INTRODUCTION
Hepatitis C virus (HCV) genome has only one open reading frame, which encodes for a precursor protein polymer containing 3000 amino acids. The protein products, including core, NS3, NS4B and NS5A, have an effect on hepatocellular carcinoma (HCC) induced by HCV. The non-structural protein 3 (NS3) segment plays a key role in replication of virus and is closely related with the growth of host cells. NS3 has attracted attention in many research fields of HCV. In this experiment, we constructed the eukaryotic expression vector containing HCV NS3 segment, which could express NS3 protein in normal human hepatocytes in vitro.
MATERIALS AND METHODS
Enzymes and reagents
PCR primers were synthesized by Shanghai Sangon Biological Engineering and Technology and Service (Shanghai, China). Restriction endonucleases EcoRI and BamHI, T4 DNA ligase, RNase A and Ex Taq DNA polymerase and markers DL 15 000 and DL 2000 were purchased from TaKaRa Biological Technology (Dalian, China) . Protein radiant markers were purchased from Bioin Technology (Zhuhai, China). Plasmid purification kit and transmaster were purchased from BoLi Bioin Technology (Beijing, China). Anti-HCV NS3 monoclonal antibody was purchased from Biodesign(American). Donkey anti-sheep IgG-HRP was purchased from Kangchen Bio-tech (Shanghai, China). Immuno histochemical SP-HRP kit was purchased from Dingguo Biological Technology (Beijing, China). Western blot ECL kit was purchased from Bioin Technology (Zhuhai, China).
Vectors and experimental cells
Recombination plasmid pBRTM/HCV1-3011 containing the whole genome of HCV was donated by Professor Rice(Washington, USA). Expression vector pcDNA3.1(-) containing CMV promoter ahead of multiple cloning sites, replication initiating site SV40ori as well as anti-ampicillin and anti-G418 sites, was donated by Doctor Shun Fenyong (College of Life Science and Technology , Jinan University). pMD 18-T simple vectors were purchased from TaKaRa Biological Technology (Dalian, China). E.coli JM109 was provided by Biochemistry Department Jinan University. HL-7702 cells were purchased from Shanghai Institute of Cell Biology (Shanghai, China).
Experimental groups
Non-transfected HL-7702, HL-7702 transfected with pcDNA3.1 (-)/NS3 and HL-7702 transfected with pcDNA3.1(-) .
Construction of expression vector pcDNA3.1 (-)/NS3
Based on published sequences, the primers [5’-GAA TTC ATG GCG CCC ATC ACG GCG TAC GC-3’ (upriver) and 5’-CTG GAC CTC CAG CAG TGC ATT CCT AGG-3’ (downriver)] were used to amplify the whole sequence of HCV NS3 region. Thirty cycles of PCR were performed at 94 oC for 60 s, at 55 oC for 30 s, at 72 oC for 120 s, and at 72 oC for 10 min in 50μl reaction mixture containing 0.3 μL (1 U/μL) Taq polymerase, 5 μL10 ×PCR buffer,1 μL dNTPs (20 mmol/L), 0.625 μL each primer (20 μmol/L), 1μL DNA as template, and 41.45 μL distilled water. The PCR products were subjected to electrophoresis on 0.8% agarose gel for 60 min(voltage:80V),visualized by DNA viewer staining. The PCR products were connected to clone vector pMD 18-T after purification. Plasmid pMD 18-T/NS3 was transferred into E. coli with calcium chloride. The E.coli was cultured to amplify the plasmid. Successfully transferred E.coli was screened at 50 mg/L ampicillin. Then the constructed pMD 18-T/NS3 was recombined with expression vector pcDNA3.1(-)/NS3, thus the eukaryotic expression plasmid pcDNA3.1(-)/NS3 came into being .
Cell culture and growth curve
HL-7702 cells were cultured and passaged in RPMI 1 640 medium supplemented with 10% fetal calf serum in an incubator containing 50 mL/L CO2 at 37 oC Cells were digested by 0.25% trypsin and seeded (2.5×104 cells per well) in 48-well culture plates. Cells were digested and counted at an interval of 24h with the average number of cells in 3 wells calculated per time.The detection was continued for 8 d. Then the population doubling time was calculated and growth curve of the cells was plotted.
Immunohistochemical staining
We used S-P method to detect the expression of HCV NS3 protein in HL-7702 cells transfected with plasmid pcDNA3.1 (-)/NS3. Cells transfected with and without plasmid pcDNA3.1 (-) served as negative and blank control groups. Cells were cultured in 25-flasks till 85% convergence. The cells were washed three times with pre-cooled PBS before harvested. Then 1 milliliter lysis reagent containing 50 mmol/L TrisHCl (pH8.0),150 mmol/L NaCl,0.02% sodium ajide, 0.1%SDS, 100 mg/LPMSF(dissolved in isopropyl alcohol, 1 mg/L aprotinin,1%Nonidet P-40, 0.5% sodium deoxycholate) was added and incubated at 4 oC for 20 min. Then the lysis reagent mixed with broken cells was harvested and centrifuged at 12 000 r/min for 5 min. Twenty microliters of supernatant protein was resolved by SDS-8% polyacrylamide gel electrophoresis. After electrophoresis, the proteins were transferred onto a PVDF membrane, blocked in 5% skimmed milk for 1h, probed with monoclonal antibody to HCV NS3(1:1000) overnight at 4 oC. After washed three times with TBS containing 0.1% Tween 20, the membrane was treated with horseradish peroxidease-conjugated donkey anti-goat antibody for 3 h. Protein binding was detected by chemiluminescence reagent (ECL). Then the bands on X-film were assayed by densitometric scanning.
RESULTS
Construction of expression vector pcDNA3.1(-)/NS3
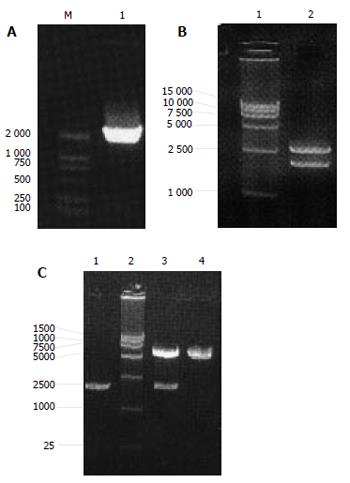
The PCR product of NS3 had correct molecule weight. Recycled PCR fragments were connected to T vector, E.coli was used to amplify the recombined vector pMD 18-T/NS3. Plasmid extracted from the positive clones could be digested by EcoRI and BamHI to two fragments (3000bp and 1900kb) as expected. DNA sequence analysis confirmed that the inserted fragment was a HCV NS3 segment containing 1893 nucleotides with right direction and reading frame. Initial codon, terminal codon and restriction sites of EcoRI , BamHI were successfully added to the ends. Vector pMD 18-T/NS3 was successfully recombined with plasmid pcDNA3.1(-). The length of the recombined vector pcDNA3.1(-)/NS3 was 7 300bp. After digested by EcoRI and BamHI, pcDNA3.1(-)/NS3 was cloven into two fragments (5 400 bp and 1900 kb) as expected (Figure 1A-1C).
Figure 1 PCR results of NS3 gene(A), pMD 18-T/NS3 (B) and pcDNA3.
1(-)/NS3 (C) digested with Eco and BamHI.
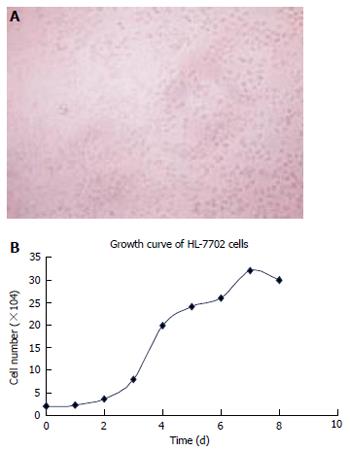
The incubation condition of HL-7702 cells and the cell growth curve were obtained (Figure 2 A, 2B)
Figure 2 Convergence (A) and growth curve (B) of HL-7702 cells.
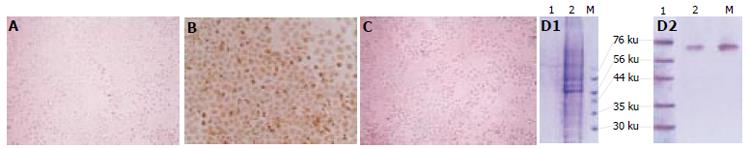
Immunohistochemical analysis showed that the blank control group was negative, pcDNA3.1 (-)/NS3 transfection group was positive for HCV NS3 protein in cytoplasm. SDS-PAGE analysis showed that among the separated protein bands (distilled from cytoplasm), a band was found at molecular weight 70 000 as expected (Figure 3A-3D)
Figure 3 Immunohistochemical staining of NS3 protein expressed in non-transfected (A), transfected with pcDNA3.
1(-)/NS3 (B), pcDNA3.1(-) (C) HL-7702 cells SDS-PAGE and Western blot as well as of NS3 protein expression in HL-7702 cells (D1-D2).
DISSCUSION
Hepatitis C virus(HCV) , the major etiological agent of non-A non-B hepatitis identified by the end of 1 980s, has infected more than 170 million persons worldwide and 85% of them could develop chronic hepatitis, liver function failure or hepatocellular carcinoma. Among all the gene segments of HCV, NS3 segment has many important physiological functions. Genetic variability of NS3 protein lies in different kinds of HCV-infected patients[2]. Nowadays little has been known about the nonstructural protein of HCV. NS3 protein contains many enzymes such as helicase , NTPase and serine protease, which can degrade the nonstructural segments of HCV polyprotein[3-5]. HCV core, NS3, NS5A and NS5B proteins can modulate cell proliferation independently by expressing p53[6]. Chronic hepatitis C induced by HCV could do great harm to human body, yet little has been known about the pathogenetic mechanism. HCV can impair the innate immunity system. It was reported that interaction between HCV NS3 protein and host TBK1 protein leads to inhibition of cellular antiviral responses[7] . Conserved C-terminal threonine of HCV NS3 can regulate autoproteolysis[8]. NS3 protein can down-regulate activities of protease peptide which could interfere with the activity of viral antigens, which may protect HCV from host immune surveillance, leading to persistent viral infection[9]. Besides, NS3/4A- mediated cleavage of TIR domain-containing adaptor protein potentially limits expression of multiple host defense genes, promoting persistent infections with this medically important virus [10].
Dolganiuc et al [11] showed that core protein and nonstructural protein 3 of HCV can induce production of IL-10 and mediate immunosuppression of against HCV. Some HCV proteins, such as core protein, NS3 and NS5A, not only regulate cell growth and interact with other cellular proteins, but also participate in programmed cell death under certain circumstances[12] . HCV can interfere with JAK/STAT signal pathway, leading to IFN production by down-regulating the expression of STAT1[13]. Additionally , NS3 can bind to a host protein (Sm-D) which interferes with the formation of small nuclear ribonuclear protein (snRNP) complexes ,so that occurrence of auto immune disease can be reduced[14].
It was reported that nonstructural protein 3 of HCV can inhibit dendritic cell differentiation by inducing pro- and anti-inflammatory cytokines [15]. Core, NS3, NS5A, NS5B proteins of hepatitis C virus can induce apoptosis of mature dendritic cells[16] and inhibit cellular immune responses through impairment of maturation of dendritic cells[17] . Dolganiuc et al[18] found that HCV core and NS3 proteins could activate monocytes. NS3 protein can activate innate immunity to recognize virus more efficiently through inducing inflammation [18]. Studies indicate that persistent HCV infection is closely related with hepatocellular carcinoma, replication and protein expression of HCV have been observed in HCC tissues[19-21], suggesting that interaction between HCV gene segments and expressed protein plays a key role in malignant transformation of hepatocytes [22]. Feng et al[23] showed that HCV NS3 N-terminal peptide may up-regulate the activation of MAPK, but not affect the expression of MAPK in normal liver cells. Additionally, Sakamuro et al[24] reported that NS3 gene-expressed protein in NIH3T3 cells and mouse fibroblasts can be transfected with NS3 gene segment , which induces tumors in nude mice. NS3 protein can inhibit actinomycin D-induced apoptosis[25] and inhibit phosphorylation mediated by cAMP-dependent protein kinase[26]. Activation of c-Jun NH2-terminal kinase (JNK) signaling pathway is essential for the stimulation of HCV non-structural protein 3 (NS3)-mediated cell growth[27]. It has been shown that cells consistently expressing low concentration of NS3 protein is prone to apoptosis induced by the Fas molecule[28].
In this experiment, we used pcDNA3.1(-) vector to construct the eukaryotic expression vector pcDNA3.1(-)/NS3 . DNA sequence analysis revealed that the inserted NS3 gene had correct reading frame and length(1893nt). Initial and terminal codons were successfully inserted into the ends of NS3 segment, which assured the correct expression of NS3 protein. We chose adult human HL-7702 hepatocytes as the host cells. The cells originate from embryonic liver, and were continuously cultivated in vitro. HL-7702 cells are slightly larger than normal human hepatocytes in vitro. The nuclei/cytoplasm ratio is high and cells are joined to each other tightly[29]. We transfected the constructed eukaryotic plasmid pcDNA3.1(-)/NS3 to HL-7702 cells, and detected the expressed protein by immunohistochemical staining. NS3 protein was expressed in the transfected cells, but not in blank cells and cells transfected with pcDNA3.1 (-). Western blot analysis showed that the expressed NS3 protein had correct molecular weight (70 000). The successful construction of eukaryotic plasmid pcDNA3.1 (-)/NS3 and correct expression of HCV NS3 protein in HL-7702 cells may contribute to the establishment of a system that can produce NS3 protein. These results may help further investigation of how NS3 protein affects immunity against virus.