Copyright
©The Author(s) 2018.
World J Hepatol. Feb 27, 2018; 10(2): 186-212
Published online Feb 27, 2018. doi: 10.4254/wjh.v10.i2.186
Published online Feb 27, 2018. doi: 10.4254/wjh.v10.i2.186
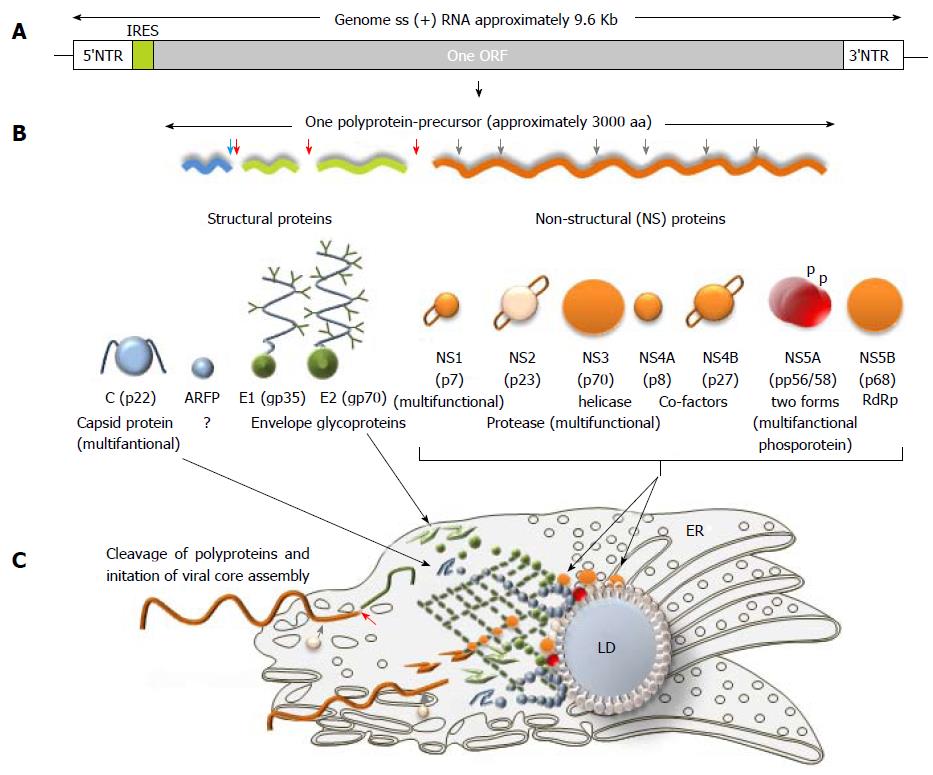
Figure 3 Hepatitis C virus genome, polyprotein precursor and the initial steps of core assembly in endoplasmic reticulum.
A: Being structurally identical, the genomes of seven hepatitis C virus (HCV) genotypes demonstrated approximately 30% of sequence diversity[140]. Two non-translated regions (5’- and 3’-NTR) are flanking a single open reading frame (ORF) shown in grey; B: Polyprotein precursor composed of about 3000 amino acids translated from a single ORF. Three structural proteins - a core protein (shown in blue) and two envelope proteins (shown in green) and seven non-structural (NS) proteins NS1, NS3, NS4A, NS4B, NS5A, and NS5B (shown in orange) that are involved in cleavage, assembly, transcription and some other functions are shown. Alternative reading frame protein (ARFP) that overlaps with the core protein sequence is given in blue. Serine protease NS2 is given in pale yellow. Transmembrane fragments in non-structural proteins are shown as staples; p- phosphoprotein; C: A model of initial steps of the virion core assembly in the endoplasmic reticulum (membranous web is given in dark green) on lipid droplet (LD). Polyprotein precursor is cleaved by cellular C-terminal signal peptidase (red arrow) and cellular signal peptidase (blue arrow) to release capsid protein. NS3-NS4A serine protease cleaves the remaining proteins, while NS2-NS3 (grey arrow) protease cleaves itself. Pre-assembled cores are transported to the LD, where the final steps of assembly take place[183].
- Citation: Morozov VA, Lagaye S. Hepatitis C virus: Morphogenesis, infection and therapy. World J Hepatol 2018; 10(2): 186-212
- URL: https://www.wjgnet.com/1948-5182/full/v10/i2/186.htm
- DOI: https://dx.doi.org/10.4254/wjh.v10.i2.186