age range.
RESULTS: Of the examined HCV-infected patients, 62% were substance abusers, although the route of transmission could not be determined in approximately 30% of these patients. HCV genotyping revealed that Gt1 was the most prevalent genotype among the drug users as well as among patients with thalassemia, hemophilia, solid organ recipients and those on hemodialysis. Mixed infections were only seen in addict groups, where Gt2 genotype was also found. The highest frequencies in HCV-positive addict patients were observed in the 31-40 age group. Our research also showed that the addiction age has increased, whereas the addiction rate has dropped in this region. Most illicit drug users had more than one risk factor such as tattoo and/or a history of imprisonment.
CONCLUSION: This study revealed that the most common HCV-infection route and HCV-genotype in southern and southwest Iran was illicit drug abuse and Gt1, respectively.
Core tip: The primary treatment method in hepatitis C virus (HCV) infection, determination of evolution pathways, assessment of epidemiological status, and knowledge of HCV genotype distribution among high-risk groups such as addicts are very important. We assessed the different HCV genotypes among illicit drug users and other high-risk groups during a 4-year period from 2009 to 2012 using real-time PCR. We found that the most affected high-risk groups were illicit drug users and specified the respective age distribution and risk factors. An important finding in this research was the genotype pattern shift from 3 to 1, especially among addicts.
- Citation: Jamalidoust M, Namayandeh M, Asaei S, Aliabadi N, Ziyaeyan M. Determining hepatitis C virus genotype distribution among high-risk groups in Iran using real-time PCR. World J Gastroenterol 2014; 20(19): 5897-5902
- URL: https://www.wjgnet.com/1007-9327/full/v20/i19/5897.htm
- DOI: https://dx.doi.org/10.3748/wjg.v20.i19.5897
The increasing use of different types of injections, including herbal injections, particularly chemical drugs, in many developed and developing countries, including Iran, and the lack of an effective hepatitis C virus (HCV) vaccine have resulted in HCV infection becoming a major public health concern[1,2]. A lower prevalence of HCV infection has been reported in Iran compared to other parts of the world, particularly the Middle East[1,2]. Given that Injection Drug Users (IDUs) constitute a major group of HCV patients, planning their treatment is essential for community health[3,4].
HCV genotyping is the most significant predictor of treatment duration and evaluation of the course of infection, with different genotypes showing different treatment responses and varying epidemiological as well as virological features[5]. HCV has a single-stranded RNA genome and at least six different genotypes, more than 90 subtypes and many types of quasi-species variants based on scattered and local variations of its genome. This genetic heterogeneity is due to the lack of fidelity of viral RNA-dependent RNA polymerases[6]. HCV-RNA diversity is concentrated in the E1 and E2 glycoprotein-coding regions of the genome, while least heterogeneity is found in the regions encoding core and NS3 proteins, which represent structural and non-structural proteins, respectively. In total, the nucleotide sequences of the different genotypes differ from each other by 30%-35%, with a divergence of 92% in the 5′-UTR regions[1].
The HCV genotype pattern varies among different groups of infected individuals, particularly IDUs, in different areas. For example, the predominant genotypes are Gt3a and Gt4 in European countries and Gt3a in Argentina[1,7-9]. Mahfoud et al[10] showed that HCV Gt3 was distributed among Lebanese IDUs at a frequency of 57%. The aim of this study was to assess HCV genotype distribution among drug-addicted HCV-infected patients in Iran and compare this distribution with other high-risk groups of infected patients using real-time PCR, which simultaneously detects target HCV genomes with genotype-specific primer sets and probes.
From March 2009 to December 2012, a total of 888 HCV-RNA positive patients from southern and southwest Iran were enrolled in this cross-sectional study and underwent HCV genotyping. These patients completed a preliminary assessment, which allowed the identification of individuals with parenteral risk factors. Of these patients, 550 (61.9%) IDUs aged 18-74 years (mean ± SD, 37.89 ± 10.192 years) were identified, including 546 (99.27%) males and 4 (0.72%) females. None of the patients had previously been treated for HCV infection. Ethylenediaminetetraacetic acid (EDTA) plasma was prospectively collected, and aliquots were stored in 1.5 mL vials at -70 °C. HCV infection was reaffirmed by the detection of antibodies using an immunochromatographic assay (Artron, Burnaby, BC, Canada).
Viral RNA was isolated from 200 μL aliquots of serum samples using the InviTrap® Spin Blood RNA Mini kit (Invitec, Berlin) as per the manufacturer’s instructions, and eluted using 50 μL of nuclease-free water. The concentration of the isolated HCV-RNA was determined for the various samples and then genotyped, followed by real-time PCR using commercially available HCV kits (Genome Diagnostics Pvt. Ltd., Hague, Netherland) to determine HCV genotypes Gt1 to Gt4. All tests were carried out in accordance with the manufacturer’s instructions. The PCR conditions were as follows: 50 °C for 25 min and 95 °C for 10 min, followed by 50 cycles of 94 °C for 10 s, 55 °C for 32 s and 72 °C for 25 s. The reactions were performed in a 7500 real-time PCR system (Applied Biosystems, United States).
This study was performed using SPSS for Windows systems (Version 16.0, 2007, SPSS Inc, Chicago, IL, United States). Comparison of HCV genotypes, their distribution among different high-risk groups and among different age groups of addicts were analysed by the χ2 test.
A total of 888 HCV RNA-positive samples, collected between 2009 and 2012 from patients with chronic HCV infection, were included in this study [787 (83%) men and 99 (11.2%) women, 2 misplaced]. Of these, 738 (83%) HCV-positive serum samples were genotyped successfully using HCV genotype real-time PCR kits from Applied Biosystems. The frequency of HCV infection among various high-risk groups and HCV genotype distribution among the patients are presented in Table 1 and 2.
| High-risk group | Frequency | M/F |
| Addict | 550 (62.2) | 546/4 |
| Thalassemia | 38 (4.3) | 17/21 |
| Hemophilia | 8 (0.9) | 7/1 |
| Kidney graft | 4 (0.5) | 2/2 |
| Dialysis | 6 (0.7) | 5/1 |
| Liver graft | 3 (0.3) | 1/2 |
| Unknown | 277 (31.3) | 209/68 |
| Total | 886 (100.0) | 787/99 |
As shown in Table 1, HCV infection was most prevalent among substance abusers (62.2%), with the highest frequency (> 99%) among male drug users. The high prevalence of HCV infection can also be attributed to unknown factors compromising approximately 31% of cases. The other groups at high-risk for HCV infection included patients with thalassemia, hemophilia and dialysis as well as recipients of solid organs.
Overall, the most prevalent HCV genotypes were Gt1 (51.1%) and Gt3 (30.1%), whereas Gt4 (0.5%) and Gt2 (0.3%) were less prevalent. Mixed HCV genotypes were reported in only 8 addicts and included Gt1/3 and Gt1/2 co-infections with 0.7% and 0.2% prevalence, respectively (Table 2).
| High-risk group | Frequency | Undetectable | Gt1 | Gt2 | Gt3 | Gt4 | Gt1/2 | Gt1/3 |
| Addict | 550 | 67 (12.2) | 283 (51.5) | 0 (0.0) | 192 (34.9) | 0 (0.0) | 2 (0.4) | 6 (1.1) |
| Thalassemia | 38 | 15 (39.5) | 17 (44.7) | 0 (0.0) | 6 (15.8) | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| Hemophilia | 8 | 2 (25.0) | 5 (62.5) | 0 (0.0) | 1 (12.5) | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| Kidney graft | 4 | 1 (25.0) | 2 (50) | 0 (0.0) | 1 (25.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| Dialysis | 6 | 1 (16.7) | 4 (66.7) | 0 (0.0) | 0 (0.0) | 1 (16.7) | 0 (0.0) | 0 (0.0) |
| Liver graft | 3 | 1 (33.3) | 2 (66.7) | 0 (0.0) | 0 (0.) | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| Unknown | 277 | 64 (23.1) | 140 (50.5) | 3 (1.1) | 67 (24.2) | 3 (1.1) | 0 (0.0) | 0 (0.0) |
| Total | 886 | 151 (17.0) | 453 (51.1) | 3 (0.3) | 267 (30.1) | 4 (0.5) | 2 (0.2) | 6 (0.7) |
A past or current history of illicit drug use was the predominant risk factor for HCV infection. A total of 550 addicts [546 (99.27%) men and 4 (0.72%) women] underwent HCV genotyping; 483 (87%) addicts were genotyped, and included 475 (87.8%) patients with a single HCV genotype and 8 (12.2%) patients with a mixed genotype distribution. The most prevalent HCV genotype was Gt1, which was found in 283 addicts (51.5%), followed by Gt3 in 192 (34.9%), Gt1/3 in 6 (1.1%) and Gt1/2 in 2 (0.4%) addicts. Single or mixed genotypes of Gt2 and Gt4 were not detected in any of the addicts.
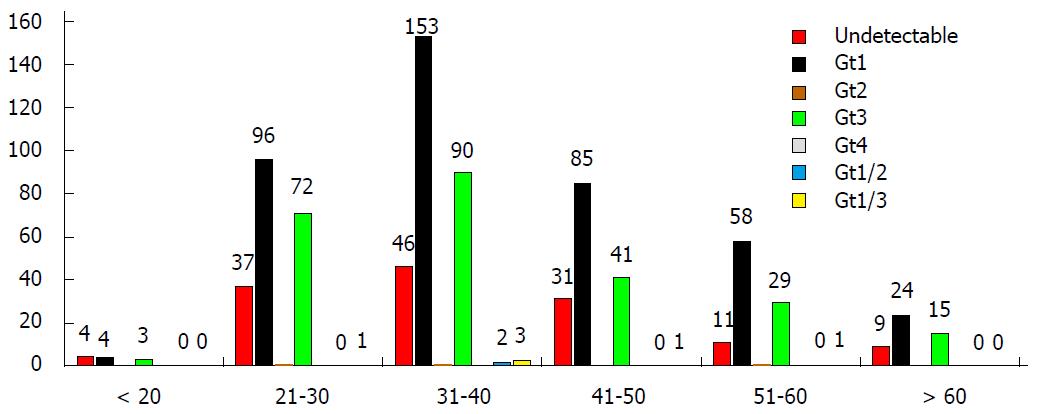
The prevalence of HCV infection and HCV genotype distribution varied between different age groups; as shown in Table 3 and Figure 1, the highest frequency of HCV-positive patients was observed in the 31-40 age group.
| Undetectable | Gt1 | Gt2 | Gt3 | Gt4 | Gt1/2 | Gt1/3 | Total | |
| < 20 | 0 (0.0) | 1 (33.3) | 0 (0.0) | 2 (66.66) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 3 (0.57) |
| 21-30 | 14 (11.02) | 59 (46.45) | 0 (0.0) | 53 (41.73) | 0 (0.0) | 0 (0.0) | 1 (0.78) | 127 (24.47) |
| 31-40 | 23 (10.95) | 113 (53.80) | 0 (0.0) | 69 (32.85) | 0 (0.0) | 2 (0.95) | 3 (1.42) | 210 (40.46) |
| 41-50 | 18 (16.36) | 58 (52.72) | 0 (0.0) | 33 (30) | 0 (0.0) | 0 (0.0) | 1 (0.9) | 110 (21.19) |
| 51-60 | 6 (11.53) | 25 (48.07) | 0 (0.0) | 20 (38.46) | 0 (0.0) | 0 (0.0) | 1 (1.92) | 52 (10.01) |
| > 60 | 1 (5.88) | 10 (58.82) | 0 (0.0) | 6 (35.29) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 17 (3.27) |
| Total | 62 (11.9) | 266 (51.25) | 0 (0.0) | 183 (35.26) | 0 (0.0) | 2 (0.38) | 6 (1.15) | 519 (100.0) |
As the most significant route of HCV infection in both developed and developing countries is addiction and the use of illicit drugs[10,11], the consideration of issues related to HCV infection such as viral load, genotyping, treatment response rate and epidemiological status is very important[2].
HCV genotyping among a defined population serves as a useful tool for determining HCV evolution in different geographical regions and in identifying the respective high-risk groups[12]. Specific HCV genotypes have also been reported as primary tools for determining infection course and assessing the duration of treatment[13]. However, certain investigations have implicated body mass index, IL28 genotype, gamma glutamyl transpeptidase, triglycerides and the level of miR-122 as other predictors of treatment duration[14].
Unfortunately, even in developed countries, only a small proportion of IDUs receive HCV treatment[15], and clinicians are often reluctant to treat them. The reasons for this trend may include high risk of reinfection, concomitant excessive alcohol intake and high rates of concurrent mental health problems[16].
Various studies suggest that the SVR Should you explain what SVR is? rate among IDUs is comparable to that among non-IDUs and that it is not significantly related to IDU status[10,17].
In clinical trials where peg interferon and ribavirin were prescribed, the median SVR rate among IDUs with chronic HCV infection was 54.3% (range, 18.1%-94.1%), comparable to 54%-63% seen in their non-IDU counterparts, whereas it was 68.5% in IDUs with acute HCV infection as opposed to 81.5% among non-IDUs[18,19].
HCV prevalence rate and genotype distribution among Asian and Middle Eastern countries is very diverse. Although Iran is surrounded by countries with a high prevalence of HCV of various genotypes, HCV prevalence in Iran (< 2%) is much lower than that of its neighbouring nations, Pakistan (5.31%) and Egypt (14.7%)[20]. Genotypes 1, 2 and 3 (Gt1-Gt3) have been found to have a global prevalence, while Gt4-Gt6 have a restricted pattern[20]. Gt4 has been chiefly detected in Egypt, some Arab countries such as Saudi Arabia, Syria and United Arab Emirates, and recently, also in certain parts of Europe[21]. Gt5 is restricted to South Africa, and Gt6 is primarily found in Southeast Asian countries such as China, Hong Kong and Taiwan[22,23].
Different studies have shown that Gt4 and Gt1 are the predominant genotypes in Arab and non-Arab countries, respectively, in the Middle East region[21]. A recent study carried out in a neighbouring country, Pakistan, identified Gt3 as the predominant HCV genotype[24,25]. Several investigations have also been conducted in different parts of Iran to determine HCV genotype distribution. The latest studies have indicated that HCV Gt among IDUs in south, southeast and north Iran (Mazandaran province), Gt1 in northeast and Gt3 in Isfahan province in central Iran, are the predominant genotypes[26,27].
The current study was performed to determine the distribution of HCV genotypes based on the route of infection, and revealed a significantly high prevalence (approximately 62%) of HCV infection among drug addicts. Many studies have been carried out in IDUs in different regions of Iran. In general, HCV seropositivity in Iranian addicts is higher than that in the general population (30%-90% vs 0.2%-2.0%)[28-31].
Our data also show that Gt1 is the most common genotype among thalassemia and hemophilia patients, solid organ recipients and hemodialysis patients as well as IDUs. A discrepancy between HCV genotype distribution among the addict and non-addict groups was observed, with Gt3 frequency in IDUs being less than in non-IDUs (Table 4). Among IDUs, Gt2 and Gt4 were rarely seen; the latter was not detected in any IDU patient, while Gt2 was detected in just 2 patients with mixed infections of Gt1/2. Overall, among the 888 patients, mixed HCV genotypes were detected in 8 serum samples from addicts and not in non-addict groups. This could be due to multiple injections in the addict group.
| Genotype | Non-addict | Addict |
| Gt1 | 50.7% | 51.5% |
| Gt2 | 1.0% | 0.0% |
| Gt3 | 22.0% | 34.9% |
| Gt4 | 1.3% | 0.0% |
| Gt1/2 | 0.0% | 0.4% |
| G1/3 | 0.0% | 1.1% |
| Unknown | 25.0% | 12.2% |
A comparison of these results with our previous study[32] revealed an impressive shift in the distribution of HCV genotypes, from the dominant prevalence of HCV Gt3a seen previously[31] to an increased prevalence of Gt1, particularly among IDUs, in the current study.
As shown in Table 3, most of the HCV-infected addicts belonged to the 31-40 age group and accounted for 35% of the infected addicts, followed by the 21-30, 41-50 and 51-60 age groups accounting for 25%, 19% and 12% infected addicts, respectively. These results indicate that unlike the study conducted in northern Iran[28], the addiction age may not have decreased in southern and southwest Iran.
According to Table 5, although all IDUs in this study had an addictive-drug use history of a few months to several years, 72% of them had more than one risk factor. More than 70% of these patients had a history of imprisonment of varying lengths. Tattoos were another crucial risk factor that was seen only in patients who had a history of imprisonment. Drinking alcohol and having multiple sex partners were other risk factors, however, there was little data on these risk factors.
| HCV infection | +Imprisonment and tattoo | +Imprisonment | + No other risk factor | Total |
| Injection Drug Users | 70 | 235 | 134 | 439 (80) |
| Non-Injection Drug Users | 63 | 23 | 25 | 111 (20) |
| Total | 133 (24) | 265 (48) | 159 (28) | 550 (100) |
We found that the genotype distribution also varied with respect to underlying conditions among patients from the same geographical area, and it is noteworthy that the least prevalent unknown genotypes were found among IDUs (Table 4), probably due to high mean quantitative viral load.
We would like to express our sincere thanks to Hassan Khajehei, PhD, for copy editing the manuscript. We would also like to thank Miss. Roosta and Statistics Center at Nemazi Hospital, Shiraz, for statistically analyzing the collected data.
From 2009 to 2012, hepatitis C virus (HCV) genotype patterns among different high-risk groups were determined using real-time PCR, a very sensitive and fast assay.
In this research, 888 HCV patients in different high-risk groups underwent genotyping. Genotype 1 was determined to be the predominant genotype in both non-addicts (thalassemia, hemophilia, and hemodialysis patients) and addicts, the most prevalent high-risk group.
This is the first report on HCV genotypes among HCV patients with different exposure categories, residing in south and southwest Iran, where genotype 1 was found to be the most frequent genotype. Compared to the results of the authors' previous study, they found that addiction age had increased. An important finding in this research was that the genotype pattern had shifted from 3 to 1, especially among addicts.
This study demonstrated changing HCV genotype patterns in Iran and it can serve as basic research for prospective studies on HCV genome variation. This study could also be helpful for health authorities and decision makers.
In the present study, authors examined HCV genotype in 888 Iranian patients by real-time PCR assay and evaluated the relation between infective root and HCV genotype.
P- Reviewer: Tamori A S- Editor: Qi Y L- Editor: Webster JR E- Editor: Liu XM
| 1. | Hellard M, Sacks-Davis R, Gold J. Hepatitis C treatment for injection drug users: a review of the available evidence. Clin Infect Dis. 2009;49:561-573. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 183] [Cited by in RCA: 197] [Article Influence: 12.3] [Reference Citation Analysis (0)] |
| 2. | Seeff LB, Hoofnagle JH. Appendix: The National Institutes of Health Consensus Development Conference Management of Hepatitis C 2002. Clin Liver Dis. 2003;7:261-287. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 128] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 3. | Kamal S. Genotypic variations around the world: Is hepatitis C virus evolving? Curr Hepat Rep. 2006;5:142-149. |
| 4. | Ramia S, Melhem NM, Kreidieh K. Hepatitis C virus infection in the Middle East and North Africa “MENA” region: injecting drug users (IDUs) is an under-investigated population. Infection. 2012;40:1-10. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 10] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 5. | Kassaian N, Adibi P, Kafashaian A, Yaran M, Nokhodian Z, Shoaei P, Hassannejad R, Babak A, Ataei B. Hepatitis C Virus and Associated Risk Factors among Prison Inmates with History of Drug Injection in Isfahan, Iran. Int J Prev Med. 2012;3:S156-S161. [PubMed] |
| 6. | Poordad F, Reddy KR, Martin P. Rapid virologic response: a new milestone in the management of chronic hepatitis C. Clin Infect Dis. 2008;46:78-84. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 87] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 7. | Nakatani SM, Santos CA, Riediger IN, Krieger MA, Duarte CA, Lacerda MA, Biondo AW, Carrilho FJ, Ono-Nita SK. Development of hepatitis C virus genotyping by real-time PCR based on the NS5B region. PLoS One. 2010;5:e10150. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 16] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 8. | Ashfaq UA, Javed T, Rehman S, Nawaz Z, Riazuddin S. An overview of HCV molecular biology, replication and immune responses. Virol J. 2011;8:161. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 104] [Cited by in RCA: 128] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 9. | Aceijas C, Rhodes T. Global estimates of prevalence of HCV infection among injecting drug users. Int J Drug Policy. 2007;18:352-358. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 237] [Cited by in RCA: 253] [Article Influence: 14.1] [Reference Citation Analysis (0)] |
| 10. | Mahfoud Z, Kassak K, Kreidieh K, Shamra S, Ramia S. Distribution of hepatitis C virus genotypes among injecting drug users in Lebanon. Virol J. 2010;7:96. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 36] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 11. | Hutchinson SJ, Roy KM, Wadd S, Bird SM, Taylor A, Anderson E, Shaw L, Codere G, Goldberg DJ. Hepatitis C virus infection in Scotland: epidemiological review and public health challenges. Scott Med J. 2006;51:8-15. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 55] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 12. | Cantaloube JF, Gallian P, Attoui H, Biagini P, De Micco P, de Lamballerie X. Genotype distribution and molecular epidemiology of hepatitis C virus in blood donors from southeast France. J Clin Microbiol. 2005;43:3624-3629. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 41] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 13. | Poynard T, Marcellin P, Lee SS, Niederau C, Minuk GS, Ideo G, Bain V, Heathcote J, Zeuzem S, Trepo C. Randomised trial of interferon alpha2b plus ribavirin for 48 weeks or for 24 weeks versus interferon alpha2b plus placebo for 48 weeks for treatment of chronic infection with hepatitis C virus. International Hepatitis Interventional Therapy Group (IHIT). Lancet. 1998;352:1426-1432. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1667] [Cited by in RCA: 1640] [Article Influence: 60.7] [Reference Citation Analysis (0)] |
| 14. | Su TH, Liu CH, Liu CJ, Chen CL, Ting TT, Tseng TC, Chen PJ, Kao JH, Chen DS. Serum microRNA-122 level correlates with virologic responses to pegylated interferon therapy in chronic hepatitis C. Proc Natl Acad Sci USA. 2013;110:7844-7849. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 55] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 15. | Wasley A, Grytdal S, Gallagher K. Surveillance for acute viral hepatitis--United States, 2006. MMWR Surveill Summ. 2008;57:1-24. [PubMed] |
| 16. | Astone JM, Strauss SM, Hagan H, Des Jarlais DC. Outpatient drug treatment program directors’ hepatitis C-related beliefs and their relationship to the provision of HCV services. Am J Drug Alcohol Abuse. 2004;30:783-797. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 15] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 17. | Kurelac I, Papic N, Sakoman S, Orban M, Dusek D, Coric M, Vince A. Intravenous Drug Users Can Achieve a High Sustained Virological Response Rate: experience From Croatian Reference Center for Viral Hepatitis. Hepat Mon. 2011;11:986-992. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 18. | Brunner N, Senn O, Rosemann T, Falcato L, Bruggmann P. Hepatitis C treatment for multimorbid patients with substance use disorder in a primary care-based integrated treatment centre: a retrospective analysis. Eur J Gastroenterol Hepatol. 2013;25:1300-1307. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 15] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 19. | Bojovic K, Simonovic J, Katanic N, Milosevic I, Pesic I, Delic D, Svirtlih N, Jevtovic DJ. The comparison of chronic hepatitis C treatment outcome between intravenous drug users and non-intravenous drug users. Biomed Pharmacother. 2013;67:517-520. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 4] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 20. | Nguyen LH, Nguyen MH. Systematic review: Asian patients with chronic hepatitis C infection. Aliment Pharmacol Ther. 2013;37:921-936. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 94] [Cited by in RCA: 95] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 21. | Kamal SM, Nasser IA. Hepatitis C genotype 4: What we know and what we don’t yet know. Hepatology. 2008;47:1371-1383. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 198] [Cited by in RCA: 207] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 22. | Kamal SM. Improving outcome in patients with hepatitis C virus genotype 4. Am J Gastroenterol. 2007;102:2582-2588. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 16] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 23. | Zhao L, Feng Y, Xia XS. [The different epidemic and evolution of HCV genotypes]. Yi Chuan. 2012;34:666-672. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 9] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 24. | Waqar M, Khan AU, Rehman HU, Idrees M, Wasim M, Ali A, Niaz Z, Ismail Z, Rehman MU, Tariq M. Determination of hepatitis C virus genotypes circulating in different districts of Punjab (Pakistan). Eur J Gastroenterol Hepatol. 2014;26:59-64. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 12] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 25. | Butt S, Idrees M, Akbar H, ur Rehman I, Awan Z, Afzal S, Hussain A, Shahid M, Manzoor S, Rafique S. The changing epidemiology pattern and frequency distribution of hepatitis C virus in Pakistan. Infect Genet Evol. 2010;10:595-600. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 40] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 26. | Kabir A, Alavian SM, Keyvani H. Distribution of hepatitis C virus genotypes in patients infected by different sources and its correlation with clinical and virological parameters: a preliminary study. Comp Hepatol. 2006;5:4. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 48] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 27. | Zarkesh-Esfahani SH, Kardi MT, Edalati M. Hepatitis C virus genotype frequency in Isfahan province of Iran: a descriptive cross-sectional study. Virol J. 2010;7:69. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 41] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 28. | Vossughinia H, Goshayeshi L, Bayegi HR, Sima H, Kazemi A, Erfani S, Abedini S, Goshayeshi L, Ghaffarzadegan K, Nomani H. Prevalence of Hepatitis C Virus Genotypes in Mashhad, Northeast Iran. Iran J Public Health. 2012;41:56-61. [PubMed] |
| 29. | Farivar TN, Nezam MK, Johari P. Genotyping of hepatitis C virus isolated from hepatitis patients in Southeast of Iran by Taqman Realtime PCR. J Pak Med Assoc. 2011;61:586-588. [PubMed] |
| 30. | Rafiei A, Darzyani AM, Taheri S, Haghshenas MR, Hosseinian A, Makhlough A. Genetic diversity of HCV among various high risk populations (IDAs, thalassemia, hemophilia, HD patients) in Iran. Asian Pac J Trop Med. 2013;6:556-560. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 10] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 31. | Samimi-Rad K, Nasiri Toosi M, Masoudi-Nejad A, Najafi A, Rahimnia R, Asgari F, Shabestari AN, Hassanpour G, Alavian SM, Asgari F. Molecular epidemiology of hepatitis C virus among injection drug users in Iran: a slight change in prevalence of HCV genotypes over time. Arch Virol. 2012;157:1959-1965. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 23] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 32. | Ziyaeyan M, Alborzi A, Jamalidoust M, Badiee P, Moeini M, Kadivar A, Prevalence of hepatitis C virus genotypes in chronic infected patients, southern Iran. Jundishapur J Microbiol. 2011;4:141-146. |