Published online May 7, 2007. doi: 10.3748/wjg.v13.i17.2406
Revised: December 28, 2006
Accepted: February 14, 2007
Published online: May 7, 2007
Hepatitis C virus (HCV) encodes a single polyprotein, which is processed by cellular and viral proteases to generate 10 polypeptides. The HCV genome also contains an overlapping +1 reading frame that may lead to the synthesis of an additional protein. Until recently, studies of HCV have been hampered by the lack of a productive cell culture system. Since the identification of HCV genome approximately 17 years ago, structural, biochemical and biological information on HCV proteins has mainly been obtained with proteins produced by heterologous expression systems. In addition, some functional studies have also been confirmed with replicon systems or with retroviral particles pseudotyped with HCV envelope glycoproteins. The data that have accumulated on HCV proteins begin to provide a framework for understanding the molecular mechanisms involved in the major steps of HCV life cycle. Moreover, the knowledge accumulated on HCV proteins is also leading to the development of antiviral drugs among which some are showing promising results in early-phase clinical trials. This review summarizes the current knowledge on the functions and biochemical features of HCV proteins.
- Citation: Dubuisson J. Hepatitis C virus proteins. World J Gastroenterol 2007; 13(17): 2406-2415
- URL: https://www.wjgnet.com/1007-9327/full/v13/i17/2406.htm
- DOI: https://dx.doi.org/10.3748/wjg.v13.i17.2406
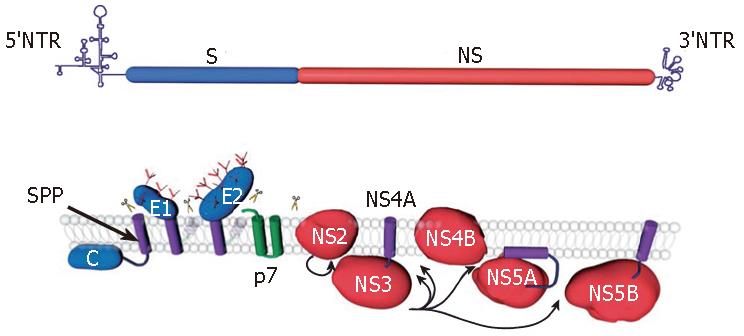
As for the other members of the Flaviviridae family the genome of Hepatitis C virus (HCV) encodes a single polyprotein. This 3010 amino acid polyprotein is processed by cellular and viral proteases to generate 10 polypeptides[1] (Figure 1). The nonstructural proteins are released from the polyprotein after cleavage by HCV proteases NS2-3 and NS3-4A, whereas the structural proteins are released by host endoplasmic reticulum (ER) signal peptidase(s)[2]. Further processing mediated by a signal peptide peptidase also occurs at the C-terminus of the capsid protein[3]. In addition to the large open reading frame encoding the polyprotein, the HCV genome contains an overlapping +1 reading frame that may lead to the synthesis of an additional protein[4]. Despite the difficulties in propagating the virus in cell culture, a large body of data has accumulated on HCV proteins since the identification of HCV genome 17 years ago. A detailed knowledge of the functions of HCV proteins is important for the development of new antiviral drugs. This review summarizes the current knowledge of the functions and biochemical features of HCV proteins. A brief summary of the functions of HCV proteins is presented in Table 1.
| Protein | Molecular Mass1 | Function |
| Core | 21 kDa | RNA binding; nucleocapsid |
| E1 | 31-35 kDa | Envelope glycoprotein; associate with E2 |
| E2 | 70 kDa | Envelope glycoprotein; receptor binding; associate with E1 |
| p7 | 7 kDa | Ion channel |
| NS2 | 21 kDa | Component of NS2-3 proteinase |
| NS3 | 69 kDa | N-terminal proteinase domain; |
| C-terminal NTPase/helicase domain | ||
| NS4A | 6 kDa | NS3-4A proteinase cofactor |
| NS4B | 27 kDa | Induces membrane alterations |
| NS5A | 56-58 kDa | Phosphoprotein |
| NS5B | 68 kDa | RNA-dependent RNA polymerase |
The core protein is an RNA-binding protein that is supposed to form the viral nucleocapsid. It is removed from the polyprotein by a host signal peptidase cleavage at the C-terminus, yielding the immature form of the protein[5], and the signal peptide present at the C-terminus of the core is processed further by a host signal peptide peptidase, yielding the mature form of the protein[3] (Figure 1). It has been shown that the mature form of core is a dimeric alpha-helical protein, which behaves as a membrane protein[6]. This protein can be separated into two domains: an N-terminal two-thirds hydrophilic domain (D1) and a C-terminal one-third hydrophobic domain (D2)[7]. The D1 domain includes numerous positively charged amino acids and has similar characteristics to the capsid proteins of related pestiviruses and flaviviruses[6,7]. The D2 domain is required for proper folding of domain D1 and is critical for the membrane characteristics of the core[6,8]. It is worth noting that this domain is absent in the pestiviruses and flaviviruses but is found in GB virus B[6,9].
Little is known about the mechanisms of HCV nucleo-capsid assembly. In vitro nucleocapsid reconstitution ex-periments with core segments have thus far yielded irregular particles larger than those isolated from infected subjects[10]. Full-length core protein has also been shown to assemble into nucleocapsid-like particles upon de novo synthesis in cell-free systems made of rabbit reticulocyte lysate or wheat germ extracts[11]. It has also been suggested that the attachment of a core protein to a phospholipid layer is required as a template for proper assembly of the nucleocapsid[6]. Although, little is known on the assembly of the nucleocapsid, developing small molecules that block the signal peptide peptidase cleavage might be a way of inhibiting HCV assembly.
When expressed in the context of heterologous expression systems or HCV replicons, core is found both attached to the ER and at the surface of lipid droplets[7,12]. In some conditions, a minor proportion of the core protein has also been found to be located in the nucleus[13]. More recently, the core protein has also been found to colocalize with mitochondrial markers in Huh-7 cells containing a full-length HCV replicon[14]. However, in the context of an infectious virus, the core protein was only found in association with lipid droplets[15]. It has been reported that the traffic between rough ER membranes, the site of capsid protein synthesis, and lipid droplets is regulated by signal peptide peptidase cleavage in the C-terminal region of the core protein[3]. It is therefore likely that in the context of HCV-infected cells, transport of the C protein to the site of lipid droplet assembly is rapid due to rapid cleavage by the signal peptide peptidase.
The core protein has been reported to interact with a variety of cellular proteins and to influence numerous host cell functions[7,16,17]. It has indeed been proposed to be involved in cell signaling, apoptosis, carcinogenesis and lipid metabolism. However, in most cases, it is unclear if these interactions occur in the course of a normal infection or are artifacts of ectopic expression or protein over-expression. Further studies with the recently developed cell culture system for HCV[18-20] should help clarify whether all the functions identified for HCV core protein can be observed in the context of infected cells.
HCV glycoproteins, E1 and E2, are released from the polyprotein by a host signal peptidase cleavage[12] (Figure 1). They are type-I transmembrane proteins with a large N-terminal ectodomain and a C-terminal transmembrane domain, and they assemble as noncovalent heterodimers[21]. The ectodomains of HCV envelope glycoproteins E1 and E2 are highly modified by N-linked glycans. Indeed, E1 and E2 possess up to 6 and 11 potential glycosylation sites, respectively, and most of them are well conserved[22,23]. It is worth noting that some glycans have been shown to play a role in HCV glycoprotein folding or in virus entry[24]. Because they are essential for virus entry, HCV envelope glycoproteins are a good target for the development of antiviral molecules that block HCV entry[25].
Hypervariable regions (HVR) have been identified in the E2 envelope glycoprotein sequence[26]. The first 27 amino acids of the E2 ectodomain form HVR1. The apparent variability of this region seems to be driven by antibody selection of immune-escape variants. An HCV clone lacking HVR1 was found to be infectious but strongly attenuated in chimpanzees[27], supporting a functional role of this domain, likely in virus entry[28,29]. Despite the sequence variability of HVR1, the physico-chemical properties of the residues at each position and the conformation of HVR1 are highly conserved among the various genotypes[30]. In addition, HVR1 is a global-ly basic region and basic residues of HVR1 have been shown to play a role in modulating virus entry[29]. Another hypervariable region, HVR2, has also been described in E2[26], and this region has been proposed to modulate E2 receptor binding[31].
Although HCV glycoproteins can be detected at the plasma membrane when they are over-expressed[32-34], the E1E2 heterodimer is mainly retained in the ER[15,35]. The determinants for ER retention of HCV envelope glyco-proteins have been mapped in the transmembrane domains of E1 and E2[36,37]. In addition to a membrane-proximal heptad repeat sequence in E2[38], these domains have also been shown to be essential for heterodimerization[39]. The transmembrane domains of HCV envelope glycoproteins are not canonical transmembrane domains[40], and dynamic changes have been shown to occur in these domains after cleavage by the signal peptidase[41]. Indeed, before cleavage by a host signal peptidase, the transmembrane domains of E1 and E2 adopt a hairpin structure, and after cleavage, the signal-like sequence is reoriented toward the cytosol, leading to a single transmembrane passage.
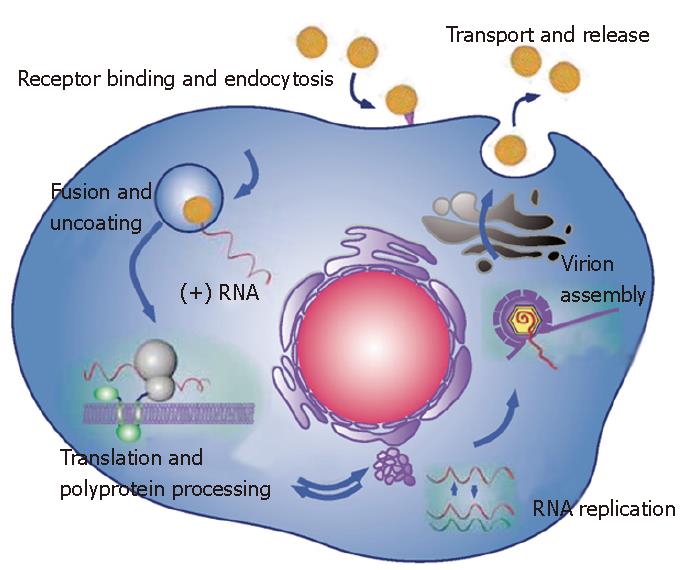
The two envelope glycoproteins, E1 and E2, play major roles at different steps of the HCV life cycle (Figure 2). They are essential for virus entry[42,43], and they partici-pate in the assembly of infectious particles[19]. The E1 E2 heterodimer is the viral component present at the sur-face of HCV particles and it is therefore the obvious candidate ligand for cellular receptors. As a first approach to identify potential HCV receptor(s), a soluble form of HCV glycoprotein E2 has been used. This led to the identification of a series of putative receptors for HCV: CD81 tetraspanin[44], scavenger receptor class B type I (SR-BI)[45], heparan sulfate[46] and the mannose binding lectins DC-SIGN and L-SIGN[47-49]. An approach using virus-like particles produced in insect cells has also led to the identification of the asialoglycoprotein receptor as another candidate receptor for HCV[50]. In addition, because of the physical association of HCV with low- or very-low-density lipoproteins (LDL or VLDL) in serum, the LDL receptor has also been proposed as another candidate receptor for HCV[51,52]. Among these molecules, only CD81 and SR-BI have been shown to play a role in HCV entry[43]. However, co-expression of CD81 and SR-BI in non-hepatic cell lines does not lead to virus entry, indicating that other molecule(s) expressed only in hepatic cells, are necessary for HCV entry.
Interactions between viral envelope glycoproteins and potential receptors can have other consequences than a direct effect on virus entry. For instance, L-SIGN and DC-SIGN are not expressed on hepatocytes, and HCV interactions with these molecules may contribute to es-tablishment or persistence of infection both by the cap-ture and delivery of virus to the liver and by modulating dendritic cell functions as recently suggested[53,54]. It has also been shown that intracellular interaction bet-ween HCV envelope glycoproteins and CD81 can lead to secretion of exosomes containing E1 and E2 glyco-proteins[55]. A soluble form of E2 is also able to bind CD81 at the surface of natural killer cells, and this in-teraction inhibits cytotoxicity and cytokine production by these cells[56,57]. Binding of a soluble form of E2 to CD81 can also provide a co-stimulatory signal for T cells[58,59], activate B lymphocytes[58] and up-regulate matrix metalloproteinase-2 in human hepatic stellate cells[60]. It remains however to be determined whether HCV glycoprotein expressed in the context of native particles will also have the same effects on cell functions.
Because they are exposed at the surface of the virion, the envelope proteins are targets of neutralizing antibodies. The recent development of retroviral particles pseudotyped by unmodified HCV E1 and E2 envelope glycoproteins (HCVpp)[32-34] has allowed to initiate studies on neutralizing antibodies. As determined with HCVpp, it seems that the majority of chronically infected patients have cross-reactive neutralizing antibodies[61,62]. In contrast, neutralizing antibodies have not been detected in several cases of acute resolving infection[61,62], and the detection of neutralizing antibodies in acutely infected individuals did not seem to be associated with viral clearance[61]. However, another study has shown in some patients a progressive emergence of a relatively strong neutralizing response in correlation with a decrease in viremia[63]. Further investigations on a large number of acutely infected patients will be necessary to determine the role of neutralizing antibodies in controlling HCV infections. Importantly, the majority of neutralizing anti-HCV monoclonal antibodies that have been described recognize E2[32,34,64-66]. In addition, some of the epitopes recognized by these antibodies have been mapped in the CD81 binding region of E2 and in the C-terminus of HVR1[34]. Studies with these neutralizing monoclonal anti-bodies will be essential to understand the mechanisms leading to HCV neutralization.
The p7 polypeptide is located within the HCV polyprotein at the junction between the structural and nonstructural proteins[67,68]. It is released from the polyprotein by a host signal peptidase cleavage[12] (Figure 1). The p7 polypeptide is a small polytopic membrane protein composed of two transmembrane domains with both its N- and C-termini oriented toward the lumen of the ER[69]. The C-terminus of p7 contains a sequence for reinitiation of translocation, and when fused to a reporter protein, this sequence func-tions as a signal peptide[69,70]. The double membrane span-ning topology of p7 with few residues accessible at one or the other side of the membrane suggests that p7 like-ly exerts its function(s) on membrane structures. When expressed by heterologous expression systems, p7 can be found in association with ER and/or mitochondrial mem-branes[69,71,72]. In addition, a small proportion of p7 can also be detected at the plasma membrane[69]. However, further investigations in the context of an infectious virus will be necessary to confirm these subcellular localizations. The p7 polypeptide is not required for RNA replication, and it is uncertain whether it is a virion component. Interestingly, the p7 polypeptide has been shown to have an ion channel activity in artificial lipid membranes[72-75]. In addition, it has been shown to be essential for infectivity of HCV in chimpanzees[76]. These observations suggest that screening for small molecules that block the ion channel activity of p7 might be an approach to develop new anti-HCV molecules.
NS2 is an integral membrane protein that is not essential for the formation of the replication complex[77,78]. The function of NS2 in its mature form is unknown; however, before cleavage from the polyprotein, NS2 participates in a protease activity responsible for the cleavage at the NS2/NS3 junction[79] (Figure 1). The first 180 residues of NS3 are also required for this cleavage. In addition, the NS2-3 enzyme has been described as a cysteine proteinase[80]. The structure of NS2 reveals a dimeric cysteine protease with two composite active sites[81]. Surprisingly, for each active site, the catalytic histidine and glutamate residues are contributed by one monomer, and the nucleophilic cysteine by the other. The host-cell chaperone Hsp90 seems to be required to activate the NS2-3 proteinase[82]. Cleavage of the NS2 N-terminus from p7 is mediated by a signal peptidase within the ER[69,70]. When expressed alone, NS2 is found located in association with ER membranes[83]. NS2 contains several stretches of hydrophobic amino acids and is predicted to be a polytopic membrane protein[84,85]. The membrane topology of NS2 is unclear, but the presence of two internal signal-like sequences points to the existence of four transmembrane segments[85]. However, since the processing at the NS2/NS3 junction has to take place in the cytosolic space, the presence of the C-terminus of NS2 in the ER lumen suggests that a reorientation of this region would have to occur after cleavage between NS2 and NS3. Interestingly, crossover sites for natural or infectious artificial inter-genotypic HCV chimeras have been mapped in NS2[18,86,87]. These data suggest that in addition to its role in the processing at the NS2/NS3 cleavage site, NS2 is also involved in virus assembly and release. It remains however to be determined by which mechanism NS2 contributes to the latter process. Due to its involvement in NS2-3 protease activity, NS2 is an interesting target for the development of anti-HCV molecules.
NS2 has been shown to be a short-lived protein whose degradation by the proteasome is regulated in a phosphorylation-dependent manner through the protein kinase CK2[83]. In addition, it has been shown to interact with the liver-specific pro-apoptotic CIDE-B protein and to be an inhibitor of CIDE-B-induced apoptosis[88]. NS2 might also potentially affect cellular gene transcription[89]. However, all these properties need to be further investi-gated in the context of the newly developed cell culture system for HCV[18-20].
NS3 is a multifunctional protein with an N-terminal serine-type protease domain and a C-terminal RNA helicase/NTPase domain. The NS3 protease domain has a typical chymotrypsin-like fold and is composed of two beta-barrel domains[90,91]. The protease activity of NS3 is enhanced by the NS4A cofactor. Indeed, NS4A contributes one beta-strand to the N-terminal protease domain and thereby allows its complete folding[90]. In addition, it induces a conformational change that leads to a repositioning of the catalytic triad. NS3 by itself has no transmembrane domain, but it associates non-covalently with the central domain of NS4A, which is a membrane protein. When co-expressed with NS4A, NS3 is found in association with ER or ER-like membranes whereas it is diffusely distributed in the cytoplasm and nucleus when expressed alone[92]. Deletion analyses have revealed that the hydrophobic N-terminal domain of NS4A is required for ER targeting of NS3. Interestingly, NS4A also stabilizes the protease against proteolytic degradation. The NS3-4A protease has an unusually shallow substrate-binding pocket and therefore requires rather long interaction surfaces with the substrate (reviewed in[1,93]). This made the design of efficient inhibitors of this protease challenging[94]. The NS3-4A protease is responsible for the polyprotein cleavage in the region downstream of NS3 (Figure 1), and this activity is essential for the generation of components of the viral RNA replication complex[95] (Figure 2). It is therefore not surprising that this protease has been the first target for the development of new anti-HCV molecules[94].
In addition to its role in the processing of the poly-protein, the NS3-4A protease activity is also involved in blocking the ability of the host cell to mount an innate antiviral response[96]. The NS3-4A has indeed been shown to interfere with double-stranded RNA signaling pathways. It disrupts the cellular RNA helicase retinoic acid-inducible gene I (RIG- I ) pathway through proteolysis of a newly discovered essential adaptor protein of interferon regulatory factor-3 (IRF-3) activation[97]. Due to its recent simultaneous discovery by four different groups, this adaptor protein has received four different names: IPS-1, Cardif, VISA and MAVS[98]. NS3-4A cleavage of MAVS/IPS-1/VISA/Cardif results in its dissociation from the mitochondrial membrane and disruption of signaling to the antiviral immune response[99]. NS3-4A also cleaves the TRIF (also called TICAM-1) adaptor protein to ablate Toll-like receptor-3 (TLR-3) signaling of IRF-3 activation by extracellular double-stranded RNA[100]. However, this pathway has a minimal role in triggering the interferon antiviral response[101].
The C terminus of NS3 encodes a DexH/D-box RNA helicase[102]. Enzymes of this superfamily are capable of unwinding RNA-RNA duplexes in an ATP-dependent manner. The crystal structure of the HCV helicase shows a Y-shaped molecule composed of 3 nearly equally sized subdomains[103,104]. Although monomeric NS3 can bind RNA with high affinity, RNA unwinding requires an NS3 dimer[105]. Kinetic analyses indicate that this enzyme undergoes highly coordinated cycles of fast double-stranded RNA unwinding[105-107]. More recently, it has been reported that the cyclic movement of NS3 helicase is coordinated by ATP in discrete steps of 11 base pairs, and that actual unwinding occurs in rapid smaller sub-steps of 2 to 5 base pairs, also triggered by ATP binding, indicating that NS3 might move like an inchworm[108]. The NS3 helicase activity can be modulated by interactions between the serine protease and helicase domains. Indeed the kinetics of duplex RNA unwinding is slower for the isolated helicase domain as compared with the full-length NS3 protein[109]. In addition, the presence of NS4A enhances productive RNA binding of a full-length NS3-4A complex[107]. The function of the NS3 helicase in the HCV life cycle is not known. It may be involved in initiation of RNA replication by unwinding stable stem-loop structures at the termini of positive and/or negative strand of HCV RNA. It may also contribute to the process of the replicase complex by removing stable RNA secondary structures and/or by displacing bound proteins that might interfere with RNA synthesis. Finally, it may also be required for dissociation of the replicative form. Due to its enzymatic activity, the helicase domain of NS3 is another potential target for the development of anti-HCV molecules.
The NS3 protein has been reported to interact with several cellular proteins[17], and it has been proposed to be involved in carcinogenesis[110]. However, the relevance of these interactions needs to be confirmed in the context of the recently developed cell culture system for HCV[18-20].
The NS4B protein is a highly hydrophobic nonstructural protein, which is predicted to contain four transmembrane domains[111,112]. It has recently been shown that NS4B is palmitoylated in the C-terminal region of the protein[113]. The N- and C-termini of NS4B are localized in the cytosol; however, a fraction of the N-terminus can also be found in the ER lumen[112]. A putative amphipathic helix in the N-terminus of NS4B has been proposed to mediate membrane association[114]. The NS4B protein is detected in association with ER membranes[111,112,115]. In addition, NS4B also induces intracellular membrane alterations, suggesting that one of its functions is to induce the formation of membranous structures supporting RNA replication[116]. However, the structure of NS4B-induced membranes appears to be slightly distinct from the membranous web observed when all the HCV proteins are expressed, suggesting that other component(s) contribute to these membrane alterations. A nucleotide binding motif has been found in NS4B[117]. This structural motif binds and hydrolyzes GTP. Interestingly, mutation of this nucleotide binding motif affects HCV RNA replication[117]. The potential presence of NS4B domains on both sides of the ER membrane suggests that this protein plays a role in crosstalk between the ER lumen and the cytosol. Although a function can be attributed to this protein, it remains challenging to develop a high-throughput screening for small molecules targeting NS4B.
NS5A is a membrane-associated protein containing a unique amphipathic alpha-helix at its N-terminus, which serves as an in-plane membrane anchor[118,119] (Figure 1). Like most HCV proteins, NS5A is detected in association with ER or ER-derived membranes[118]. Besides its membrane anchor sequence, NS5A contains three distinct domains that are separated by low complexity sequences (LCs) I and II [120]. Recently, the x-ray crystal structure of domain I was solved[121]. It is composed of a basic N-terminal subdomain IA and a predominantly acidic C-terminal subdomain IB. In subdomain IA a zinc ion is coordinated by a unique motif of 4 fully conserved cysteine residues, which are absolutely essential for RNA replication[120,121]. In subdomain IB an unusual disulfide bond linking 2 cysteine residues near the C-terminal subdomain border was found. However, this disulfide bond does not seem to be essential for HCV RNA repli-cation. Domain I forms homodimers via contacts near the N-terminal end of the molecules. This dimerization results in the formation of a basic groove facing the cytosol at the surface of the membrane. This 'claw like' structure is believed to provide an RNA binding site that might be involved in regulated genome targeting within the replication complex[121]. In line with this observation, NS5A has been shown to bind to HCV RNA and more specifically to the 3'-ends of HCV plus and minus strand RNAs, with a preference for the polypyrimidine tract in the 3' non-translated region of positive strand RNA[122]. Therefore, the structure of domain I of NS5A provides a framework for the rational design of small antiviral molecules. The other two domains of NS5A are less characterized. Domain II has been proposed to be involved in inhibition of the interferon-induced double stranded RNA activated protein kinase PKR[123], and domain III is a less conserved region, which can tolerate insertions or partial deletions[124,125].
NS5A is a protein which is essential for genome re-plication[126,127]. Indeed, mutations that enhance RNA replication in cell culture map to the NS5A-coding sequence. In addition, NS5A has been shown to interact with NS5B, and this interaction is essential for maintenance of sub-genomic replicons in Huh-7 cells[128,129]. NS5A is expressed as a basally phosphorylated and a hyperphosphorylated forms[93]. The functional relevance of the different pho-sphorylated forms is unknown. However, mutations that reduce NS5A hyperphosphorylation can lead to a dra-matic enhancement of HCV genomic replication[124,130]. Furthermore, treatment of cells carrying non-adapted repli-cons with an inhibitor of the cellular kinase(s) responsible for NS5A hyperphosphorylation leads to an increase in HCV genomic replication[131]. In addition to its role in HCV genomic replication, NS5A has initially attracted considerable interest because of its potential role in modu-lating the interferon response[132]. NS5A has also been shown to interact with components of numerous cellular signaling pathways[17,133,134]. Among the potential cellular partners identified for NS5A, human vesicle-associated membrane protein-associated protein A (hVAP-A) is of particular interest because it is regulated by NS5A phosphorylation[130,135]. Indeed, NS5A hyperphosphorylation disrupts interaction with hVAP-A and negatively regulates viral RNA replication. VAP-A is a protein found on ER and Golgi membranes, which is involved in intracellular vesicle trafficking. It remains however to be determined why NS5A hijacks hVAP-A at some step of its life cycle. Another potentially important host cell factor interacting with NS5A is the geranylgeranylated protein FBL-2[136]. In line with this observation, it has been shown that inhibition of geranylgeranylation in cells abolishes HCV RNA replication[137].
NS5B is a membrane-associated protein containing a C-terminal transmembrane domain[138], which is essential for RNA replication in cell culture[139] (Figure 1). Like most HCV proteins, NS5B is detected in association with ER or ER-derived membranes[140]. NS5B is an RNA-dependent RNA polymerase, which is the catalytic component of the HCV RNA replication machinery. This enzyme synthesizes RNA using an RNA template. NS5B can initiate RNA synthesis de novo, at least in vitro, and it is assumed that de novo initiation is also operating in vivo[93]. The crystal structure of the NS5B catalytic domain shows a structural fold comparable with other polymerases with palm, finger, and thumb subdomains[141,142]. The palm domain contains the active site of the enzyme, whereas the fingers and the thumb modulate the interaction with the RNA chain. One structural peculiarity of the enzyme is the fully encircled active site, which is due to multiple interactions between the finger and thumb subdomains creating a tunnel in which a single-stranded RNA molecule is directly guided to the active site. NTPs enter the active site via another positively charged tunnel. Binding of the RNA template and initiation of RNA synthesis are supposed to be regu-lated by a highly flexible beta-hairpin loop located in the thumb domain and pointing toward the active site[126]. As observed for other viral polymerases, NS5B is an inter-esting and promising target for the development of new antiviral molecules targeting HCV[94].
The RNA-dependent RNA polymerase activity appears to be modulated by interaction with some other viral proteins (NS3 and NS5A)[93]. It has been shown that cyclophilin B, a peptidyl-prolyl cis-trans-isomerase, interacts with the C-terminal region of NS5B and appears to stimulate its RNA binding activity[143]. In addition, cyclosporin A, an inhibitor of cyclophilin B, inhibits HCV replication in cell culture[144]. However, how cyclophilin B activates replication remains to be determined. Furthermore, cyclophilin B does not seem to stimulate the RNA binding activity of NS5B in all genotypes[145]. NS5B has also been shown to interact with other cellular proteins[146-148].
In addition to the large open reading frame encoding the polyprotein, HCV genome contains an overlapping +1 reading frame that overlaps the sequence of the core protein[4]. This alternative reading frame (ARF) lacks an in-frame AUG start codon, suggesting that its expression involves unusual translation-level events. In vitro studies indicate that ribosomal frameshifting may the process leading to translation of the ARF. Frameshifting yields chimeric proteins that have segments encoded in the core sequence covalently attached to amino acids encoded in the ARF. Based on experiments with reporter gene constructs, the frameshift efficiency is in the range of 1% to 2%. The development of an immune response against the ARF protein in HCV infected patients indicates that this protein is expressed during natural HCV infections and stimulates specific immune responses[149]. The role of ARF protein in the HCV life cycle and/or pathogenesis is not yet known. However, the ARF protein is not required for HCV RNA replication. One cannot exclude that the ARF protein may be responsible for some of the effects attributed to the core protein. Indeed, most studies seeking to define the function of the core protein have used sequences likely to contain a combination of the core protein and ARF protein. Due to the lack of knowledge of its function, the ARF protein is not currently considered as a target for the development of new antiviral molecules.
I thank Sophana Ung for preparing the illustrations. Research projects in the laboratory are supported by the "Agence Nationale de Recherche sur le Sida et les hépatites virales". J.D. is an international scholar of the Howard Hughes Medical Institute.
S- Editor Liu Y L- Editor Alpini GD E- Editor Liu Y
| 1. | Penin F, Dubuisson J, Rey FA, Moradpour D, Pawlotsky JM. Structural biology of hepatitis C virus. Hepatology. 2004;39:5-19. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 458] [Cited by in F6Publishing: 490] [Article Influence: 24.5] [Reference Citation Analysis (0)] |
| 2. | Reed KE, Rice CM. Overview of hepatitis C virus genome structure, polyprotein processing, and protein properties. Curr Top Microbiol Immunol. 2000;242:55-84. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 92] [Cited by in F6Publishing: 250] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 3. | McLauchlan J, Lemberg MK, Hope G, Martoglio B. Intramembrane proteolysis promotes trafficking of hepatitis C virus core protein to lipid droplets. EMBO J. 2002;21:3980-3988. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 379] [Cited by in F6Publishing: 365] [Article Influence: 16.6] [Reference Citation Analysis (0)] |
| 4. | Branch AD, Stump DD, Gutierrez JA, Eng F, Walewski JL. The hepatitis C virus alternate reading frame (ARF) and its family of novel products: the alternate reading frame protein/F-protein, the double-frameshift protein, and others. Semin Liver Dis. 2005;25:105-117. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 94] [Cited by in F6Publishing: 99] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 5. | Santolini E, Migliaccio G, La Monica N. Biosynthesis and biochemical properties of the hepatitis C virus core protein. J Virol. 1994;68:3631-3641. [PubMed] [Cited in This Article: ] |
| 6. | Boulant S, Vanbelle C, Ebel C, Penin F, Lavergne JP. Hepatitis C virus core protein is a dimeric alpha-helical protein exhibiting membrane protein features. J Virol. 2005;79:11353-11365. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 112] [Cited by in F6Publishing: 121] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 7. | McLauchlan J. Properties of the hepatitis C virus core protein: a structural protein that modulates cellular processes. J Viral Hepat. 2000;7:2-14. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 227] [Cited by in F6Publishing: 244] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 8. | Boulant S, Montserret R, Hope RG, Ratinier M, Targett-Adams P, Lavergne JP, Penin F, McLauchlan J. Structural determinants that target the hepatitis C virus core protein to lipid droplets. J Biol Chem. 2006;281:22236-22247. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 158] [Cited by in F6Publishing: 167] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 9. | Hope RG, Murphy DJ, McLauchlan J. The domains required to direct core proteins of hepatitis C virus and GB virus-B to lipid droplets share common features with plant oleosin proteins. J Biol Chem. 2002;277:4261-4270. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 130] [Cited by in F6Publishing: 141] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 10. | Kunkel M, Lorinczi M, Rijnbrand R, Lemon SM, Watowich SJ. Self-assembly of nucleocapsid-like particles from recombinant hepatitis C virus core protein. J Virol. 2001;75:2119-2129. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 130] [Cited by in F6Publishing: 135] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 11. | Klein KC, Polyak SJ, Lingappa JR. Unique features of hepatitis C virus capsid formation revealed by de novo cell-free assembly. J Virol. 2004;78:9257-9269. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 57] [Cited by in F6Publishing: 60] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 12. | Dubuisson J, Penin F, Moradpour D. Interaction of hepatitis C virus proteins with host cell membranes and lipids. Trends Cell Biol. 2002;12:517-523. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 94] [Cited by in F6Publishing: 100] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 13. | Yasui K, Wakita T, Tsukiyama-Kohara K, Funahashi SI, Ichikawa M, Kajita T, Moradpour D, Wands JR, Kohara M. The native form and maturation process of hepatitis C virus core protein. J Virol. 1998;72:6048-6055. [PubMed] [Cited in This Article: ] |
| 14. | Schwer B, Ren S, Pietschmann T, Kartenbeck J, Kaehlcke K, Bartenschlager R, Yen TS, Ott M. Targeting of hepatitis C virus core protein to mitochondria through a novel C-terminal localization motif. J Virol. 2004;78:7958-7968. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 123] [Cited by in F6Publishing: 131] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 15. | Rouillé Y, Helle F, Delgrange D, Roingeard P, Voisset C, Blanchard E, Belouzard S, McKeating J, Patel AH, Maertens G. Subcellular localization of hepatitis C virus structural proteins in a cell culture system that efficiently replicates the virus. J Virol. 2006;80:2832-2841. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 159] [Cited by in F6Publishing: 164] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 16. | Ray RB, Ray R. Hepatitis C virus core protein: intriguing properties and functional relevance. FEMS Microbiol Lett. 2001;202:149-156. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 112] [Cited by in F6Publishing: 116] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 17. | Tellinghuisen TL, Rice CM. Interaction between hepatitis C virus proteins and host cell factors. Curr Opin Microbiol. 2002;5:419-427. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 140] [Cited by in F6Publishing: 148] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 18. | Lindenbach BD, Evans MJ, Syder AJ, Wölk B, Tellinghuisen TL, Liu CC, Maruyama T, Hynes RO, Burton DR, McKeating JA. Complete replication of hepatitis C virus in cell culture. Science. 2005;309:623-626. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1843] [Cited by in F6Publishing: 1817] [Article Influence: 95.6] [Reference Citation Analysis (0)] |
| 19. | Wakita T, Pietschmann T, Kato T, Date T, Miyamoto M, Zhao Z, Murthy K, Habermann A, Kräusslich HG, Mizokami M. Production of infectious hepatitis C virus in tissue culture from a cloned viral genome. Nat Med. 2005;11:791-796. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2241] [Cited by in F6Publishing: 2239] [Article Influence: 117.8] [Reference Citation Analysis (0)] |
| 20. | Zhong J, Gastaminza P, Cheng G, Kapadia S, Kato T, Burton DR, Wieland SF, Uprichard SL, Wakita T, Chisari FV. Robust hepatitis C virus infection in vitro. Proc Natl Acad Sci USA. 2005;102:9294-9299. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1427] [Cited by in F6Publishing: 1441] [Article Influence: 75.8] [Reference Citation Analysis (0)] |
| 21. | Deleersnyder V, Pillez A, Wychowski C, Blight K, Xu J, Hahn YS, Rice CM, Dubuisson J. Formation of native hepatitis C virus glycoprotein complexes. J Virol. 1997;71:697-704. [PubMed] [Cited in This Article: ] |
| 22. | Goffard A, Dubuisson J. Glycosylation of hepatitis C virus envelope proteins. Biochimie. 2003;85:295-301. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 113] [Cited by in F6Publishing: 114] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 23. | Zhang M, Gaschen B, Blay W, Foley B, Haigwood N, Kuiken C, Korber B. Tracking global patterns of N-linked glycosylation site variation in highly variable viral glycoproteins: HIV, SIV, and HCV envelopes and influenza hemagglutinin. Glycobiology. 2004;14:1229-1246. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 360] [Cited by in F6Publishing: 376] [Article Influence: 18.8] [Reference Citation Analysis (0)] |
| 24. | Goffard A, Callens N, Bartosch B, Wychowski C, Cosset FL, Montpellier C, Dubuisson J. Role of N-linked glycans in the functions of hepatitis C virus envelope glycoproteins. J Virol. 2005;79:8400-8409. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 199] [Cited by in F6Publishing: 200] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 25. | Helle F, Wychowski C, Vu-Dac N, Gustafson KR, Voisset C, Dubuisson J. Cyanovirin-N inhibits hepatitis C virus entry by binding to envelope protein glycans. J Biol Chem. 2006;281:25177-25183. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 129] [Cited by in F6Publishing: 129] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 26. | Weiner AJ, Brauer MJ, Rosenblatt J, Richman KH, Tung J, Crawford K, Bonino F, Saracco G, Choo QL, Houghton M. Variable and hypervariable domains are found in the regions of HCV corresponding to the flavivirus envelope and NS1 proteins and the pestivirus envelope glycoproteins. Virology. 1991;180:842-848. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 433] [Cited by in F6Publishing: 442] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 27. | Forns X, Thimme R, Govindarajan S, Emerson SU, Purcell RH, Chisari FV, Bukh J. Hepatitis C virus lacking the hypervariable region 1 of the second envelope protein is infectious and causes acute resolving or persistent infection in chimpanzees. Proc Natl Acad Sci USA. 2000;97:13318-13323. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 120] [Cited by in F6Publishing: 127] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 28. | Bartosch B, Vitelli A, Granier C, Goujon C, Dubuisson J, Pascale S, Scarselli E, Cortese R, Nicosia A, Cosset FL. Cell entry of hepatitis C virus requires a set of co-receptors that include the CD81 tetraspanin and the SR-B1 scavenger receptor. J Biol Chem. 2003;278:41624-41630. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 452] [Cited by in F6Publishing: 471] [Article Influence: 22.4] [Reference Citation Analysis (0)] |
| 29. | Callens N, Ciczora Y, Bartosch B, Vu-Dac N, Cosset FL, Pawlotsky JM, Penin F, Dubuisson J. Basic residues in hypervariable region 1 of hepatitis C virus envelope glycoprotein e2 contribute to virus entry. J Virol. 2005;79:15331-15341. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 60] [Cited by in F6Publishing: 64] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 30. | Penin F, Combet C, Germanidis G, Frainais PO, Deléage G, Pawlotsky JM. Conservation of the conformation and positive charges of hepatitis C virus E2 envelope glycoprotein hypervariable region 1 points to a role in cell attachment. J Virol. 2001;75:5703-5710. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 138] [Cited by in F6Publishing: 136] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 31. | Roccasecca R, Ansuini H, Vitelli A, Meola A, Scarselli E, Acali S, Pezzanera M, Ercole BB, McKeating J, Yagnik A. Binding of the hepatitis C virus E2 glycoprotein to CD81 is strain specific and is modulated by a complex interplay between hypervariable regions 1 and 2. J Virol. 2003;77:1856-1867. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 129] [Cited by in F6Publishing: 134] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 32. | Bartosch B, Dubuisson J, Cosset FL. Infectious hepatitis C virus pseudo-particles containing functional E1-E2 envelope protein complexes. J Exp Med. 2003;197:633-642. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 885] [Cited by in F6Publishing: 868] [Article Influence: 41.3] [Reference Citation Analysis (0)] |
| 33. | Drummer HE, Maerz A, Poumbourios P. Cell surface expression of functional hepatitis C virus E1 and E2 glycoproteins. FEBS Lett. 2003;546:385-390. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 157] [Cited by in F6Publishing: 164] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 34. | Hsu M, Zhang J, Flint M, Logvinoff C, Cheng-Mayer C, Rice CM, McKeating JA. Hepatitis C virus glycoproteins mediate pH-dependent cell entry of pseudotyped retroviral particles. Proc Natl Acad Sci USA. 2003;100:7271-7276. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 655] [Cited by in F6Publishing: 636] [Article Influence: 30.3] [Reference Citation Analysis (0)] |
| 35. | Duvet S, Cocquerel L, Pillez A, Cacan R, Verbert A, Moradpour D, Wychowski C, Dubuisson J. Hepatitis C virus glycoprotein complex localization in the endoplasmic reticulum involves a determinant for retention and not retrieval. J Biol Chem. 1998;273:32088-32095. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 117] [Cited by in F6Publishing: 120] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 36. | Cocquerel L, Duvet S, Meunier JC, Pillez A, Cacan R, Wychowski C, Dubuisson J. The transmembrane domain of hepatitis C virus glycoprotein E1 is a signal for static retention in the endoplasmic reticulum. J Virol. 1999;73:2641-2649. [PubMed] [Cited in This Article: ] |
| 37. | Cocquerel L, Meunier JC, Pillez A, Wychowski C, Dubuisson J. A retention signal necessary and sufficient for endoplasmic reticulum localization maps to the transmembrane domain of hepatitis C virus glycoprotein E2. J Virol. 1998;72:2183-2191. [PubMed] [Cited in This Article: ] |
| 38. | Drummer HE, Poumbourios P. Hepatitis C virus glycoprotein E2 contains a membrane-proximal heptad repeat sequence that is essential for E1E2 glycoprotein heterodimerization and viral entry. J Biol Chem. 2004;279:30066-30072. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 63] [Cited by in F6Publishing: 64] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 39. | Op De Beeck A, Montserret R, Duvet S, Cocquerel L, Cacan R, Barberot B, Le Maire M, Penin F, Dubuisson J. The transmembrane domains of hepatitis C virus envelope glycoproteins E1 and E2 play a major role in heterodimerization. J Biol Chem. 2000;275:31428-31437. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 128] [Cited by in F6Publishing: 132] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 40. | Cocquerel L, Wychowski C, Minner F, Penin F, Dubuisson J. Charged residues in the transmembrane domains of hepatitis C virus glycoproteins play a major role in the processing, subcellular localization, and assembly of these envelope proteins. J Virol. 2000;74:3623-3633. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 133] [Cited by in F6Publishing: 139] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 41. | Cocquerel L, Op de Beeck A, Lambot M, Roussel J, Delgrange D, Pillez A, Wychowski C, Penin F, Dubuisson J. Topological changes in the transmembrane domains of hepatitis C virus envelope glycoproteins. EMBO J. 2002;21:2893-2902. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 97] [Cited by in F6Publishing: 100] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 42. | Bartosch B, Cosset FL. Cell entry of hepatitis C virus. Virology. 2006;348:1-12. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 112] [Cited by in F6Publishing: 117] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 43. | Cocquerel L, Voisset C, Dubuisson J. Hepatitis C virus entry: potential receptors and their biological functions. J Gen Virol. 2006;87:1075-1084. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 132] [Cited by in F6Publishing: 141] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 44. | Pileri P, Uematsu Y, Campagnoli S, Galli G, Falugi F, Petracca R, Weiner AJ, Houghton M, Rosa D, Grandi G. Binding of hepatitis C virus to CD81. Science. 1998;282:938-941. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1572] [Cited by in F6Publishing: 1521] [Article Influence: 58.5] [Reference Citation Analysis (0)] |
| 45. | Scarselli E, Ansuini H, Cerino R, Roccasecca RM, Acali S, Filocamo G, Traboni C, Nicosia A, Cortese R, Vitelli A. The human scavenger receptor class B type I is a novel candidate receptor for the hepatitis C virus. EMBO J. 2002;21:5017-5025. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 888] [Cited by in F6Publishing: 868] [Article Influence: 39.5] [Reference Citation Analysis (0)] |
| 46. | Barth H, Schafer C, Adah MI, Zhang F, Linhardt RJ, Toyoda H, Kinoshita-Toyoda A, Toida T, Van Kuppevelt TH, Depla E. Cellular binding of hepatitis C virus envelope glycoprotein E2 requires cell surface heparan sulfate. J Biol Chem. 2003;278:41003-41012. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 356] [Cited by in F6Publishing: 368] [Article Influence: 17.5] [Reference Citation Analysis (0)] |
| 47. | Lozach PY, Lortat-Jacob H, de Lacroix de Lavalette A, Staropoli I, Foung S, Amara A, Houles C, Fieschi F, Schwartz O, Virelizier JL. DC-SIGN and L-SIGN are high affinity binding receptors for hepatitis C virus glycoprotein E2. J Biol Chem. 2003;278:20358-20366. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 281] [Cited by in F6Publishing: 285] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 48. | Pöhlmann S, Zhang J, Baribaud F, Chen Z, Leslie GJ, Lin G, Granelli-Piperno A, Doms RW, Rice CM, McKeating JA. Hepatitis C virus glycoproteins interact with DC-SIGN and DC-SIGNR. J Virol. 2003;77:4070-4080. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 296] [Cited by in F6Publishing: 310] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 49. | Gardner JP, Durso RJ, Arrigale RR, Donovan GP, Maddon PJ, Dragic T, Olson WC. L-SIGN (CD 209L) is a liver-specific capture receptor for hepatitis C virus. Proc Natl Acad Sci USA. 2003;100:4498-4503. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 228] [Cited by in F6Publishing: 230] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 50. | Saunier B, Triyatni M, Ulianich L, Maruvada P, Yen P, Kohn LD. Role of the asialoglycoprotein receptor in binding and entry of hepatitis C virus structural proteins in cultured human hepatocytes. J Virol. 2003;77:546-559. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 99] [Cited by in F6Publishing: 101] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 51. | Agnello V, Abel G, Elfahal M, Knight GB, Zhang QX. Hepatitis C virus and other flaviviridae viruses enter cells via low density lipoprotein receptor. Proc Natl Acad Sci USA. 1999;96:12766-12771. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 702] [Cited by in F6Publishing: 688] [Article Influence: 27.5] [Reference Citation Analysis (0)] |
| 52. | Monazahian M, Böhme I, Bonk S, Koch A, Scholz C, Grethe S, Thomssen R. Low density lipoprotein receptor as a candidate receptor for hepatitis C virus. J Med Virol. 1999;57:223-229. [PubMed] [DOI] [Cited in This Article: ] [Cited by in F6Publishing: 5] [Reference Citation Analysis (0)] |
| 53. | Cormier EG, Durso RJ, Tsamis F, Boussemart L, Manix C, Olson WC, Gardner JP, Dragic T. L-SIGN (CD209L) and DC-SIGN (CD209) mediate transinfection of liver cells by hepatitis C virus. Proc Natl Acad Sci USA. 2004;101:14067-14072. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 161] [Cited by in F6Publishing: 165] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 54. | Lozach PY, Amara A, Bartosch B, Virelizier JL, Arenzana-Seisdedos F, Cosset FL, Altmeyer R. C-type lectins L-SIGN and DC-SIGN capture and transmit infectious hepatitis C virus pseudotype particles. J Biol Chem. 2004;279:32035-32045. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 146] [Cited by in F6Publishing: 148] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 55. | Masciopinto F, Giovani C, Campagnoli S, Galli-Stampino L, Colombatto P, Brunetto M, Yen TS, Houghton M, Pileri P, Abrignani S. Association of hepatitis C virus envelope proteins with exosomes. Eur J Immunol. 2004;34:2834-2842. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 142] [Cited by in F6Publishing: 147] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 56. | Crotta S, Stilla A, Wack A, D'Andrea A, Nuti S, D'Oro U, Mosca M, Filliponi F, Brunetto RM, Bonino F. Inhibition of natural killer cells through engagement of CD81 by the major hepatitis C virus envelope protein. J Exp Med. 2002;195:35-41. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 342] [Cited by in F6Publishing: 354] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 57. | Tseng CT, Klimpel GR. Binding of the hepatitis C virus envelope protein E2 to CD81 inhibits natural killer cell functions. J Exp Med. 2002;195:43-49. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 337] [Cited by in F6Publishing: 348] [Article Influence: 15.8] [Reference Citation Analysis (0)] |
| 58. | Wack A, Soldaini E, Tseng C, Nuti S, Klimpel G, Abrignani S. Binding of the hepatitis C virus envelope protein E2 to CD81 provides a co-stimulatory signal for human T cells. Eur J Immunol. 2001;31:166-175. [PubMed] [DOI] [Cited in This Article: ] [Cited by in F6Publishing: 3] [Reference Citation Analysis (0)] |
| 59. | Rosa D, Saletti G, De Gregorio E, Zorat F, Comar C, D'Oro U, Nuti S, Houghton M, Barnaba V, Pozzato G. Activation of naïve B lymphocytes via CD81, a pathogenetic mechanism for hepatitis C virus-associated B lymphocyte disorders. Proc Natl Acad Sci USA. 2005;102:18544-18549. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 211] [Cited by in F6Publishing: 207] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 60. | Mazzocca A, Sciammetta SC, Carloni V, Cosmi L, Annunziato F, Harada T, Abrignani S, Pinzani M. Binding of hepatitis C virus envelope protein E2 to CD81 up-regulates matrix metalloproteinase-2 in human hepatic stellate cells. J Biol Chem. 2005;280:11329-11339. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 102] [Cited by in F6Publishing: 90] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 61. | Logvinoff C, Major ME, Oldach D, Heyward S, Talal A, Balfe P, Feinstone SM, Alter H, Rice CM, McKeating JA. Neutralizing antibody response during acute and chronic hepatitis C virus infection. Proc Natl Acad Sci USA. 2004;101:10149-10154. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 333] [Cited by in F6Publishing: 324] [Article Influence: 16.2] [Reference Citation Analysis (0)] |
| 62. | Meunier JC, Engle RE, Faulk K, Zhao M, Bartosch B, Alter H, Emerson SU, Cosset FL, Purcell RH, Bukh J. Evidence for cross-genotype neutralization of hepatitis C virus pseudo-particles and enhancement of infectivity by apolipoprotein C1. Proc Natl Acad Sci USA. 2005;102:4560-4565. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 195] [Cited by in F6Publishing: 207] [Article Influence: 10.9] [Reference Citation Analysis (0)] |
| 63. | Lavillette D, Morice Y, Germanidis G, Donot P, Soulier A, Pagkalos E, Sakellariou G, Intrator L, Bartosch B, Pawlotsky JM. Human serum facilitates hepatitis C virus infection, and neutralizing responses inversely correlate with viral replication kinetics at the acute phase of hepatitis C virus infection. J Virol. 2005;79:6023-6034. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 209] [Cited by in F6Publishing: 209] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 64. | Keck ZY, Op De Beeck A, Hadlock KG, Xia J, Li TK, Dubuisson J, Foung SK. Hepatitis C virus E2 has three immunogenic domains containing conformational epitopes with distinct properties and biological functions. J Virol. 2004;78:9224-9232. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 117] [Cited by in F6Publishing: 119] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 65. | Owsianka A, Tarr AW, Juttla VS, Lavillette D, Bartosch B, Cosset FL, Ball JK, Patel AH. Monoclonal antibody AP33 defines a broadly neutralizing epitope on the hepatitis C virus E2 envelope glycoprotein. J Virol. 2005;79:11095-11104. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 224] [Cited by in F6Publishing: 234] [Article Influence: 12.3] [Reference Citation Analysis (0)] |
| 66. | Schofield DJ, Bartosch B, Shimizu YK, Allander T, Alter HJ, Emerson SU, Cosset FL, Purcell RH. Human monoclonal antibodies that react with the E2 glycoprotein of hepatitis C virus and possess neutralizing activity. Hepatology. 2005;42:1055-1062. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 49] [Cited by in F6Publishing: 56] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 67. | Lin C, Lindenbach BD, Prágai BM, McCourt DW, Rice CM. Processing in the hepatitis C virus E2-NS2 region: identification of p7 and two distinct E2-specific products with different C termini. J Virol. 1994;68:5063-5073. [PubMed] [Cited in This Article: ] |
| 68. | Mizushima H, Hijikata M, Asabe S, Hirota M, Kimura K, Shimotohno K. Two hepatitis C virus glycoprotein E2 products with different C termini. J Virol. 1994;68:6215-6222. [PubMed] [Cited in This Article: ] |
| 69. | Carrère-Kremer S, Montpellier-Pala C, Cocquerel L, Wychowski C, Penin F, Dubuisson J. Subcellular localization and topology of the p7 polypeptide of hepatitis C virus. J Virol. 2002;76:3720-3730. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 159] [Cited by in F6Publishing: 159] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 70. | Carrère-Kremer S, Montpellier C, Lorenzo L, Brulin B, Cocquerel L, Belouzard S, Penin F, Dubuisson J. Regulation of hepatitis C virus polyprotein processing by signal peptidase involves structural determinants at the p7 sequence junctions. J Biol Chem. 2004;279:41384-41392. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 50] [Cited by in F6Publishing: 54] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 71. | Griffin S, Clarke D, McCormick C, Rowlands D, Harris M. Signal peptide cleavage and internal targeting signals direct the hepatitis C virus p7 protein to distinct intracellular membranes. J Virol. 2005;79:15525-15536. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 57] [Cited by in F6Publishing: 61] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 72. | Griffin SD, Harvey R, Clarke DS, Barclay WS, Harris M, Rowlands DJ. A conserved basic loop in hepatitis C virus p7 protein is required for amantadine-sensitive ion channel activity in mammalian cells but is dispensable for localization to mitochondria. J Gen Virol. 2004;85:451-461. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 131] [Cited by in F6Publishing: 136] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 73. | Griffin SD, Beales LP, Clarke DS, Worsfold O, Evans SD, Jaeger J, Harris MP, Rowlands DJ. The p7 protein of hepatitis C virus forms an ion channel that is blocked by the antiviral drug, Amantadine. FEBS Lett. 2003;535:34-38. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 351] [Cited by in F6Publishing: 338] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 74. | Pavlović D, Neville DC, Argaud O, Blumberg B, Dwek RA, Fischer WB, Zitzmann N. The hepatitis C virus p7 protein forms an ion channel that is inhibited by long-alkyl-chain iminosugar derivatives. Proc Natl Acad Sci USA. 2003;100:6104-6108. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 286] [Cited by in F6Publishing: 274] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 75. | Premkumar A, Wilson L, Ewart GD, Gage PW. Cation-selective ion channels formed by p7 of hepatitis C virus are blocked by hexamethylene amiloride. FEBS Lett. 2004;557:99-103. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 153] [Cited by in F6Publishing: 159] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 76. | Sakai A, Claire MS, Faulk K, Govindarajan S, Emerson SU, Purcell RH, Bukh J. The p7 polypeptide of hepatitis C virus is critical for infectivity and contains functionally important genotype-specific sequences. Proc Natl Acad Sci USA. 2003;100:11646-11651. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 166] [Cited by in F6Publishing: 182] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 77. | Blight KJ, Kolykhalov AA, Rice CM. Efficient initiation of HCV RNA replication in cell culture. Science. 2000;290:1972-1974. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1154] [Cited by in F6Publishing: 1139] [Article Influence: 47.5] [Reference Citation Analysis (0)] |
| 78. | Lohmann V, Körner F, Koch J, Herian U, Theilmann L, Bartenschlager R. Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science. 1999;285:110-113. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2294] [Cited by in F6Publishing: 2224] [Article Influence: 89.0] [Reference Citation Analysis (0)] |
| 79. | Lindenbach BD, Rice CM. Flaviviridae: The Viruses and Their Replication. Fields Virology. 4th ed. Philadelphia, Pa: Lippincott Williams & Wilkins 2001; 991-1042. [Cited in This Article: ] |
| 80. | Pallaoro M, Lahm A, Biasiol G, Brunetti M, Nardella C, Orsatti L, Bonelli F, Orrù S, Narjes F, Steinkühler C. Characterization of the hepatitis C virus NS2/3 processing reaction by using a purified precursor protein. J Virol. 2001;75:9939-9946. [PubMed] [DOI] [Cited in This Article: ] [Cited by in F6Publishing: 56] [Reference Citation Analysis (0)] |
| 81. | Lorenz IC, Marcotrigiano J, Dentzer TG, Rice CM. Structure of the catalytic domain of the hepatitis C virus NS2-3 protease. Nature. 2006;442:831-835. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 148] [Cited by in F6Publishing: 154] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 82. | Waxman L, Whitney M, Pollok BA, Kuo LC, Darke PL. Host cell factor requirement for hepatitis C virus enzyme maturation. Proc Natl Acad Sci USA. 2001;98:13931-13935. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 74] [Cited by in F6Publishing: 73] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 83. | Franck N, Le Seyec J, Guguen-Guillouzo C, Erdtmann L. Hepatitis C virus NS2 protein is phosphorylated by the protein kinase CK2 and targeted for degradation to the proteasome. J Virol. 2005;79:2700-2708. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 60] [Cited by in F6Publishing: 67] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 84. | Santolini E, Pacini L, Fipaldini C, Migliaccio G, Monica N. The NS2 protein of hepatitis C virus is a transmembrane polypeptide. J Virol. 1995;69:7461-7471. [PubMed] [Cited in This Article: ] |
| 85. | Yamaga AK, Ou JH. Membrane topology of the hepatitis C virus NS2 protein. J Biol Chem. 2002;277:33228-33234. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 70] [Cited by in F6Publishing: 78] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 86. | Kalinina O, Norder H, Mukomolov S, Magnius LO. A natural intergenotypic recombinant of hepatitis C virus identified in St. Petersburg. J Virol. 2002;76:4034-4043. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 235] [Cited by in F6Publishing: 260] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 87. | Pietschmann T, Kaul A, Koutsoudakis G, Shavinskaya A, Kallis S, Steinmann E, Abid K, Negro F, Dreux M, Cosset FL. Construction and characterization of infectious intragenotypic and intergenotypic hepatitis C virus chimeras. Proc Natl Acad Sci USA. 2006;103:7408-7413. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 602] [Cited by in F6Publishing: 593] [Article Influence: 32.9] [Reference Citation Analysis (0)] |
| 88. | Erdtmann L, Franck N, Lerat H, Le Seyec J, Gilot D, Cannie I, Gripon P, Hibner U, Guguen-Guillouzo C. The hepatitis C virus NS2 protein is an inhibitor of CIDE-B-induced apoptosis. J Biol Chem. 2003;278:18256-18264. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 83] [Cited by in F6Publishing: 83] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 89. | Dumoulin FL, von dem Bussche A, Li J, Khamzina L, Wands JR, Sauerbruch T, Spengler U. Hepatitis C virus NS2 protein inhibits gene expression from different cellular and viral promoters in hepatic and nonhepatic cell lines. Virology. 2003;305:260-266. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 43] [Cited by in F6Publishing: 50] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 90. | Kim JL, Morgenstern KA, Lin C, Fox T, Dwyer MD, Landro JA, Chambers SP, Markland W, Lepre CA, O'Malley ET. Crystal structure of the hepatitis C virus NS3 protease domain complexed with a synthetic NS4A cofactor peptide. Cell. 1996;87:343-355. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 566] [Cited by in F6Publishing: 507] [Article Influence: 18.1] [Reference Citation Analysis (0)] |
| 91. | Love RA, Parge HE, Wickersham JA, Hostomsky Z, Habuka N, Moomaw EW, Adachi T, Hostomska Z. The crystal structure of hepatitis C virus NS3 proteinase reveals a trypsin-like fold and a structural zinc binding site. Cell. 1996;87:331-342. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 406] [Cited by in F6Publishing: 409] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 92. | Wölk B, Sansonno D, Kräusslich HG, Dammacco F, Rice CM, Blum HE, Moradpour D. Subcellular localization, stability, and trans-cleavage competence of the hepatitis C virus NS3-NS4A complex expressed in tetracycline-regulated cell lines. J Virol. 2000;74:2293-2304. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 212] [Cited by in F6Publishing: 225] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 93. | Bartenschlager R, Frese M, Pietschmann T. Novel insights into hepatitis C virus replication and persistence. Adv Virus Res. 2004;63:71-180. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 213] [Cited by in F6Publishing: 227] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 94. | De Francesco R, Migliaccio G. Challenges and successes in developing new therapies for hepatitis C. Nature. 2005;436:953-960. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 331] [Cited by in F6Publishing: 350] [Article Influence: 18.4] [Reference Citation Analysis (0)] |
| 95. | Lindenbach BD, Rice CM. Unravelling hepatitis C virus replication from genome to function. Nature. 2005;436:933-938. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 602] [Cited by in F6Publishing: 583] [Article Influence: 30.7] [Reference Citation Analysis (0)] |
| 96. | Foy E, Li K, Wang C, Sumpter R, Ikeda M, Lemon SM, Gale M. Regulation of interferon regulatory factor-3 by the hepatitis C virus serine protease. Science. 2003;300:1145-1148. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 624] [Cited by in F6Publishing: 649] [Article Influence: 30.9] [Reference Citation Analysis (0)] |
| 97. | Meylan E, Curran J, Hofmann K, Moradpour D, Binder M, Bartenschlager R, Tschopp J. Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus. Nature. 2005;437:1167-1172. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1933] [Cited by in F6Publishing: 1854] [Article Influence: 97.6] [Reference Citation Analysis (0)] |
| 98. | Sen GC, Sarkar SN. Hitching RIG to action. Nat Immunol. 2005;6:1074-1076. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 21] [Cited by in F6Publishing: 22] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 99. | Li XD, Sun L, Seth RB, Pineda G, Chen ZJ. Hepatitis C virus protease NS3/4A cleaves mitochondrial antiviral signaling protein off the mitochondria to evade innate immunity. Proc Natl Acad Sci USA. 2005;102:17717-17722. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 630] [Cited by in F6Publishing: 657] [Article Influence: 34.6] [Reference Citation Analysis (0)] |
| 100. | Li K, Foy E, Ferreon JC, Nakamura M, Ferreon AC, Ikeda M, Ray SC, Gale M, Lemon SM. Immune evasion by hepatitis C virus NS3/4A protease-mediated cleavage of the Toll-like receptor 3 adaptor protein TRIF. Proc Natl Acad Sci USA. 2005;102:2992-2997. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 847] [Cited by in F6Publishing: 869] [Article Influence: 45.7] [Reference Citation Analysis (0)] |
| 101. | Lin R, Lacoste J, Nakhaei P, Sun Q, Yang L, Paz S, Wilkinson P, Julkunen I, Vitour D, Meurs E. Dissociation of a MAVS/IPS-1/VISA/Cardif-IKKepsilon molecular complex from the mitochondrial outer membrane by hepatitis C virus NS3-4A proteolytic cleavage. J Virol. 2006;80:6072-6083. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 177] [Cited by in F6Publishing: 183] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 102. | Tai CL, Chi WK, Chen DS, Hwang LH. The helicase activity associated with hepatitis C virus nonstructural protein 3 (NS3). J Virol. 1996;70:8477-8484. [PubMed] [Cited in This Article: ] |
| 103. | Kim JL, Morgenstern KA, Griffith JP, Dwyer MD, Thomson JA, Murcko MA, Lin C, Caron PR. Hepatitis C virus NS3 RNA helicase domain with a bound oligonucleotide: the crystal structure provides insights into the mode of unwinding. Structure. 1998;6:89-100. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 480] [Cited by in F6Publishing: 519] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 104. | Yao N, Hesson T, Cable M, Hong Z, Kwong AD, Le HV, Weber PC. Structure of the hepatitis C virus RNA helicase domain. Nat Struct Biol. 1997;4:463-467. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 343] [Cited by in F6Publishing: 375] [Article Influence: 13.9] [Reference Citation Analysis (0)] |
| 105. | Serebrov V, Pyle AM. Periodic cycles of RNA unwinding and pausing by hepatitis C virus NS3 helicase. Nature. 2004;430:476-480. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 100] [Cited by in F6Publishing: 109] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 106. | Levin MK, Gurjar M, Patel SS. A Brownian motor mechanism of translocation and strand separation by hepatitis C virus helicase. Nat Struct Mol Biol. 2005;12:429-435. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 101] [Cited by in F6Publishing: 104] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 107. | Pang PS, Jankowsky E, Planet PJ, Pyle AM. The hepatitis C viral NS3 protein is a processive DNA helicase with cofactor enhanced RNA unwinding. EMBO J. 2002;21:1168-1176. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 176] [Cited by in F6Publishing: 180] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 108. | Dumont S, Cheng W, Serebrov V, Beran RK, Tinoco I, Pyle AM, Bustamante C. RNA translocation and unwinding mechanism of HCV NS3 helicase and its coordination by ATP. Nature. 2006;439:105-108. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 293] [Cited by in F6Publishing: 277] [Article Influence: 15.4] [Reference Citation Analysis (0)] |
| 109. | Frick DN, Rypma RS, Lam AM, Gu B. The nonstructural protein 3 protease/helicase requires an intact protease domain to unwind duplex RNA efficiently. J Biol Chem. 2004;279:1269-1280. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 88] [Cited by in F6Publishing: 88] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 110. | Deng L, Nagano-Fujii M, Tanaka M, Nomura-Takigawa Y, Ikeda M, Kato N, Sada K, Hotta H. NS3 protein of Hepatitis C virus associates with the tumour suppressor p53 and inhibits its function in an NS3 sequence-dependent manner. J Gen Virol. 2006;87:1703-1713. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 67] [Cited by in F6Publishing: 70] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 111. | Hügle T, Fehrmann F, Bieck E, Kohara M, Kräusslich HG, Rice CM, Blum HE, Moradpour D. The hepatitis C virus nonstructural protein 4B is an integral endoplasmic reticulum membrane protein. Virology. 2001;284:70-81. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 172] [Cited by in F6Publishing: 180] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 112. | Lundin M, Monné M, Widell A, Von Heijne G, Persson MA. Topology of the membrane-associated hepatitis C virus protein NS4B. J Virol. 2003;77:5428-5438. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 156] [Cited by in F6Publishing: 162] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 113. | Yu GY, Lee KJ, Gao L, Lai MM. Palmitoylation and polymerization of hepatitis C virus NS4B protein. J Virol. 2006;80:6013-6023. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 95] [Cited by in F6Publishing: 99] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 114. | Elazar M, Liu P, Rice CM, Glenn JS. An N-terminal amphipathic helix in hepatitis C virus (HCV) NS4B mediates membrane association, correct localization of replication complex proteins, and HCV RNA replication. J Virol. 2004;78:11393-11400. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 123] [Cited by in F6Publishing: 129] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 115. | Gretton SN, Taylor AI, McLauchlan J. Mobility of the hepatitis C virus NS4B protein on the endoplasmic reticulum membrane and membrane-associated foci. J Gen Virol. 2005;86:1415-1421. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 48] [Cited by in F6Publishing: 51] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 116. | Egger D, Wölk B, Gosert R, Bianchi L, Blum HE, Moradpour D, Bienz K. Expression of hepatitis C virus proteins induces distinct membrane alterations including a candidate viral replication complex. J Virol. 2002;76:5974-5984. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 639] [Cited by in F6Publishing: 617] [Article Influence: 28.0] [Reference Citation Analysis (0)] |
| 117. | Einav S, Elazar M, Danieli T, Glenn JS. A nucleotide binding motif in hepatitis C virus (HCV) NS4B mediates HCV RNA replication. J Virol. 2004;78:11288-11295. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 97] [Cited by in F6Publishing: 104] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 118. | Brass V, Bieck E, Montserret R, Wölk B, Hellings JA, Blum HE, Penin F, Moradpour D. An amino-terminal amphipathic alpha-helix mediates membrane association of the hepatitis C virus nonstructural protein 5A. J Biol Chem. 2002;277:8130-8139. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 281] [Cited by in F6Publishing: 295] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 119. | Penin F, Brass V, Appel N, Ramboarina S, Montserret R, Ficheux D, Blum HE, Bartenschlager R, Moradpour D. Structure and function of the membrane anchor domain of hepatitis C virus nonstructural protein 5A. J Biol Chem. 2004;279:40835-40843. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 221] [Cited by in F6Publishing: 233] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 120. | Tellinghuisen TL, Marcotrigiano J, Gorbalenya AE, Rice CM. The NS5A protein of hepatitis C virus is a zinc metalloprotein. J Biol Chem. 2004;279:48576-48587. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 256] [Cited by in F6Publishing: 269] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 121. | Tellinghuisen TL, Marcotrigiano J, Rice CM. Structure of the zinc-binding domain of an essential component of the hepatitis C virus replicase. Nature. 2005;435:374-379. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 381] [Cited by in F6Publishing: 373] [Article Influence: 19.6] [Reference Citation Analysis (0)] |
| 122. | Huang L, Hwang J, Sharma SD, Hargittai MR, Chen Y, Arnold JJ, Raney KD, Cameron CE. Hepatitis C virus nonstructural protein 5A (NS5A) is an RNA-binding protein. J Biol Chem. 2005;280:36417-36428. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 194] [Cited by in F6Publishing: 200] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 123. | Gale M, Blakely CM, Kwieciszewski B, Tan SL, Dossett M, Tang NM, Korth MJ, Polyak SJ, Gretch DR, Katze MG. Control of PKR protein kinase by hepatitis C virus nonstructural 5A protein: molecular mechanisms of kinase regulation. Mol Cell Biol. 1998;18:5208-5218. [PubMed] [Cited in This Article: ] |
| 124. | Appel N, Pietschmann T, Bartenschlager R. Mutational analysis of hepatitis C virus nonstructural protein 5A: potential role of differential phosphorylation in RNA replication and identification of a genetically flexible domain. J Virol. 2005;79:3187-3194. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 184] [Cited by in F6Publishing: 185] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 125. | Moradpour D, Evans MJ, Gosert R, Yuan Z, Blum HE, Goff SP, Lindenbach BD, Rice CM. Insertion of green fluorescent protein into nonstructural protein 5A allows direct visualization of functional hepatitis C virus replication complexes. J Virol. 2004;78:7400-7409. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 202] [Cited by in F6Publishing: 216] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 126. | Appel N, Schaller T, Penin F, Bartenschlager R. From structure to function: new insights into hepatitis C virus RNA replication. J Biol Chem. 2006;281:9833-9836. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 144] [Cited by in F6Publishing: 151] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 127. | Seeger C. Salient molecular features of hepatitis C virus revealed. Trends Microbiol. 2005;13:528-534. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 7] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 128. | Shimakami T, Hijikata M, Luo H, Ma YY, Kaneko S, Shimotohno K, Murakami S. Effect of interaction between hepatitis C virus NS5A and NS5B on hepatitis C virus RNA replication with the hepatitis C virus replicon. J Virol. 2004;78:2738-2748. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 72] [Cited by in F6Publishing: 76] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 129. | Shirota Y, Luo H, Qin W, Kaneko S, Yamashita T, Kobayashi K, Murakami S. Hepatitis C virus (HCV) NS5A binds RNA-dependent RNA polymerase (RdRP) NS5B and modulates RNA-dependent RNA polymerase activity. J Biol Chem. 2002;277:11149-11155. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 174] [Cited by in F6Publishing: 181] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 130. | Evans MJ, Rice CM, Goff SP. Phosphorylation of hepatitis C virus nonstructural protein 5A modulates its protein interactions and viral RNA replication. Proc Natl Acad Sci USA. 2004;101:13038-13043. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 248] [Cited by in F6Publishing: 249] [Article Influence: 12.5] [Reference Citation Analysis (0)] |
| 131. | Neddermann P, Quintavalle M, Di Pietro C, Clementi A, Cerretani M, Altamura S, Bartholomew L, De Francesco R. Reduction of hepatitis C virus NS5A hyperphosphorylation by selective inhibition of cellular kinases activates viral RNA replication in cell culture. J Virol. 2004;78:13306-13314. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 108] [Cited by in F6Publishing: 107] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 132. | Tan SL, Katze MG. How hepatitis C virus counteracts the interferon response: the jury is still out on NS5A. Virology. 2001;284:1-12. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 143] [Cited by in F6Publishing: 158] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 133. | Reyes GR. The nonstructural NS5A protein of hepatitis C virus: an expanding, multifunctional role in enhancing hepatitis C virus pathogenesis. J Biomed Sci. 2002;9:187-197. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 65] [Cited by in F6Publishing: 68] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 134. | Macdonald A, Harris M. Hepatitis C virus NS5A: tales of a promiscuous protein. J Gen Virol. 2004;85:2485-2502. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 280] [Cited by in F6Publishing: 280] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 135. | Tu H, Gao L, Shi ST, Taylor DR, Yang T, Mircheff AK, Wen Y, Gorbalenya AE, Hwang SB, Lai MM. Hepatitis C virus RNA polymerase and NS5A complex with a SNARE-like protein. Virology. 1999;263:30-41. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 184] [Cited by in F6Publishing: 193] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 136. | Wang C, Gale M, Keller BC, Huang H, Brown MS, Goldstein JL, Ye J. Identification of FBL2 as a geranylgeranylated cellular protein required for hepatitis C virus RNA replication. Mol Cell. 2005;18:425-434. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 217] [Cited by in F6Publishing: 226] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 137. | Ye J, Wang C, Sumpter R, Brown MS, Goldstein JL, Gale M. Disruption of hepatitis C virus RNA replication through inhibition of host protein geranylgeranylation. Proc Natl Acad Sci USA. 2003;100:15865-15870. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 283] [Cited by in F6Publishing: 301] [Article Influence: 14.3] [Reference Citation Analysis (0)] |
| 138. | Ivashkina N, Wölk B, Lohmann V, Bartenschlager R, Blum HE, Penin F, Moradpour D. The hepatitis C virus RNA-dependent RNA polymerase membrane insertion sequence is a transmembrane segment. J Virol. 2002;76:13088-13093. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 70] [Cited by in F6Publishing: 73] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 139. | Moradpour D, Brass V, Bieck E, Friebe P, Gosert R, Blum HE, Bartenschlager R, Penin F, Lohmann V. Membrane association of the RNA-dependent RNA polymerase is essential for hepatitis C virus RNA replication. J Virol. 2004;78:13278-13284. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 104] [Cited by in F6Publishing: 109] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 140. | Schmidt-Mende J, Bieck E, Hugle T, Penin F, Rice CM, Blum HE, Moradpour D. Determinants for membrane association of the hepatitis C virus RNA-dependent RNA polymerase. J Biol Chem. 2001;276:44052-44063. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 157] [Cited by in F6Publishing: 163] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 141. | Bressanelli S, Tomei L, Roussel A, Incitti I, Vitale RL, Mathieu M, De Francesco R, Rey FA. Crystal structure of the RNA-dependent RNA polymerase of hepatitis C virus. Proc Natl Acad Sci USA. 1999;96:13034-13039. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 471] [Cited by in F6Publishing: 488] [Article Influence: 19.5] [Reference Citation Analysis (0)] |
| 142. | Lesburg CA, Cable MB, Ferrari E, Hong Z, Mannarino AF, Weber PC. Crystal structure of the RNA-dependent RNA polymerase from hepatitis C virus reveals a fully encircled active site. Nat Struct Biol. 1999;6:937-943. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 597] [Cited by in F6Publishing: 621] [Article Influence: 24.8] [Reference Citation Analysis (0)] |
| 143. | Watashi K, Ishii N, Hijikata M, Inoue D, Murata T, Miyanari Y, Shimotohno K. Cyclophilin B is a functional regulator of hepatitis C virus RNA polymerase. Mol Cell. 2005;19:111-122. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 351] [Cited by in F6Publishing: 366] [Article Influence: 19.3] [Reference Citation Analysis (0)] |
| 144. | Watashi K, Hijikata M, Hosaka M, Yamaji M, Shimotohno K. Cyclosporin A suppresses replication of hepatitis C virus genome in cultured hepatocytes. Hepatology. 2003;38:1282-1288. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 425] [Cited by in F6Publishing: 402] [Article Influence: 19.1] [Reference Citation Analysis (0)] |
| 145. | Ishii N, Watashi K, Hishiki T, Goto K, Inoue D, Hijikata M, Wakita T, Kato N, Shimotohno K. Diverse effects of cyclosporine on hepatitis C virus strain replication. J Virol. 2006;80:4510-4520. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 122] [Cited by in F6Publishing: 131] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 146. | Choi SH, Park KJ, Ahn BY, Jung G, Lai MM, Hwang SB. Hepatitis C virus nonstructural 5B protein regulates tumor necrosis factor alpha signaling through effects on cellular IkappaB kinase. Mol Cell Biol. 2006;26:3048-3059. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 35] [Cited by in F6Publishing: 37] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 147. | Shimakami T, Honda M, Kusakawa T, Murata T, Shimotohno K, Kaneko S, Murakami S. Effect of hepatitis C virus (HCV) NS5B-nucleolin interaction on HCV replication with HCV subgenomic replicon. J Virol. 2006;80:3332-3340. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 32] [Cited by in F6Publishing: 33] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 148. | Munakata T, Nakamura M, Liang Y, Li K, Lemon SM. Down-regulation of the retinoblastoma tumor suppressor by the hepatitis C virus NS5B RNA-dependent RNA polymerase. Proc Natl Acad Sci USA. 2005;102:18159-18164. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 106] [Cited by in F6Publishing: 112] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 149. | Bain C, Parroche P, Lavergne JP, Duverger B, Vieux C, Dubois V, Komurian-Pradel F, Trépo C, Gebuhrer L, Paranhos-Baccala G. Memory T-cell-mediated immune responses specific to an alternative core protein in hepatitis C virus infection. J Virol. 2004;78:10460-10469. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 56] [Cited by in F6Publishing: 61] [Article Influence: 3.1] [Reference Citation Analysis (0)] |