INTRODUCTION
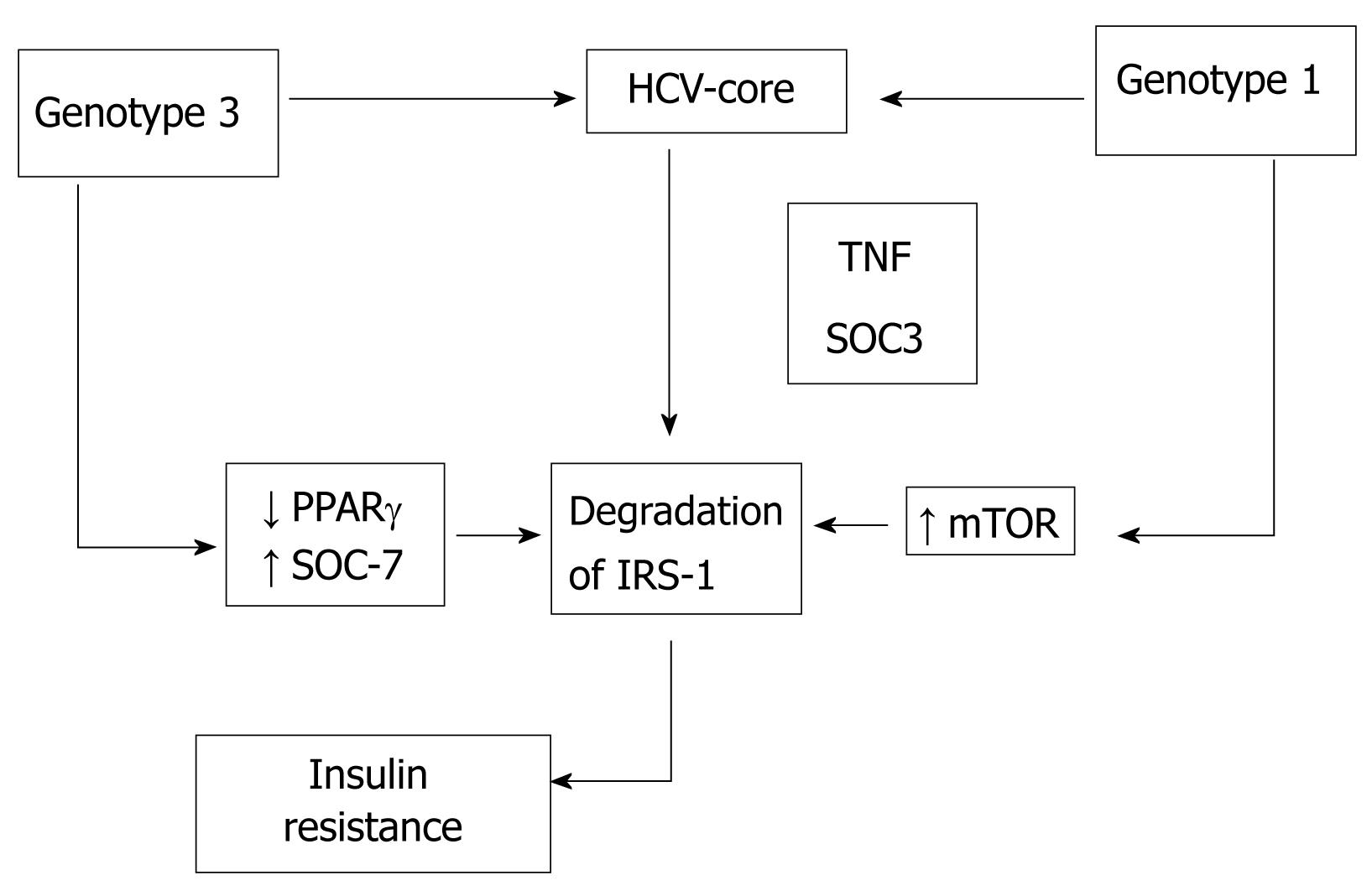
Figure 1 Hepatitis C core induces insulin resistance promoting proteasomal degradation of insulin receptor substrate-1 (IRS-1).
Hepatocyte steatosis, defined as accumulation of fat droplets in hepatocytes, is a histological feature of a group of liver diseases including not only metabolic or alcoholic disorders but also chronic hepatitis C virus (HCV) and drug-induced liver disease[1,2]. Steatosis is a very common lesion in chronic HCV, seen in more than half of patients, with a prevalence of 40% to 86% according to the genotype. The majority of patients show simple steatosis, but features of non-alcoholic steatohepatitis have been found in approximately 10% of patients with chronic HCV[3]. Two main types of steatosis have been defined in HCV: metabolic steatosis found in patients infected by genotype 1 and associated with metabolic syndrome, and viral steatosis reported in patients with genotype 3a, without other known steatogenic cofactors and directly linked to a cytopathic effect of the virus. In the same way, chronic HCV infection can also induce insulin resistance[4,5]. Epidemiological data support an association between HCV infection and a risk for the development of type 2 diabetes mellitus, both in cross-sectional and longitudinal cohorts. The majority of transgenic mice expressing the HCV core protein developed insulin resistance and type 2 diabetes. Indeed, patients with HCV showed a higher insulin resistance index than healthy controls or patients with liver diseases other than HCV matched by sex, body mass index and age. Insulin resistance could be promoted by viral proteins and the intrinsic mechanism seems to be genotype-specific[5]. In this sense, HCV replicons - genomic or subgenomic constructs expressing the viral replicase complex and capable of autonomous viral replication - constitute a powerful tool to investigate the molecular mechanism leading to steatosis and insulin resistance in host cells.
In this review, we aim to analyse the role of HCV in the pathogenesis of steatosis and insulin resistance as a possible “way out” mechanism of the virus, together with the impact of both metabolic abnormalities on the clinical course of the disease.
PATHOGENESIS OF STEATOSIS IN HEPATITIS C
HCV infection is characterized by a high rate of progression to fibrosis, chronic hepatitis, leading to cirrhosis and ultimately to hepatocellular carcinoma. In addition, various observations suggest that hepatic steatosis is a common histological feature of chronic HCV infection. Furthermore, increasing evidence indicates that hepatic steatosis is a more vulnerable factor that leads to liver inflammation and fibrosis. These suggest that HCV has a direct role in the development of steatosis and/or that the presence of steatosis affects the progression of HCV-related liver disease. The core protein component of HCV is known to contribute to hepatic steatosis[6], hepatic fibrosis, and hepatic carcinogenesis[7]. Some studies suggest that HCV core protein causes hepatic steatosis through inhibition of microsomal triglyceride transfer protein (MTP) activity and very low density lipoprotein (VLDL) secretion, and impairment of the expression and transcriptional activity of peroxisome proliferator-activated receptor (PPAR)α[8]. Sterol regulatory element binding protein 1s (SREBP1s) belong to the basic helix-loop-helix-leucine zipper family of transcription factors. A key role of two SREBP1 isoforms in regulating fatty acid synthesis in liver is suggested by study of transgenic mice overexpressing the constitutively active mature forms of SREBP1 isoforms[9]. The transgenic mice study suggests that SREBP1s increase the transcription of genes involved in hepatic fatty acid synthesis [including fatty acid synthase (FAS); acetyl-CoA carboxylase (ACC); and stearoyl-CoA desaturase (SCD)], inducing massive hepatic steatosis through increased accumulation of triglycerides.
PPARγ is a transcription factor, belonging to the nuclear receptor superfamily, is a master regulator of adipocyte differentiation, and is important in regulation of a number of genes involved in fatty acid and glucose metabolism[10]. There is evidence suggesting that liver PPARγ increases the transcription of genes involved in hepatic fatty acid synthesis (including FAS, ACC and SCD) and fatty acid uptake (including FAT/CD36 and fatty acid translocase). Thus, liver PPARγ contributes to regulation of lipid synthesis, transport and storage within hepatocytes, causing the development of hepatic steatosis.
Using a cell culture based model, Kim and coworkers[11] have shown that HCV core protein is able to induce the gene expression and transcriptional activity of SREBP1, thereby causing the increase of fatty acid synthesis. They also observed that HCV core protein elevates PPARγ activity, inducing the expression of fatty acid uptake-associated gene. These results suggest that SREBP1 and PPARγ may represent a new potential therapeutic target in the pathogenesis in HCV infection.
Steatosis development is due to both viral and host factors. Viral steatosis is mostly reported in patients with genotype 3a, in whom fat accumulation correlates with HCV replication levels in serum and liver and disappears after successful antiviral therapy, strongly suggesting a direct role of specific viral products in the fat deposition. To address the role of specific HCV genotype on lipid accumulation in cells, Abid and coworkers[12] developed an in vitro model to study the effect of the core protein belonging to several viral genotypes (1b, 2a, 3a, 3h, 4h and 5a). They concluded that the pattern observed in Huh7 cells upon expression of the six core proteins largely corroborated the phenotype seen in vivo. The genotype 3a-derived core protein was about three-fold more efficient than the corresponding protein from genotype 1b at inducing triglycerides accumulation in transfected cells, and is proposed as the ideal candidate to study the pathogenesis of HCV-induced steatosis. This group also reported the role of PPARγ expression on triglyceride accumulation in Huh7 cells transfected with genotypes 1b and 3a core proteins[13]. They found that the expression of HCV 3a core protein was associated with an increase in triglyceride accumulation and with a significant reduction of PPAR-γ mRNA compared with HCV 1b. Moreover, Fukasawa and coworkers[14] showed that ACC1 and FAS, enzymes responsible for de novo lipid biosynthesis, are induced in Huh7 cells transfected with HCV core protein. Using microarray analysis Pazienza et al[15] compared the gene expression profile of Huh7 cells transfected with the core protein of HCV genotype 1b and 3a, leading to the conclusion that several genes involving lipid transport and metabolism were up- or down-regulated in a genotype-specific manner. This fact could explain the variable disease expression associated with HCV infection. Taking all these observations together, we can conclude that there is a direct link between virus infection and steatosis development.
It has been shown that genotype 3 infected patients had lower expression of PPARα or MTP mRNA in the liver and fat accumulation was three times higher in comparison with non-genotype 3 patients[16]. Although the mechanism by which genotype 3a induces steatosis more efficiently than others genotypes is not completely understood, some differences in the amino acid sequence in the core protein could explain, at least in part, these differences. Indeed, a specific polymorphism in core protein from genotype 3 has been associated with lipid accumulation in hepatocytes. Amino acid substitution at positions 182 and 186 caused intracellular lipid accumulation in hepatic cells and contributed to steatosis development[17]. Indeed, single polymorphisms in the core gene promoting an amino acid change from tyrosine to phenylalanine (Y164F) has been associated with greater cumulative lipid droplet area in cultured cells than in cells producing the wild-type core protein[18].
The immune response against HCV releases reactive oxygen species (ROS) from sequestered phagocytes and activated Kupffer cells in the liver. Increased oxidative stress caused by HCV may result in the activation of Kupffer cells[19]. The change in H+ concentration alters the balance in the Na+/H+ exchanger, causing the Kupffer cells to swell and eventually burst. This releases ROS and an arsenal of inflammatory mediators such as TNF-α, TGF-β, IL-6 and IL-8. The rising concentrations of ROS induce lipid peroxidation and damage triglycerides. The process of lipid peroxidation disrupts cellular membranes and can induce mitochondrial dysfunction[20].
Metabolic and viral steatosis share the same pathophysiological pathways, although metabolic abnormalities are more often seen in genotype 3a resulting in greater steatosis. A predominant but not exclusive metabolic or viral mechanism could be associated with each genotype, and steatosis could appear as a consequence of the interplay between both host and viral factors.
Recently, steatosis has been implicated in viral replication. The disruption of the association between core protein and fat droplets impairs viral fitness[21]. In infected cells, core protein is targeted to lipid droplets, which serve as intracellular storage organelles. According to their results, two facts show the relationship between lipids droplets and viral replication: first, the change in the distribution of the core protein from wild sites juxtaposed to lipids droplets at an early stage, which later agreed with a peak production of virus. Secondly, JFH1DP, a mutant strain obtained from JFH1, which did not give rise to virus progeny, expressed a core protein that was targeted to punctuate sites indistinguishable from those identified for the wild type protein at early times, but JFH1DP core did not proceed to coat lipid droplets. In such cases, an association between core protein and lipid droplets would be essential for virus assembly. Furthermore, alteration of a phenylalanine in domain 2 of the core protein generates an unstable form of protein associated with reduced replication rates. Lastly, Shavinskaya et al[22] identified the lipid droplet binding domain of HCV core as the major determinant for efficient virus production. They show that D2 in HCV core is a critical determinant for efficient virus assembly and that small numbers of variations (mutations) in this highly conserved domain can exert a significant effect on production of infectious HCV. Thus, HCV promotes steatosis as an efficient mechanism for stable viral replication.
From a clinical perspective, the negative impact of severe hepatic steatosis on graft dysfunction during the immediate post-transplant period has long been recognized[23]. Donor livers containing greater than 30%-50% steatosis are at increased risk of developing primary nonfunction and delayed function and are associated with reduced graft and patient survival. Since chronic HCV is the most common indication for liver transplantation (LT), several studies have examined the impact of steatosis within the donor graft on the severity of recurrent HCV and/or survival following LT[24,25]. From those studies, a direct relationship between marginal donors, graft steatosis and more frequent and earlier recurrence for HCV-related cirrhosis has been established[25]. The putative relationship between steatosis and viral replication could explain this fact.
INSULIN RESISTANCE IN HEPATITIS C
Liver fibrosis has been considered for a long time to be responsible for the appearance of insulin resistance and type 2 diabetes in patients with chronic liver diseases. Hyperinsulinemia in liver cirrhosis has been reported to be due to diminished hepatic insulin extraction by liver dysfunction and not to pancreatic hypersecretion. C-peptide (a peptide resulting from the split of proinsulin into insulin and C-peptide) and insulin are secreted in equimolar quantities, and more than 50% of insulin is degraded in the liver at first pass, whereas C-peptide is degraded in the kidneys. Simultaneous measurements of C-peptide and insulin revealed that both insulin resistance and insulin secretion contribute to glucose intolerance in patients with chronic HCV[26]. From a clinical point of view, the insulin resistance index was higher in patients with chronic HCV showing mild or no fibrosis than matched healthy controls. Moreover, insulin resistance was found to be higher in patients with chronic HCV than patients with other causes of chronic hepatitis matched by age, sex, body mass index, family history of type 2 diabetes and fibrosis staging[27]. On the other hand, Homeostasis Model Assessment of Insulin Resistance (HOMA-IR) correlated with HCV RNA level and was found to be higher in patients with HCV than healthy controls in spite of a lower body mass index. Indeed, in patients with HCV, insulin resistance is more strongly associated with viral load than visceral obesity[4]. Insulin resistance is a common metabolic disorder in the pre-diabetic state. Thus, if HCV promotes insulin resistance it should be linked to type 2 diabetes. Indeed, diabetes was more often seen in HCV than other liver diseases (20%-25% in patients with HCV and in 10% of those with hepatitis B)[28]. Moreover, in a recent systematic review an increased risk of type 2 diabetes mellitus in patients with hepatitis C in comparison with non-infected people has been reported[29].
HCV seems to cause metabolic syndrome, but the mechanisms by which HCV promotes insulin resistance have not been completely understood[30]. HCV induces several complex mechanisms that lead to oxidative stress, insulin resistance, steatosis, fibrosis, apoptosis, altered gene expression and hepatocellular carcinoma. HCV seems to lead to insulin resistance through interference of intracellular insulin signalling by HCV proteins, mainly the serine phosphorylation of IRS-1 and impairment of the downstream Akt signalling pathway[31].
Transgenic mice expressing core HCV protein developed insulin resistance, which does not occur in wild-type animals[32]. During HCV replication, core protein has been found to localize to the outer mitochondrial membrane and is also associated with the endoplasmic reticulum (ER)[33]. In the mitochondria, HCV core protein induces mitochondrial permeability transition, calcium accumulation, stimulation of electron transport and ROS production, as well as promoting glutathione depletion and release of cytochrome C[34]. Moreover, HCV proteins are assembled and correctly folded by chaperones in the ER, but in some circumstances the ER fails to export synthesized proteins properly leading to an accumulation of misfolded proteins[35]. The misfolded protein response causes ER dysfunction and promotes inflammation and ER stress[36]. Of the HCV nonstructural proteins, NS3 and NS5A act as key mediators in the induction of oxidative stress and inflammation. The association of NS5A with the ER has been suggested to stimulate mitochondrial ROS production by releasing calcium from the ER[37]. In addition, NS3 has been shown to activate nicotinamide adenine dinucleotide phosphate (NADPH) oxidase 2 (Nox2) that generates ROS[38]. Nox2 can nitrosylate proteins within cells and can lead to a large number of pathological processes. Consequently, a role for NOX enzymes in hepatic fibrosis, characterized by hepatic stellate cell (HSC) proliferation and accumulation of extracellular matrix proteins, has been suggested[39,40].
HCV core protein inhibits PPAR α and γ expressed in hepatocytes and adipocytes promoting IRS-1 degradation and insulin resistance[13]. HCV core protein induces the over production of TNFα, responsible for phosphorylation of serine residues of IRS-1 and IRS-2 and down-regulation of glucose transporter gene expression. TNF correlates with the hyperinsulinemic state and the blockade of TNF production by anti-TNF drugs like infliximab inhibits the development of insulin resistance. Thus, TNF promotes hyperinsulinemia and hyperglycaemia and has been linked to an increased risk of diabetes development[41]. Moreover, non-structural proteins such as NS3 and NS5 interact with the ER. NS3 enhances Nox2 activity and increases nitrosylated proteins and ROS[42]. NS5A and NS5B proteins activate toll-like receptor-4 and the NFΚB pathway enhances TNF and IL-6 production and promotes insulin resistance[43]. IL-6 is a cytokine that is secreted from Kupffer cells, adipocytes, B cells, and hepatocytes. HCV-infected patients are known to have elevated levels of IL-6 due to the virus-induced inflammatory state[44]. Increased IL-6 derived from adipocytes leads to an ongoing acute-phase response that acts on hepatocytes and promotes hepatic insulin resistance. IL-6 is able to inhibit the expression of LPL in mice[45]. Unlike TNF-α, IL-6 circulates at high levels in plasma, perhaps representing a hormonal role of IL-6 that may induce insulin resistance in other tissues besides liver.
The HCV core protein interferes with in vitro insulin signalling by genotype-specific mechanisms (Figure 1). Pazienza et al[46] used the previously described model of transient expression of the core protein of different genotypes of HCV to assess the interaction of HCV genotype 3a with the insulin signaling, using as comparison the genotype 1b. They found that Insulin Receptor-1 (IRS-1) protein level was significantly reduced in Huh-7 cells expressing the core protein of both genotypes 3a and 1b. However, while the core protein of genotype 3a promoted IRS-1 degradation through the down-regulation of PPARγ and by up-regulating the suppressor of cytokine signal 7 (SOCS-7), the core protein of genotype 1b activated the mammalian target of rapamycin (mTOR), demonstrating that interaction between viral core protein and IRS-1 degradation is genotype-specific.
Enhanced SOCS production after HCV infection seems to play a crucial role in inducing interferon resistance by inhibiting interferon-α intracellular signalling[47]. Moreover, SOCS inhibit the phosphorylation of Akt and phosphatidyl inositol 3 kinase (PI3K), impairing intracellular insulin signalling, blocking the transactivation of GLUT-4 and avoiding glucose uptake by cells. Over-expression of SOCS-3 has also been linked to interferon and insulin resistance[48]. Indeed, in transgenic mice unable to express SOCS-3 and expressing HCV core protein, insulin resistance did not appear in the absence of SOCS-3[49]. Therefore, as previously reported in steatosis, patients with HCV could suffer from viral or metabolic insulin resistance. HCV itself induces insulin resistance by several factors also implicated in interferon resistance, allowing the virus to resist antiviral treatment and to promote fibrosis progression[50]. However, some controversial results support the presence of additional mechanisms in the development of insulin resistance in patients with chronic HCV and metabolic or viral type insulin resistance. Pazienza and coworkers[46] have shown that SOCS-1 and SOCS-3 mRNA levels did not change following transfection with both core proteins from genotypes 1b and 3a. However, SOCS-7 mRNA levels were found to be significantly higher in cells expressing the core protein 3a (but not in those transfected with core protein 1b). Their results were corroborated at the protein level by immunoblot. The role of SOCS-7 in IRS-1 downregulation in genotype 3a-transfected cells was confirmed using siRNA. They found that the mechanism of IRS-1 degradation by genotype 3a seems to be quite different from that possessed by genotype 1b. The latter is apparently mediated neither by PPARγ or SOCS-7 nor by SOCS-1 or SOCS-3, as suggested by other authors.
In chronic HCV infection, steatosis up-regulates hepatocyte CD95/Fas and thus increases apoptosis, which facilitates inflammation and fibrosis[51]. It has been recently shown that adiponectin reduced FFA-induced CD95/Fas expression and apoptosis of HepG2 hepatoma cells, which suggests a protective role for this hormone with promising therapeutic implications[52].
There are still many molecular mechanisms and open questions to uncover on the HCV-host interaction for the development of effective drugs for the treatment of this disease. In this regard, cell culture based replication systems described so far will help in this task. The availability of systems for replication of all known HCV genotypes together with animal models is highly desirable, in order to find out the importance of the virus genome in disease development.
CONCLUSION
Steatosis development is linked to HCV infection. There is evidence for the accumulation of lipids in the infected cell that could play a determinant role for efficient virus assembly. Thus, HCV promotes steatosis as an efficient mechanism for stable viral replication. HCV itself induces insulin resistance by several factors also implicated in interferon resistance, allowing the virus to resist antiviral treatment and to promote fibrosis progression.
Peer reviewers: Nahum Méndez-Sánchez, MD, PhD. Departments of Biomedical Research, Gastroenterology & Liver Unit, Medica Sur Clinic & Foundation, Puente de Piedra 150, Col. Toriello Guerra, Tlalpan 14050, Mexico City, Mexico; Frank J Burczynski, Faculty of Pharmacy, University of Manitoba, 50 Sifton Road, Winnipeg, Manitoba, R3T 2N2, Canada
S- Editor Tian L L- Editor O’Neill M E- Editor Ma WH