Published online Jun 25, 1996. doi: 10.3748/wjg.v2.i2.115
Revised: May 18, 1996
Accepted: June 10, 1996
Published online: June 25, 1996
AIM: To explore the relation between level of hepatitis C virus (HCV) viremia and HCV-related chronic liver disease.
METHODS: Serum HCV RNA was measured by competitive reverse transcription polymerase chain reaction (CRT-PCR) in 27 patients with chronic HCV infection.
RESULTS: Levels of serum HCV RNA were low (102-106 copies/50 μL serum) in patients with chronic HCV infection. Patients with chronic active hepatitis (105.739 ± 0.25 copies/50 μL serum) and with cirrhosis (105.803 ± 0.76 copies/50 μL serum) had higher levels of serum HCV RNA than patients with chronic persistent hepatitis (105.068 ± 1.04 copies/50 μL serum) (P < 0.05). There was a positive relation between levels of serum HCV RNA and alanine aminotransferase.
CONCLUSION: All of these results suggest that viremia level is low in chronic HCV infection. HCV itself plays an important role in progress of chronic liver disease, and HCV replication is related to liver damage.
- Citation: Zhang SL, Liang XS, Lin SM, Qiu PC. Relation between viremia level and liver disease in patients with chronic HCV infection. World J Gastroenterol 1996; 2(2): 115-117
- URL: https://www.wjgnet.com/1007-9327/full/v2/i2/115.htm
- DOI: https://dx.doi.org/10.3748/wjg.v2.i2.115
The pathogenesis of hepatitis C virus (HCV) infection is complex. Replication of HCV as well as host immune responses are involved in the liver cell damage that manifests in HCV infected individuals. Some studies have demonstrated that during the late stage of chronic liver disease, HCV still replicates actively[1]. We developed a competitive reverse transcription-polymerase chain reaction (CRT-PCR) for quantitative analysis of HCV RNA and investigated 27 patients with chronic HCV infection to explore the relation between HCV viremia level and chronic liver disease.
Twenty-seven patients (17 males and 10 females, aged 28-60 years) were hospitalized in our Department of Infectious Disease during the period from June 1993 to July 1995. All of these patients tested positive for anti-HCV and serum HCV RNA, and had shown abnormal levels of alanine aminotransferase (ALT) for at least 6 months. Thirteen of the cases were diagnosed as chronic persistent hepatitis (CPH), 10 as chronic active hepatitis (CAH) and 4 as cirrhosis, according to the criteria made at the National Conference on Viral Hepatitis, Shanghai, 1990. All 27 patients tested negative for hepatitis B surface antigen (HBsAg) and antibodies to the hepatitis A, D an E viruses (anti-HAV-IgM, anti-HDV IgM and anti-HEV IgM respectively). Serum samples were taken upon the first finding of HCV RNA positivity and stored at -20 °C until use.
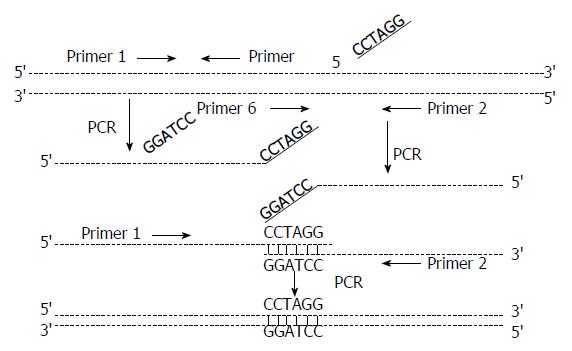
RNA was extracted from 50 μL of each serum sample by using the acid quanidium thiocyanate phenol-chloroform method. HCV cDNA was synthesized from each extract by reverse transcription using primer 2 (Table 1). A competitive DNA plate was constructed by recombinant PCR. The restriction enzyme target site recognized by BamH I (GGATCC) was inserted into the HCV cDNA products by using primers 1 and 5, and primers 2 and 6 (Figure 1)[2,3]. Quantitative detection of serum HCV RNA was then carried out. The HCV cDNA from each serum sample was added to the competitive plate DNA (103-5 copies), respectively, and amplified by nested PCR using primers 1 and 2, and primers 3 and 4. The DNA product was then cut with endonuclease BamH I, electrophoresed through a 1.5% agarose gel, and stained with ethidium bromide; the staining pattern included three bands of 286 bp, 208 bp and 78 bp. The DNA was isolated from the 286-bp and 208-bp sized bands, respectively, by electro-osmosis and the OD260 value was detected. The number of HCV RNA copies in the 50 μL serum sample was calculated according to the formula: C1 = OD1/OD2 × C2 × 1.2, where C1 was the number of HCV RNA copies in 50 μL serum sample, C2 was the number of competitive plate copies, OD1 was the OD260 value of DNA from the 286 bp band, OD2 was the OD260 value of DNA from the 208 bp band, and 1.2 was the transformation coefficient.
Anti-HCV was detected by second generation radioimmunoassay (RIA-2). HBsAg and IgM anti-HAV were detected by standard RIA. Anti-HDV and anti-HEV were detected by enzyme-linked immunosorbent assay.
The serum levels of HCV RNA in patients with chronic HCV infection were low (102-106 copies/50 μL serum).
The HCV RNA levels in patients with CAH and cirrhosis were much higher than those in patients with CPH (P < 0.05; Table 2).
| Clinical classification | Number of cases | HCV RNA, copies/50 μL serum |
| CPH | 13 | 105.068 ± 1.04 |
| CAH | 10 | 105.739 ± 0.25a |
| Cirrhosis | 4 | 105.803 ± 0.76 |
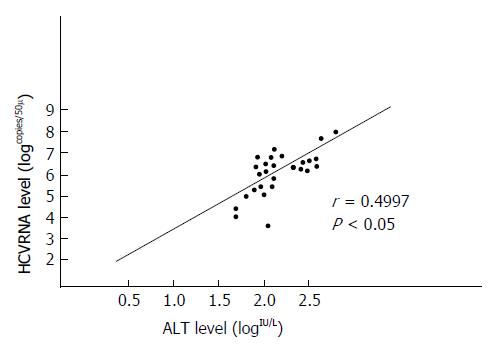
There was a positive correlation between levels of serum HCV RNA and ALT (r = 0.4997, P < 0.05; Figure 2).
In chronic HCV infection the viremia level is low. Bradley[4] conducted a chimpanzee infection experiment and found that serum titer of HCV was 102-104 CID/mL. A CID/mL of 106.5 equates to 4 × 108 copies of HCV RNA/mL in serum[4]. Ulrich et al[1] used end-point dilution PCR to show that serum HCV RNA levels in patients with chronic hepatitis C were 102-5× 107 copies/mL. Hagiwara et al[5] used CRT-PCR to demonstrate that the serum HCV RNA was 104-109copies/mL. We obtained similar results in this study, detecting 2 × 103-2 × 107 copies/mL by the CRT-PCR technique. However, Jenison et al[6] demonstrated that the serum HBV DNA level in patients with chronic hepatitis B was 5 × 108-3 × 1010 copies/mL by dot blot hybridization, which is much less sensitive than PCR. It can be postulated that the viremia level in chronic HCV infection is significantly lower than that in HBV infection.
Replication of HCV is related to the progress of chronic liver disease. Our results show that serum HCV RNA levels in patients with CAH and cirrhosis are much higher than that in patients with CPH (P < 0.01). A similar result was reported by Kato et al[7], from a study in which they used CRT-PCR to detect HCV RNA in 36 patients with chronic liver diseases; specifically, the serum levels in patients with CAH (105.6 ± 1.6 copies/50 μL) and with cirrhosis (106.0 ± 1.6 copies/50 μL) were higher than that in patients with CPH (103.3 ± 1.9copies/50 μL) (P < 0.01)[7]. Gordon et al[8] demonstrated that viremia level is positively related to liver histological score in patients with chronic HCV infection. Tsutsumi et al[9] studied the expression of NS5 antigen related to RNA-dependent RNA polymerase in liver, and found that the detection rate of NS5 antigen in patients with cirrhosis was higher than that in patients with CAH and CPH. Collectively, these data suggest that HCV itself plays an important role in the progress of HCV-related chronic liver disease.
HCV is known to replicate actively during the course of chronic HCV infection, and to replicate at high level even in the late stage of the disease. Moreover, HCV replication is well recognized as closely related to liver damage. It was demonstrated in this study that there is a positive correlation between the levels of serum HCV RNA and ALT in patients with chronic HCV infection (r = 0.4997, P < 0.05). Kurasaki et al[10] detected serum HCV RNA in 41 patients with chronic HCV infection by using PCR. Subgroup analysis of the 41 patients, by high-level HCV RNA (showing positive test results on the first amplification) and low-level HCV RNA (showing positive test results on the second amplification), indicated that the ALT in the high-level group was much higher than that in the low-level group (191.7 ± 103.4 μkat/L vs 98.4 ± 66.7 μkat/L, P < 0.05); additionally, all the patients showing ALT > 166.7 μkat/L belonged to the high-level HCV RNA group[10], suggesting that HCV may have a cytopathologenic effect.
In conclusion, the results of the current study demonstrate that the viremia level is low in chronic HCV infection, indicating that HCV replication is related to the progress of chronic liver disease and supporting the notion that HCV may have a cytopathologenic effect.
Original title:
S- Editor: Filipodia L- Editor: Jennifer E- Editor: Zhang FF
| 1. | Ulrich PP, Romeo JM, Lane PK, Kelly I, Daniel LJ, Vyas GN. Detection, semiquantitation, and genetic variation in hepatitis C virus sequences amplified from the plasma of blood donors with elevated alanine aminotransferase. J Clin Invest. 1990;86:1609-1614. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 125] [Cited by in RCA: 132] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 2. | Higuchi R, Krummel B, Saiki RK. A general method of in vitro preparation and specific mutagenesis of DNA fragments: study of protein and DNA interactions. Nucleic Acids Res. 1988;16:7351-7367. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1869] [Cited by in RCA: 2026] [Article Influence: 54.8] [Reference Citation Analysis (0)] |
| 3. | Ho SN, Hunt HD, Horton RM, Pullen JK, Pease LR. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene. 1989;77:51-59. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5970] [Cited by in RCA: 6343] [Article Influence: 176.2] [Reference Citation Analysis (0)] |
| 4. | Bradley DW. The agents of non-A, non-B viral hepatitis. J Virol Methods. 1985;10:307-319. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 40] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 5. | Hagiwara H, Hayashi N, Mita E, Naito M, Kasahara A, Fusamoto H, Kamada T. Quantitation of hepatitis C virus RNA in serum of asymptomatic blood donors and patients with type C chronic liver disease. Hepatology. 1993;17:545-550. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 137] [Cited by in RCA: 133] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 6. | Jenison SA, Lemon SM, Baker LN, Newbold JE. Quantitative analysis of hepatitis B virus DNA in saliva and semen of chronically infected homosexual men. J Infect Dis. 1987;156:299-307. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 74] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 7. | Kato N, Yokosuka O, Hosoda K, Ito Y, Ohto M, Omata M. Quantification of hepatitis C virus by competitive reverse transcription-polymerase chain reaction: increase of the virus in advanced liver disease. Hepatology. 1993;18:16-20. [PubMed] |
| 8. | Gordon SC, Kodali VP, Silverman AL, Dmuchowski CF, Urdea MS, Chan CS, Wilber JC. Levels of hepatitis C virus RNA and liver histology in chronic type C hepatitis. Am J Gastroenterol. 1994;89:1458-1461. [PubMed] |
| 9. | Tsutsumi M, Urashima S, Takada A, Date T, Tanaka Y. Detection of antigens related to hepatitis C virus RNA encoding the NS5 region in the livers of patients with chronic type C hepatitis. Hepatology. 1994;19:265-272. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 23] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 10. | Kurosaki M, Enomoto N, Sato C, Sakamoto N, Hoshino Y, Haritani H, Marumo F. Correlation of plasma hepatitis C virus RNA levels with serum alanine aminotransferase in non-A, non-B chronic liver disease. J Med Virol. 1993;39:246-250. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 23] [Article Influence: 0.7] [Reference Citation Analysis (0)] |