Published online Jun 28, 2020. doi: 10.35711/aimi.v1.i1.6
Peer-review started: June 1, 2020
First decision: June 5, 2020
Revised: June 17, 2020
Accepted: June 20, 2020
Article in press: June 20, 2020
Published online: June 28, 2020
Processing time: 38 Days and 14.2 Hours
Breast cancer represents the most common malignancy in women, being one of the most frequent cause of cancer-related mortality. Ultrasound, mammography, and magnetic resonance imaging (MRI) play a pivotal role in the diagnosis of breast lesions, with different levels of accuracy. Particularly, dynamic contrast-enhanced MRI has shown high diagnostic value in detecting multifocal, multicentric, or contralateral breast cancers. Radiomics is emerging as a promising tool for quantitative tumor evaluation, allowing the extraction of additional quantitative data from radiological imaging acquired with different modalities. Radiomics analysis may provide novel information through the quantification of lesions heterogeneity, that may be relevant in clinical practice for the characterization of breast lesions, prediction of tumor response to systemic therapies and evaluation of prognosis in patients with breast cancers. Several published studies have explored the value of radiomics with good-to-excellent diagnostic and prognostic performances for the evaluation of breast lesions. Particularly, the integrations of radiomics data with other clinical and histopathological parameters have demonstrated to improve the prediction of tumor aggressiveness with high accuracy and provided precise models that will help to guide clinical decisions and patients management. The purpose of this article in to describe the current application of radiomics in breast dynamic contrast-enhanced MRI.
Core tip: Dynamic contrast-enhanced-magnetic resonance imaging (DCE-MRI) has been evaluated in most of radiomics studies on breast cancers. However, heterogeneity in study designs related to magnetic field, contrast media used, and software available to perform radiomics challenge the comparisons of available results. In this review we will focus on the following applications of radiomics in breast DCE-MRI: characterization of breast lesions, prediction of breast cancer histological types, correlation with receptor status, prediction of lymph node metastases, prediction of tumor response to neoadjuvant systemic therapy, prognosis and recurrence risks.
- Citation: Orlando A, Dimarco M, Cannella R, Bartolotta TV. Breast dynamic contrast-enhanced-magnetic resonance imaging and radiomics: State of art. Artif Intell Med Imaging 2020; 1(1): 6-18
- URL: https://www.wjgnet.com/2644-3260/full/v1/i1/6.htm
- DOI: https://dx.doi.org/10.35711/aimi.v1.i1.6
Breast cancer represents the most common malignancy in women[1]. It is estimated that 268600 US women were newly diagnosed with invasive breast cancer in 2019, and that 41760 US women died of breast cancer[1]. Because of its incidence and clinical impact, early and accurate tumor detection with imaging is of utmost importance. Ultrasound, mammography, and magnetic resonance imaging (MRI) play a pivotal role in the diagnosis of breast lesions, with different levels of accuracy. Particularly, MRI has shown a greater sensitivity than mammography (92% vs 75%, respectively)[2] and ultrasound (90% vs 39% and 49% of ultrasound alone or associated with mam-mography, respectively)[3] for the diagnosis of breast cancer. Thanks to the ability to provide both morphologic and hemodynamic features, dynamic contrast-enhanced MRI (DCE-MRI) provides high sensitivity (over 90%) in the detection of breast cancer, although specificity for lesion characterization is still suboptimal (72%)[2,3]. DCE-MRI has shown high diagnostic value in detecting multifocal, multicentric, or contralateral disease not diagnosed on physical examination, mammography or ultrasound, recognition of ductal carcinoma in situ (DCIS), evaluation of treatment response to neoadjuvant chemotherapy, detection of occult primary breast cancer in patients with metastatic axillary nodes (the so-called “CUP syndrome”), and detection of cancer in dense breast tissue[4].
Recently, an increasing interest for the clinical utility of quantitative imaging is developing. In this scenario, radiomics is emerging as a promising tool for quantitative tumor evaluation. Radiomics allow to extract quantitative data from medical images that be combined to provide models for clinical decision support[5].
The purpose of this article in to describe the current application of radiomics in breast dynamic contrast-enhanced MRI.
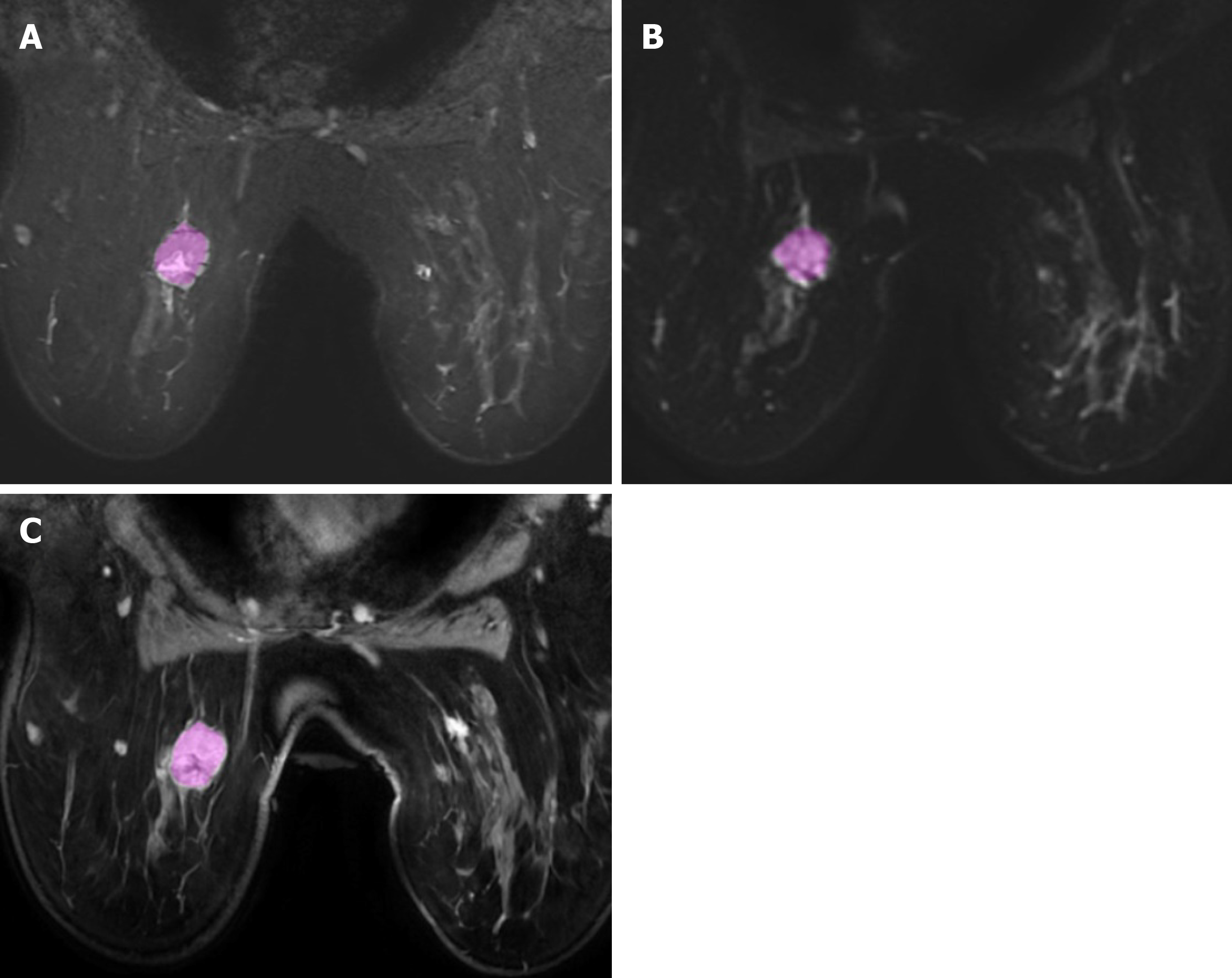
Radiomics is a complex process that articulates into distinct steps, including: Acquisition of images, tumor segmentation, feature extraction, exploratory analysis, and model building. The first step of radiomics is acquisition of high-quality images. Potentially, all the radiologic techniques may be used for radiomics analysis. In the field of breast imaging, all the techniques (mammography, ultrasound, and MRI) have shown promising results in radiomics studies. Particularly, breast MRI is commonly performed using T2-weighted images acquired to characterized diseased tissue, diffusion-weighted imaging (DWI), and apparent diffusion coefficient (ADC) that have an important clinical role in the evaluation of breast lesions, and post-contrast dynamic imaging that are mandatory for the differentiation of benign and malignant lesions. Next step is the segmentation of the lesion (Figure 1), with selection of a region of interest (ROI) and delineation of the borders of its volume. The ROI selection process is not yet standardized and it is linked to high levels of variability between different studies, as it can include the whole tumor or single slice segmentation[6].
Feature extraction may be performed with different radiomics software that are able to provide a large number of quantitative features. Quantitative radiomics features can be divided into morphological (basic features that describe the shape of the ROI and its geometric properties such as volume, diameter, sphericity), and statistical (calculated using statistical methods). These features can be further divided into first order (histogram-based) features that describe the distribution of voxel values without considering the spatial relationships (i.e. mean, median, skewness, kurtosis, and entropy); second order texture features that are obtained by calculating the relationships between neighboring voxels (i.e. grey level co-occurrence matrix, grey level run length matrix, grey level size zone matrix); and third order features that are obtained by statistical methods after applying filters or mathematical transforms to the images (i.e. wavelet transform, Laplacian transforms of Gaussian-filtered images)[7].
The next and last step in the workflow is building the statistical radiomics model with the purpose to predict an outcome or response variables. Different models can be evaluated to predict a specific outcome or a response using a variety of classifiers.
The emerging field of radiomics was applied to several breast imaging modalities[8,9]. Nevertheless, DCE-MRI was used in most studies but with heterogeneity in study designs related to magnetic field (1.5T or 3T), contrast media used, and software available to perform radiomics[10]. In this review we will focus the following applications of radiomics in breast DCE-MRI: Characterization of breast lesions, prediction of breast cancer histological types, correlation with receptor status, prediction with lymph node metastases, prediction of tumor response to neoadjuvant systemic therapy (NST), prognosis and recurrence risks.
Radiomics features extracted from multiple MRI sequences have shown to be helpful in establishing predictive models that could help differentiate between benign and malignant breast lesions. Several radiomics models were proposed with promising results, with most texture analysis performed on post-contrast T1-weighted images, alone or in association with other sequences (T2w and ADC maps).
Since the very first studies in literature, conducted on small populations analyzing different types of features extracted (dynamic, textural, spatio-temporal) from breast contrast-enhanced MRI, the dynamic subset revealed the best performance for the characterization of breast lesions for Fusco et al[11]. Testing a multi-layer perceptron neural network classifier, with an automatic ROI segmentation or ROI classification, they found an accuracy for dynamic features subset of about 80%, with the major discrimination power in differentiating benign from malignant lesions found for “basal signal”, “sum of intensities difference”, “relative enhancement slope” and “relative enhancement” features.
Nie et al[12] investigated the utility of breast lesions morphology and textural features for differentiating between benign and malignant lesions, with both manual and automated segmentation and performing diagnostic feature selection using artificial neural network. They found that among morphological features “Compactness” and “Normalized Radial Length Entropy” showed significant differences between the benign and the malignant groups, whereas among “Gray Level Co-occurrence Matrices” texture features, “Gray Level Entropy” and “Gray Level Sum Average” were significantly lower in benign compared to malignant lesions. Analyzing the diagnostic performance of individual and combined features the highest AUROC (0.86) was obtained combining the following 6 features: Compactness, NRL entropy, volume, gray level entropy, gray level sum average, and homogeneity. Entropy is an important feature associated with tumor aggressiveness. It represents one of the most reliable feature to distinguish malignant from benign lesions, with the irregularity of texture reflecting the tumor heterogeneity, and tumor aggressiveness[13-15]. Gibbs et al[16], testing texture analysis with the aim to characterize breast lesions, concluded that texture features of variance, sum entropy, and entropy were the most significant when discriminating between benign and malignant lesions.
Radiomics model of quantitative pharmacokinetic maps demonstrated a strong ability to discriminate between benign and malignant breast lesions, directly reflecting the physiological properties of tissues, such as vessel permeability, perfusion, and volume of the extravascular/extracellular space[13,17]. Nagarajan et al[18] studied texture features extracted from the lesion enhancement pattern on all five post-contrast images, thus using a dynamic texture quantification approach. In this study, the highest AUROC (0.82) was achieved with texture features responsible for capturing aspects of lesion heterogeneity. Gibbs et al[19] also assessed the efficacy of radiomics analysis with quantitative pharmacokinetic maps in small breast lesions (less than 1 cm). Their results showed that texture parameters calculated from initial enhancement, overall enhancement, and area under the enhancement curve maps offered similar discriminatory power in discriminating benign and malignant breast lesions, whereas texture features obtained from washout maps did not demonstrate any diagnostic value[19].
While many studies focused on discriminatory capacities of specific texture features extracted from combining quantitative pharmacokinetic parameters of DCE-MRI sequences, few studies used a multiparametric approach analyzing also feature extracted from other sequences, such as T2-weigthed and T1-weigthed imaging, diffusion kurtosis imaging, and ADC maps. The multimodal MRI-based radiomics model developed by Zhang et al[13] demonstrated higher diagnostic ability for differentiating benign and malignant breast lesions [Area under curve (AUC) = 0.921], increasing the discriminatory power of radiomics features extracted from DCE pharmacokinetic parameter maps alone (AUC = 0.836). In particular, analyzing textural features included in the radiomics models, malignant breast lesions had higher entropy and nonuniformity than benign lesions. The multiview IsoSVM (hybrid isomap and support vector machine) model applied by Parekh et al[20] to radiomics features extracted from multiparametric breast MR imaging at 3T, classified benign and malignant breast tumors with an AUROC of 0.91, sensitivity of 93%, and specificity of 85%. In this study, entropy features maps obtained demonstrated significantly higher entropy for malignant than benign lesions on post contrast DCE-MRI and ADC maps[20]. The same authors developed a multiparametric imaging radiomics framework for extraction of first and second order radiomics features from multiparametric radiological datasets which provided a 9%-28% increase in AUROC over single radiomics parameters. Similar results were reported by Bhooshan et al[21], who found the better performance applying a multiparametric feature vector, with T2-weighted MRI textural features added to DCE-MRI kinetic ones.
Radiomics features extracted from unenhanced MRI sequences were also evaluated for the prediction of malignancies. In the study of Bickelhaupt et al[22] an unenhanced, abbreviated DWI protocol (ueMRI), including T2-weighted, DWI, DWI with background suppression sequences, and corresponding ADC maps, was used to test three machine learning classifiers including univariate mean ADC model, unconstrained radiomics model, constrained radiomics model with mandatory inclusion of mean ADC. The last two radiomics classifiers were found to be able to distinguish benign from malignant lesions more accurately (AUROC of 0.842 and 0.851) than the mean ADC parameter alone (AUROC of 0.774)[22]. Nevertheless, the performance remained lower than that of the experienced breast radiologist using standard DCE-MRI protocol[22]. ADC radiomics features reflect the heterogeneity of diffusion in tumors, relative to the cell density and the microenvironment distribution inside the lesion. Hu et al[23] found that ADC radiomics score was more accurate than ADC values alone and they developed a prediction model based on ADC radiomics, pharmacokinetics and clinical features, which showed good diagnostic performance in differentiating benign and malignant lesions classified as BI-RADS 4. A radiomics model based on kurtosis diffusion-weighted imaging was evaluated by Bickelhaupt et al[24] who conducted a multicentric and prospective study on BI-RADS 4 and 5 lesions, by using MRI scanners from different vendors, showing reliable results, with a real benefit for BI-RADS 4a and 4b breast lesions.
Finally, more recent studies are using DCE-MRI focusing their attention on peritumoural tissues inclusion during segmentation. Zhou et al[25] found that the smallest bounding box, that included a small amount of peritumoral tissue adjacent to the tumor, had higher accuracy compared to tumor alone or larger input boxes.
Few studies employed radiomics models and texture analysis to distinguish between the heterogeneous histopathologic subtypes of breast cancer and entropy-based features from the co-occurrence matrix appear to be most crucial, with promising results. Invasive ductal (IDC) and lobular (ILC) carcinoma are the most common pathologic types. The different growth patterns may manifest with different heterogeneity of internal enhancement in DCE-MRI, and could be the basis to differentiate between these two histological types by means of textural analysis[14,26]. Holli et al[26] found that the co-occurrence matrix texture features group was statistically significant different between ductal and lobular invasive cancers on DCE-MR images. Similar conclusions were reported by Waugh et al[14] analyzing differences between IDC, ILC and in situ ductal carcinoma (DCIS). Chou et al[27] investigated the potential role of radiomics in classifying DCIS nuclear grade and found that only one heterogeneity metric, surface-to-volume ratio from the “shape and morphology” metrics group, was significantly different between “high nuclear grade” and “non-high nuclear grade” DCIS.
Expression of Ki-67, estrogen receptor (ER), progesterone receptor, human epidermal growth factor 2 receptor (HER2) are crucial factors to differentiate breast cancers into four main molecular subtypes (Luminal A, Luminal B, Her2 over-expressing, and triple negative, TN) with different outcomes and therapeutic strategies. According to the molecular subtypes different strategies, including surgery, adjuvant or neoadjuvant therapies, can be undertaken[28-31]. Current assessment of molecular subtypes is mostly based on immunohistochemistry (IHC)[32]. When IHC is tested in tissue specimens obtained by needle biopsy, could be not totally representative of the entire tumor or provide inconclusive results due to insufficient material. In this setting, according to prior studies, DCE-MRI may provide information suggesting the molecular subtype of breast cancer. In 2018, the American Joint Committee on Cancer updated the breast cancer staging guidelines to add other cancer characteristics to the TNM system to determine a cancer’s stage, including receptorial status[33]. When developing a treatment plan, a correct assessment of receptorial status is crucial. Several published studies revealed that rim enhancement, heterogeneous internal enhancement, and peritumoral edema are more frequently associated with TN than Luminal subtypes[34,35]. In the study of Blaschke et al[36] HER2-enriched tumors showed the percent volume with > 50% and > 100% early phase uptake higher than Luminal A/B lesions at kinetic assessment. TN tends to be more frequently round in shape[32,37], Her2 cancers with smooth margins than other subtypes[37]. Controversial results were reported for diffusion-weighted imaging, suggesting that high ADC values are associated with HER2 subtypes[38] or with Luminal A[39], and for spectroscopy, suggesting that high values of tCho are statistically correlated to the TN subtype for some authors[39,40], and with non-TN and Luminal B[41].
Several studies investigated the relationship between radiomics MRI features and breast cancer receptor status[42-44]. Wu et al[45] reported only few features significantly associated with Luminal A, Luminal B or TN in their study cohorts for distinguishing different molecular subtypes of breast cancers. Radiomics analysis conducted by Li et al[46] showed a statistically significant trend for the relationship between enhancement textures (entropy) and molecular subtypes in the task of distinguishing between ER+ versus ER−. Indeed, heterogeneous nature of contrast uptake within the breast tumor is related to molecular subtype. Similar observations were reported by Waugh et al[14], revealing that HER2-enriched and TN cancers showed a significant increase in entropy value. In the study of Chang et al[47] the quantitative region-based features extracted from breast DCE-MRI were used to interpret the intra-tumoral heterogeneity and correlated with ER, HER2, and TNBC, with better performance than morphological features (texture features and shape feature) and the pharmacokinetic model. Fan et al[48] investigated the use of features extracted from DCE-MRI for the prediction of the molecular subtypes of breast cancer and observed low kurtosis and skewness for the luminal A subtype, the highest enhancement values in the normal breasts for Her 2 subtypes and the lowest for luminal A and luminal B tumors. Furthermore, other studies suggested the value of the heterogeneity of the surrounding parenchyma, including background parenchymal enhancement features in differentiating TN breast cancers from others, as observed by Wang et al[49]. The evaluation of both peritumoral and intratumoral features allowed to identify HER2 subtype with better accuracy than intratumoral features alone in the study of Braman et al[50]. According to the results of Leithner et al[51] radiomics analysis from DWI with ADC mapping allows evaluation of breast cancer receptor status and molecular subtyping. For differentiating ER positive breast cancer molecular subtypes (Luminal A vs Luminal B) the two most discriminative texture parameters extracted from the dynamic T1-weighted sequences by Holli-Helenius et al[52] were sum entropy and sum variance, which also showed positive correlation with higher Ki-67 index.
High Ki-67 expression is a well-known prognostic factor, related to better neo-adjuvant therapy response but poorer prognosis. Assessment of Ki-67 based on immunohistochemistry on tissue specimens obtained by needle biopsy sample may not be representative of the whole tumor because of the relatively small tissue sample size and tumor heterogeneity. In the attempt to predict the expression of Ki-67 several studies have explored the potential of radiomics imaging features, with promising results. In their retrospective study, Ma et al[53] showed that texture features extracted on the first post-contrast images were associated with breast cancer Ki-67 expression. Similar results were obtained by Juan et al[54]. A correlation between Ki-67 expression and radiomics features were observed also performing features extraction from T2-weighted images[55] and ADC maps[56].
Involvement of axillary lymph nodes (LN) in patients with breast cancers represents a crucial prognostic factor, as it guides therapeutic management. Non-invasive methods to preoperatively evaluate LN metastasis are highly needed. Some promising studies suggested that radiomics models could be able to achieve this objective. In recent studies, specific lesions textural features extracted from anatomical and functional MRI images, improved the performance of radiomics models in predicting LN metastasis[57,58]. Liu et al[59] demonstrated that DCE-MRI radiomics features, particularly features extracted from peritumoral regions, associated with clinico-pathologic informations were able to predict LN metastasis in breast cancer patients. Indeed, the area surrounding tumors, is thought to carry informations such as peritumoral lymphatic vessel invasion, lymphocytic infiltration, and edema[59,60]. Other authors reported that the best results were obtained when the features extraction was performed in the strongest phases of tumor enhancement, probably because it shows more clearly the lesion boundaries and better reflects the tumor heterogeneity and invasiveness[61]. The radiomics nomogram developed by Han et al[62] demonstrated excellent performance to predict LN metastases, and good ability in distinguishing the number of metastatic LNs. Similar performances were reported by several other evidences[59,63-65]. Finally, only very few studies evaluated texture analysis in identified index lymph nodes in postcontrast T1-weighted images, concluding that morphologic features were more predictive than kinetic and texture features[66,67].
NST is often the first line treatment for those patients diagnosed with locally advanced breast cancer, with several potential advantages, including the reduction of tumor size to allow breast-conservative surgery instead of mastectomy, as well as a prognostic indicator[68]. The pathologic complete response (pCR) rate range from 0.3%–38.7%, depending on cancer subtype and breast cancer stage[69]. Early identification of patients who are not likely to achieve pCR is crucial as they could benefit from changes to their initial NST regimens. DCE-MRI is considered as the most reliable technique for evaluating the responses to NST. According to a meta-analysis based on 25 studies, breast MRI had high specificity (up to 90.7%), but low sensitivity (63.1%) in predicting pathologic complete remission after preoperative therapy in patients with breast cancer[70]. According to another recent meta-analysis, accuracy in detection of residual malignancy with breast MRI varies also in consideration of the treatment type, with AUC values ranging from 0.83 to 0.89, and on the basis of response definition, for instance volume reduction, absence of enhancement or enhancement equal or less than breast parenchyma[71,72]. The wide heterogeneity of studies, with controversial results, suggests to standardize definitions and primary endpoints to produce clinically significant results[73].
The identification of pCR is still a challenge and according with several studies, radiomics can be helpful in a non-invasive prediction of response to NST[74-78]. In most studies, GLCM features were the most predictive of therapy response, particularly entropy[79-81]. Noteworthy, in the study of Parikh et al[82], responders to NST showed increase in lesion homogeneity after one round of therapy. Cao et al[83] demonstrated that texture analysis may help to improve the performance of post-NST MRI in identifying pCR in mass-like breast cancer, showing that entropy was an independent risk factor. Intratumoral spatial heterogeneity at perfusion MRI appeared to be an independent prognostic factor of recurrence-free-survival in patients with locally advanced breast cancers treated with NST[84]. Significate differences between pCR and non-pCR patients were found for texture parameters also by Fusco et al[85]. Peritumoral region includes prognostic informations, such as angiogenic and lymphangiogenic activity, peritumoral invasion of lymphatics and blood vessels and peritumoral lymphocytic infiltration[86]. In their retrospective study, Braman et al[87] demonstrated that with combined intratumoral and peritumoral radiomics approach, analyzing textural features extracted from T1-weighted contrast-enhanced MRI scans, it is possible to successfully predict pCR to NST from pretreatment breast DCE-MRI, both with and without a priori knowledge of receptor status. Later, the same authors, confirmed that an intratumoral and peritumoral imaging signature was capable to predict the response to preroperative targeted therapy in another retrospective study conducted on HER2-positive breast cancers, highlighting again the relationship between immune-response and the peritumoral environment[50]. Zhou et al[88] investigated the role of wavelet-transformed textures, which can provide compre-hensive spatial, and frequency distributions for characterizing intratumoral and peritumoral regions in terms of low and high frequency signals. In their study wavelet-transformed textures outperformed volumetric and peripheral textures in the radiomics MRI prediction of pCR to NST for patients with locally advanced breast cancers.
DWI is considerably sensitive to NST-induced intratumoral changes, resulting in an additional value when associated to contrast-enhanced MRI in radiomics models. Radiomics signatures combining multi-parametric MRI achieved a good performance for predicting complete response in BC, in both Luminal and TN cancers, in the study conducted by Liu et al[89]. With a radiomics signature, combining radiomics features from DCE-MRI and ADC maps, Chen et al[90] obtained similar results, with a higher performance than the models with DCE-MRI or ADC maps alone, in predicting PCR.
Sentinel lymph node biopsy has replaced axillary lymph node dissection in patients who convert to node-negative status after NST. Several studies assessed whether breast MRI can be used to assess lymph node residual metastasis after NST allowing breast cancer patients to avoid unnecessary axillary surgery. In the study of Hyun et al[91], DCE-MRI was able to rule out the presence of advanced nodal disease with a NPV of 94% in NAC patients. Nevertheless, in the work of Mattingly et al[92], post-treatment MRI and surgical pathologic findings revealed a slight strength of agreement and DCE-MRI revealed specificity and sensitivity of 63% and 55%, respectively. Ha et al[93] found different results, with sensitivity and specificity of 57% and 72%, with positive estrogen receptor status significantly associated with misdiagnosis by MRI. These latter evidences, revealing that post-treatment MRI findings were not exactly predictive of residual axillary disease, suggest to use DCE-MRI results with caution when planning treatment and to avoid omitting sentinel lymph node biopsy or axillary lymph node dissection for staging in women determined to be node-positive pre-treatment. In this setting, convolutional neural networks (CNN), were employed to predict the likelihood of axillary LN metastasis and NAC treatment response, using MRI datasets prior to initiation of NAC in few studies with controversal results[79,94-96]. Ha et al[96] reported an accuracy of 83% with AUC of 0.93 for CNN in predicting axillary response. Nevertheless, in the study of Golden et al[79] the GLCM texture features extracted from pre- chemotherapy MRI was able to predict pCR and residual lymph node metastasis with an AUC of 0.68.
Radiomics models demonstrated promising results in predicting cancer prognosis of patients with tumors of various organs, reporting that several texture features, such as uniformity and entropy, can be used in risk stratification[15,97,98]. By using the genomic-based scores for the multigene assays MammaPrint, Oncotype DX, and PAM50 as the reference standards, Li et al[99] demonstrated that breast MRI radiomics show a promising role for image-based phenotyping in assessing the risk of recurrence. Noteworthy, enhancement texture features were consistently associated with recurrence score, highlighting how microvascular density and/or central necrosis, responsible of tumor heterogeneity, play an important biological role in recurrence. Other authors confirmed these results, finding that tumors with higher entropy on T2-weighted images and lower entropy on T1w subtraction images were associated with poorer recurrence-free survival[15]. A CNN developed by Ha et al[100] was able to predict with an accuracy of up to 84%, the Oncotype Dx Recurrence Score (ODRS), an expensive but validated recurrence score, recommended by American Society of Clinical Oncology guidelines to decide on adjuvant systemic chemotherapy in ER+/HER-/node negative lesions[101]. Nevertheless, this result was not confirmed by Saha et al[102], who tested two machine learning-based models, finding only a moderate association between imaging and ODRS. The study of Park et al[103] was the first performed, using ROIs drawn on entire tumors, to demonstrate that a radiomics signature can estimate survival in patients with BC. They generated a multivariate feature vector based on morphologic, histogram texture, and GLCM texture features to stratify patients at risk for recurrence. They also showed that a combined radiomics-clinical-pathological nomogram achieved superior prognostic performance than either the Rad-score-only or the clinico-pathological nomograms. Nevertheless, controversial results were recently reported applying radiomics models to predict prognosis for TN (triple-negative) breast cancers[104,105]. While in the study conducted by Kim et al[105] the radiomics score was significantly associated with worse disease free survival, but comparable in performance with the clinico-pathologic model, in both the training and validation sets, the work performed by Koh et al[104] showed that their Radiomics model was able to predict systemic recurrence better than the Clinical model only in the training set.
Radiomics techniques require further studies, as they have not yet achieved wide-spread, demonstrated and accepted, clinical relevance and applicability. The main challenge is the standardization of MRI acquisition protocol, method of segmentation, feature extraction and selection, or classification. Another hurdle is the current lack of evidences regarding reproducibility of feature extraction systems and radiomics models. Furthermore, most studies are retrospectively designed, with relatively small sample size and wide methodological differences. Larger and prospective studies, with standardized radiomics methods are needed to prove and improve potential clinical applications of radiomics in BC. Further studies are necessary to prove and understand the relationships between image-derived texture features and histopathologic or even genomic expression data. The main future directions include the correlation between proteomic and genomic tumor analyses with radiomics features, through the field of radiogenomics. These last investigations could have a potential role in explaining tumor biology, contributing in the main future objective of personalized diagnosis and treatment of breast cancer patients.
Manuscript source: Invited manuscript
Specialty type: Radiology, nuclear medicine and medical imaging
Country/Territory of origin: Italy
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Li JF S-Editor: Wang JL L-Editor: A E-Editor: Xing YX
| 1. | Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69:7-34. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13300] [Cited by in RCA: 15456] [Article Influence: 2576.0] [Reference Citation Analysis (2)] |
| 2. | Zhang Y, Ren H. Meta-analysis of diagnostic accuracy of magnetic resonance imaging and mammography for breast cancer. J Cancer Res Ther. 2017;13:862-868. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 27] [Article Influence: 3.9] [Reference Citation Analysis (2)] |
| 3. | Kuhl CK, Schrading S, Leutner CC, Morakkabati-Spitz N, Wardelmann E, Fimmers R, Kuhn W, Schild HH. Mammography, breast ultrasound, and magnetic resonance imaging for surveillance of women at high familial risk for breast cancer. J Clin Oncol. 2005;23:8469-8476. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 823] [Cited by in RCA: 752] [Article Influence: 37.6] [Reference Citation Analysis (0)] |
| 4. | Menezes GL, Knuttel FM, Stehouwer BL, Pijnappel RM, van den Bosch MA. Magnetic resonance imaging in breast cancer: A literature review and future perspectives. World J Clin Oncol. 2014;5:61-70. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 119] [Cited by in RCA: 105] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 5. | Gillies RJ, Kinahan PE, Hricak H. Radiomics: Images Are More than Pictures, They Are Data. Radiology. 2016;278:563-577. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4541] [Cited by in RCA: 5506] [Article Influence: 611.8] [Reference Citation Analysis (3)] |
| 6. | Tagliafico AS, Piana M, Schenone D, Lai R, Massone AM, Houssami N. Overview of radiomics in breast cancer diagnosis and prognostication. Breast. 2020;49:74-80. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 214] [Cited by in RCA: 178] [Article Influence: 35.6] [Reference Citation Analysis (0)] |
| 7. | Rizzo S, Botta F, Raimondi S, Origgi D, Fanciullo C, Morganti AG, Bellomi M. Radiomics: the facts and the challenges of image analysis. Eur Radiol Exp. 2018;2:36. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 413] [Cited by in RCA: 680] [Article Influence: 97.1] [Reference Citation Analysis (0)] |
| 8. | Valdora F, Houssami N, Rossi F, Calabrese M, Tagliafico AS. Rapid review: radiomics and breast cancer. Breast Cancer Res Treat. 2018;169:217-229. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 193] [Cited by in RCA: 179] [Article Influence: 25.6] [Reference Citation Analysis (0)] |
| 9. | Rogers W, Thulasi Seetha S, Refaee TAG, Lieverse RIY, Granzier RWY, Ibrahim A, Keek SA, Sanduleanu S, Primakov SP, Beuque MPL, Marcus D, van der Wiel AMA, Zerka F, Oberije CJG, van Timmeren JE, Woodruff HC, Lambin P. Radiomics: from qualitative to quantitative imaging. Br J Radiol. 2020;93:20190948. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 198] [Article Influence: 39.6] [Reference Citation Analysis (0)] |
| 10. | Antropova N, Huynh BQ, Giger ML. A deep feature fusion methodology for breast cancer diagnosis demonstrated on three imaging modality datasets. Med Phys. 2017;44:5162-5171. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 204] [Cited by in RCA: 225] [Article Influence: 28.1] [Reference Citation Analysis (0)] |
| 11. | Fusco R, Sansone M, Sansone C, Petrillo A. Segmentation and classification of breast lesions using dynamic and textural features in Dynamic Contrast Enhanced-Magnetic Resonance Imaging. IEEE. 2012;. [DOI] [Full Text] |
| 12. | Nie K, Chen JH, Yu HJ, Chu Y, Nalcioglu O, Su MY. Quantitative analysis of lesion morphology and texture features for diagnostic prediction in breast MRI. Acad Radiol. 2008;15:1513-1525. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 198] [Cited by in RCA: 168] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 13. | Zhang Q, Peng Y, Liu W, Bai J, Zheng J, Yang X, Zhou L. Radiomics Based on Multimodal MRI for the Differential Diagnosis of Benign and Malignant Breast Lesions. J Magn Reson Imaging. 2020;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 66] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 14. | Waugh SA, Purdie CA, Jordan LB, Vinnicombe S, Lerski RA, Martin P, Thompson AM. Magnetic resonance imaging texture analysis classification of primary breast cancer. Eur Radiol. 2016;26:322-330. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 137] [Cited by in RCA: 142] [Article Influence: 14.2] [Reference Citation Analysis (0)] |
| 15. | Kim JH, Ko ES, Lim Y, Lee KS, Han BK, Ko EY, Hahn SY, Nam SJ. Breast Cancer Heterogeneity: MR Imaging Texture Analysis and Survival Outcomes. Radiology. 2017;282:665-675. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 151] [Cited by in RCA: 181] [Article Influence: 20.1] [Reference Citation Analysis (0)] |
| 16. | Gibbs P, Turnbull LW. Textural analysis of contrast-enhanced MR images of the breast. Magn Reson Med. 2003;50:92-98. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 208] [Cited by in RCA: 194] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 17. | Kim SG, Freed M, Leite APK, Zhang J, Seuss C, Moy L. Separation of benign and malignant breast lesions using dynamic contrast enhanced MRI in a biopsy cohort. J Magn Reson Imaging. 2017;45:1385-1393. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 20] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 18. | Nagarajan MB, Huber MB, Schlossbauer T, Leinsinger G, Krol A, Wismüller A. Classification of Small Lesions in Breast MRI: Evaluating The Role of Dynamically Extracted Texture Features Through Feature Selection. J Med Biol Eng. 2013;33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 47] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 19. | Gibbs P, Onishi N, Sadinski M, Gallagher KM, Hughes M, Martinez DF, Morris EA, Sutton EJ. Characterization of Sub-1 cm Breast Lesions Using Radiomics Analysis. J Magn Reson Imaging. 2019;50:1468-1477. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 31] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 20. | Parekh VS, Jacobs MA. Integrated radiomic framework for breast cancer and tumor biology using advanced machine learning and multiparametric MRI. NPJ Breast Cancer. 2017;3:43. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 95] [Cited by in RCA: 108] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 21. | Bhooshan N, Giger M, Lan L, Li H, Marquez A, Shimauchi A, Newstead GM. Combined use of T2-weighted MRI and T1-weighted dynamic contrast-enhanced MRI in the automated analysis of breast lesions. Magn Reson Med. 2011;66:555-564. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 27] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 22. | Bickelhaupt S, Paech D, Kickingereder P, Steudle F, Lederer W, Daniel H, Götz M, Gählert N, Tichy D, Wiesenfarth M, Laun FB, Maier-Hein KH, Schlemmer HP, Bonekamp D. Prediction of malignancy by a radiomic signature from contrast agent-free diffusion MRI in suspicious breast lesions found on screening mammography. J Magn Reson Imaging. 2017;46:604-616. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 103] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 23. | Hu B, Xu K, Zhang Z, Chai R, Li S, Zhang L. A radiomic nomogram based on an apparent diffusion coefficient map for differential diagnosis of suspicious breast findings. Chin J Cancer Res. 2018;30:432-438. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 20] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 24. | Bickelhaupt S, Jaeger PF, Laun FB, Lederer W, Daniel H, Kuder TA, Wuesthof L, Paech D, Bonekamp D, Radbruch A, Delorme S, Schlemmer HP, Steudle FH, Maier-Hein KH. Radiomics Based on Adapted Diffusion Kurtosis Imaging Helps to Clarify Most Mammographic Findings Suspicious for Cancer. Radiology. 2018;287:761-770. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 79] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 25. | Zhou J, Zhang Y, Chang KT, Lee KE, Wang O, Li J, Lin Y, Pan Z, Chang P, Chow D, Wang M, Su MY. Diagnosis of Benign and Malignant Breast Lesions on DCE-MRI by Using Radiomics and Deep Learning With Consideration of Peritumor Tissue. J Magn Reson Imaging. 2020;51:798-809. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 140] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 26. | Holli K, Lääperi AL, Harrison L, Luukkaala T, Toivonen T, Ryymin P, Dastidar P, Soimakallio S, Eskola H. Characterization of breast cancer types by texture analysis of magnetic resonance images. Acad Radiol. 2010;17:135-141. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 108] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 27. | Chou SS, Gombos EC, Chikarmane SA, Giess CS, Jayender J. Computer-aided heterogeneity analysis in breast MR imaging assessment of ductal carcinoma in situ: Correlating histologic grade and receptor status. J Magn Reson Imaging. 2017;46:1748-1759. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 25] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 28. | Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thürlimann B, Senn HJ; Panel members. Strategies for subtypes--dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011;22:1736-1747. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2597] [Cited by in RCA: 2779] [Article Influence: 198.5] [Reference Citation Analysis (0)] |
| 29. | Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thürlimann B, Senn HJ; Panel members. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol. 2013;24:2206-2223. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2486] [Cited by in RCA: 2665] [Article Influence: 222.1] [Reference Citation Analysis (0)] |
| 30. | Balic M, Thomssen C, Würstlein R, Gnant M, Harbeck N. St. Gallen/Vienna 2019: A Brief Summary of the Consensus Discussion on the Optimal Primary Breast Cancer Treatment. Breast Care (Basel). 2019;14:103-110. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 126] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 31. | Öztürk VS, Polat YD, Soyder A, Tanyeri A, Karaman CZ, Taşkın F. The Relationship Between MRI Findings and Molecular Subtypes in Women With Breast Cancer. Curr Probl Diagn Radiol. 2019;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 12] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 32. | Guiu S, Michiels S, André F, Cortes J, Denkert C, Di Leo A, Hennessy BT, Sorlie T, Sotiriou C, Turner N, Van de Vijver M, Viale G, Loi S, Reis-Filho JS. Molecular subclasses of breast cancer: how do we define them? The IMPAKT 2012 Working Group Statement. Ann Oncol. 2012;23:2997-3006. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 212] [Cited by in RCA: 211] [Article Influence: 16.2] [Reference Citation Analysis (0)] |
| 33. | Giuliano AE, Connolly JL, Edge SB, Mittendorf EA, Rugo HS, Solin LJ, Weaver DL, Winchester DJ, Hortobagyi GN. Breast Cancer-Major changes in the American Joint Committee on Cancer eighth edition cancer staging manual. CA Cancer J Clin. 2017;67:290-303. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 477] [Cited by in RCA: 616] [Article Influence: 77.0] [Reference Citation Analysis (0)] |
| 34. | Net JM, Whitman GJ, Morris E, Brandt KR, Burnside ES, Giger ML, Ganott M, Sutton EJ, Zuley ML, Rao A. Relationships Between Human-Extracted MRI Tumor Phenotypes of Breast Cancer and Clinical Prognostic Indicators Including Receptor Status and Molecular Subtype. Curr Probl Diagn Radiol. 2019;48:467-472. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 11] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 35. | Jinguji M, Kajiya Y, Kamimura K, Nakajo M, Sagara Y, Takahama T, Ando M, Rai Y, Sagara Y, Ohi Y, Yoshida H. Rim enhancement of breast cancers on contrast-enhanced MR imaging: relationship with prognostic factors. Breast Cancer. 2006;13:64-73. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 32] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 36. | Blaschke E, Abe H. MRI phenotype of breast cancer: Kinetic assessment for molecular subtypes. J Magn Reson Imaging. 2015;42:920-924. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 56] [Cited by in RCA: 57] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 37. | Grimm LJ, Zhang J, Baker JA, Soo MS, Johnson KS, Mazurowski MA. Relationships Between MRI Breast Imaging-Reporting and Data System (BI-RADS) Lexicon Descriptors and Breast Cancer Molecular Subtypes: Internal Enhancement is Associated with Luminal B Subtype. Breast J. 2017;23:579-582. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 31] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 38. | Martincich L, Deantoni V, Bertotto I, Redana S, Kubatzki F, Sarotto I, Rossi V, Liotti M, Ponzone R, Aglietta M, Regge D, Montemurro F. Correlations between diffusion-weighted imaging and breast cancer biomarkers. Eur Radiol. 2012;22:1519-1528. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 167] [Cited by in RCA: 177] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 39. | Montemezzi S, Camera L, Giri MG, Pozzetto A, Caliò A, Meliadò G, Caumo F, Cavedon C. Is there a correlation between 3T multiparametric MRI and molecular subtypes of breast cancer? Eur J Radiol. 2018;108:120-127. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 33] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 40. | Shin HJ, Baek HM, Cha JH, Kim HH. Evaluation of breast cancer using proton MR spectroscopy: total choline peak integral and signal-to-noise ratio as prognostic indicators. AJR Am J Roentgenol. 2012;198:W488-W497. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 32] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 41. | Sah RG, Sharma U, Parshad R, Seenu V, Mathur SR, Jagannathan NR. Association of estrogen receptor, progesterone receptor, and human epidermal growth factor receptor 2 status with total choline concentration and tumor volume in breast cancer patients: an MRI and in vivo proton MRS study. Magn Reson Med. 2012;68:1039-1047. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 30] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 42. | Sutton EJ, Dashevsky BZ, Oh JH, Veeraraghavan H, Apte AP, Thakur SB, Morris EA, Deasy JO. Breast cancer molecular subtype classifier that incorporates MRI features. J Magn Reson Imaging. 2016;44:122-129. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 101] [Article Influence: 11.2] [Reference Citation Analysis (0)] |
| 43. | Agner SC, Rosen MA, Englander S, Tomaszewski JE, Feldman MD, Zhang P, Mies C, Schnall MD, Madabhushi A. Computerized image analysis for identifying triple-negative breast cancers and differentiating them from other molecular subtypes of breast cancer on dynamic contrast-enhanced MR images: a feasibility study. Radiology. 2014;272:91-99. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 107] [Cited by in RCA: 115] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 44. | Fan M, Yuan W, Zhao W, Xu M, Wang S, Gao X, Li L. Joint Prediction of Breast Cancer Histological Grade and Ki-67 Expression Level Based on DCE-MRI and DWI Radiomics. IEEE J Biomed Health Inform. 2020;24:1632-1642. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 73] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 45. | Wu J, Sun X, Wang J, Cui Y, Kato F, Shirato H, Ikeda DM, Li R. Identifying relations between imaging phenotypes and molecular subtypes of breast cancer: Model discovery and external validation. J Magn Reson Imaging. 2017;46:1017-1027. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 73] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 46. | Li H, Zhu Y, Burnside ES, Huang E, Drukker K, Hoadley KA, Fan C, Conzen SD, Zuley M, Net JM, Sutton E, Whitman GJ, Morris E, Perou CM, Ji Y, Giger ML. Quantitative MRI radiomics in the prediction of molecular classifications of breast cancer subtypes in the TCGA/TCIA data set. NPJ Breast Cancer. 2016;2:16012. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 195] [Cited by in RCA: 253] [Article Influence: 28.1] [Reference Citation Analysis (0)] |
| 47. | Chang RF, Chen HH, Chang YC, Huang CS, Chen JH, Lo CM. Quantification of breast tumor heterogeneity for ER status, HER2 status, and TN molecular subtype evaluation on DCE-MRI. Magn Reson Imaging. 2016;34:809-819. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 67] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 48. | Fan M, Li H, Wang S, Zheng B, Zhang J, Li L. Radiomic analysis reveals DCE-MRI features for prediction of molecular subtypes of breast cancer. PLoS One. 2017;12:e0171683. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 88] [Cited by in RCA: 107] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 49. | Wang J, Kato F, Oyama-Manabe N, Li R, Cui Y, Tha KK, Yamashita H, Kudo K, Shirato H. Identifying Triple-Negative Breast Cancer Using Background Parenchymal Enhancement Heterogeneity on Dynamic Contrast-Enhanced MRI: A Pilot Radiomics Study. PLoS One. 2015;10:e0143308. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 103] [Cited by in RCA: 108] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 50. | Braman N, Prasanna P, Whitney J, Singh S, Beig N, Etesami M, Bates DDB, Gallagher K, Bloch BN, Vulchi M, Turk P, Bera K, Abraham J, Sikov WM, Somlo G, Harris LN, Gilmore H, Plecha D, Varadan V, Madabhushi A. Association of Peritumoral Radiomics With Tumor Biology and Pathologic Response to Preoperative Targeted Therapy for HER2 (ERBB2)-Positive Breast Cancer. JAMA Netw Open. 2019;2:e192561. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 138] [Cited by in RCA: 242] [Article Influence: 40.3] [Reference Citation Analysis (0)] |
| 51. | Leithner D, Horvat JV, Marino MA, Bernard-Davila B, Jochelson MS, Ochoa-Albiztegui RE, Martinez DF, Morris EA, Thakur S, Pinker K. Radiomic signatures with contrast-enhanced magnetic resonance imaging for the assessment of breast cancer receptor status and molecular subtypes: initial results. Breast Cancer Res. 2019;21:106. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 84] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 52. | Holli-Helenius K, Salminen A, Rinta-Kiikka I, Koskivuo I, Brück N, Boström P, Parkkola R. MRI texture analysis in differentiating luminal A and luminal B breast cancer molecular subtypes - a feasibility study. BMC Med Imaging. 2017;17:69. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 59] [Cited by in RCA: 75] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 53. | Ma W, Ji Y, Qi L, Guo X, Jian X, Liu P. Breast cancer Ki67 expression prediction by DCE-MRI radiomics features. Clin Radiol. 2018;73:909.e1-909.e5. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 65] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 54. | Juan MW, Yu J, Peng GX, Jun LJ, Feng SP, Fang LP. Correlation between DCE-MRI radiomics features and Ki-67 expression in invasive breast cancer. Oncol Lett. 2018;16:5084-5090. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 19] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 55. | Liang C, Cheng Z, Huang Y, He L, Chen X, Ma Z, Huang X, Liang C, Liu Z. An MRI-based Radiomics Classifier for Preoperative Prediction of Ki-67 Status in Breast Cancer. Acad Radiol. 2018;25:1111-1117. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 79] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 56. | Zhang Y, Zhu Y, Zhang K, Liu Y, Cui J, Tao J, Wang Y, Wang S. Invasive ductal breast cancer: preoperative predict Ki-67 index based on radiomics of ADC maps. Radiol Med. 2020;125:109-116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 40] [Cited by in RCA: 65] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 57. | Tan H, Gan F, Wu Y, Zhou J, Tian J, Lin Y, Wang M. Preoperative Prediction of Axillary Lymph Node Metastasis in Breast Carcinoma Using Radiomics Features Based on the Fat-Suppressed T2 Sequence. Acad Radiol. 2019;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 47] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 58. | Dong Y, Feng Q, Yang W, Lu Z, Deng C, Zhang L, Lian Z, Liu J, Luo X, Pei S, Mo X, Huang W, Liang C, Zhang B, Zhang S. Preoperative prediction of sentinel lymph node metastasis in breast cancer based on radiomics of T2-weighted fat-suppression and diffusion-weighted MRI. Eur Radiol. 2018;28:582-591. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 153] [Cited by in RCA: 189] [Article Influence: 27.0] [Reference Citation Analysis (0)] |
| 59. | Liu C, Ding J, Spuhler K, Gao Y, Serrano Sosa M, Moriarty M, Hussain S, He X, Liang C, Huang C. Preoperative prediction of sentinel lymph node metastasis in breast cancer by radiomic signatures from dynamic contrast-enhanced MRI. J Magn Reson Imaging. 2019;49:131-140. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 94] [Cited by in RCA: 145] [Article Influence: 20.7] [Reference Citation Analysis (0)] |
| 60. | Liu Z, Feng B, Li C, Chen Y, Chen Q, Li X, Guan J, Chen X, Cui E, Li R, Li Z, Long W. Preoperative prediction of lymphovascular invasion in invasive breast cancer with dynamic contrast-enhanced-MRI-based radiomics. J Magn Reson Imaging. 2019;50:847-857. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 69] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 61. | Liu J, Sun D, Chen L, Fang Z, Song W, Guo D, Ni T, Liu C, Feng L, Xia Y, Zhang X, Li C. Radiomics Analysis of Dynamic Contrast-Enhanced Magnetic Resonance Imaging for the Prediction of Sentinel Lymph Node Metastasis in Breast Cancer. Front Oncol. 2019;9:980. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 87] [Cited by in RCA: 84] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 62. | Han L, Zhu Y, Liu Z, Yu T, He C, Jiang W, Kan Y, Dong D, Tian J, Luo Y. Radiomic nomogram for prediction of axillary lymph node metastasis in breast cancer. Eur Radiol. 2019;29:3820-3829. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 140] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 63. | Cui X, Wang N, Zhao Y, Chen S, Li S, Xu M, Chai R. Preoperative Prediction of Axillary Lymph Node Metastasis in Breast Cancer using Radiomics Features of DCE-MRI. Sci Rep. 2019;9:2240. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 57] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 64. | Burnside ES, Drukker K, Li H, Bonaccio E, Zuley M, Ganott M, Net JM, Sutton EJ, Brandt KR, Whitman GJ, Conzen SD, Lan L, Ji Y, Zhu Y, Jaffe CC, Huang EP, Freymann JB, Kirby JS, Morris EA, Giger ML. Using computer-extracted image phenotypes from tumors on breast magnetic resonance imaging to predict breast cancer pathologic stage. Cancer. 2016;122:748-757. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 51] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 65. | Reig B, Heacock L, Geras KJ, Moy L. Machine learning in breast MRI. J Magn Reson Imaging. 2019;. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 96] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 66. | Schacht DV, Drukker K, Pak I, Abe H, Giger ML. Using quantitative image analysis to classify axillary lymph nodes on breast MRI: a new application for the Z 0011 Era. Eur J Radiol. 2015;84:392-397. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 24] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 67. | Fusco R, Sansone M, Granata V, Di Bonito M, Avino F, Catalano O, Botti G, Petrillo A. Use of Quantitative Morphological and Functional Features for Assessment of Axillary Lymph Node in Breast Dynamic Contrast-Enhanced Magnetic Resonance Imaging. Biomed Res Int. 2018;2018:2610801. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 21] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 68. | Thompson AM, Moulder-Thompson SL. Neoadjuvant treatment of breast cancer. Ann Oncol. 2012;23 Suppl 10:x231-x236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 174] [Cited by in RCA: 172] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 69. | Haque W, Verma V, Hatch S, Suzanne Klimberg V, Brian Butler E, Teh BS. Response rates and pathologic complete response by breast cancer molecular subtype following neoadjuvant chemotherapy. Breast Cancer Res Treat. 2018;170:559-567. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 113] [Cited by in RCA: 209] [Article Influence: 29.9] [Reference Citation Analysis (0)] |
| 70. | Yuan Y, Chen XS, Liu SY, Shen KW. Accuracy of MRI in prediction of pathologic complete remission in breast cancer after preoperative therapy: a meta-analysis. AJR Am J Roentgenol. 2010;195:260-268. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 112] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 71. | Marinovich ML, Houssami N, Macaskill P, Sardanelli F, Irwig L, Mamounas EP, von Minckwitz G, Brennan ME, Ciatto S. Meta-analysis of magnetic resonance imaging in detecting residual breast cancer after neoadjuvant therapy. J Natl Cancer Inst. 2013;105:321-333. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 235] [Cited by in RCA: 274] [Article Influence: 22.8] [Reference Citation Analysis (0)] |
| 72. | Hylton NM, Blume JD, Bernreuter WK, Pisano ED, Rosen MA, Morris EA, Weatherall PT, Lehman CD, Newstead GM, Polin S, Marques HS, Esserman LJ, Schnall MD; ACRIN 6657 Trial Team and I-SPY 1 TRIAL Investigators. Locally advanced breast cancer: MR imaging for prediction of response to neoadjuvant chemotherapy--results from ACRIN 6657/I-SPY TRIAL. Radiology. 2012;263:663-672. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 299] [Cited by in RCA: 364] [Article Influence: 28.0] [Reference Citation Analysis (0)] |
| 73. | Yu N, Leung VWY, Meterissian S. MRI Performance in Detecting pCR After Neoadjuvant Chemotherapy by Molecular Subtype of Breast Cancer. World J Surg. 2019;43:2254-2261. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 12] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 74. | Michoux N, Van den Broeck S, Lacoste L, Fellah L, Galant C, Berlière M, Leconte I. Texture analysis on MR images helps predicting non-response to NAC in breast cancer. BMC Cancer. 2015;15:574. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 54] [Cited by in RCA: 51] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 75. | Wiechmann L, Sampson M, Stempel M, Jacks LM, Patil SM, King T, Morrow M. Presenting features of breast cancer differ by molecular subtype. Ann Surg Oncol. 2009;16:2705-2710. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 137] [Cited by in RCA: 146] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 76. | Xiong Q, Zhou X, Liu Z, Lei C, Yang C, Yang M, Zhang L, Zhu T, Zhuang X, Liang C, Liu Z, Tian J, Wang K. Multiparametric MRI-based radiomics analysis for prediction of breast cancers insensitive to neoadjuvant chemotherapy. Clin Transl Oncol. 2020;22:50-59. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 61] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 77. | Cain EH, Saha A, Harowicz MR, Marks JR, Marcom PK, Mazurowski MA. Multivariate machine learning models for prediction of pathologic response to neoadjuvant therapy in breast cancer using MRI features: a study using an independent validation set. Breast Cancer Res Treat. 2019;173:455-463. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 116] [Article Influence: 16.6] [Reference Citation Analysis (0)] |
| 78. | Granzier RWY, van Nijnatten TJA, Woodruff HC, Smidt ML, Lobbes MBI. Exploring breast cancer response prediction to neoadjuvant systemic therapy using MRI-based radiomics: A systematic review. Eur J Radiol. 2019;121:108736. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 61] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 79. | Golden DI, Lipson JA, Telli ML, Ford JM, Rubin DL. Dynamic contrast-enhanced MRI-based biomarkers of therapeutic response in triple-negative breast cancer. J Am Med Inform Assoc. 2013;20:1059-1066. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 56] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 80. | Teruel JR, Heldahl MG, Goa PE, Pickles M, Lundgren S, Bathen TF, Gibbs P. Dynamic contrast-enhanced MRI texture analysis for pretreatment prediction of clinical and pathological response to neoadjuvant chemotherapy in patients with locally advanced breast cancer. NMR Biomed. 2014;27:887-896. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 96] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 81. | Thibault G, Tudorica A, Afzal A, Chui SY, Naik A, Troxell ML, Kemmer KA, Oh KY, Roy N, Jafarian N, Holtorf ML, Huang W, Song X. DCE-MRI Texture Features for Early Prediction of Breast Cancer Therapy Response. Tomography. 2017;3:23-32. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 53] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 82. | Parikh J, Selmi M, Charles-Edwards G, Glendenning J, Ganeshan B, Verma H, Mansi J, Harries M, Tutt A, Goh V. Changes in primary breast cancer heterogeneity may augment midtreatment MR imaging assessment of response to neoadjuvant chemotherapy. Radiology. 2014;272:100-112. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 109] [Cited by in RCA: 108] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 83. | Cao K, Zhao B, Li XT, Li YL, Sun YS. Texture Analysis of Dynamic Contrast-Enhanced MRI in Evaluating Pathologic Complete Response (pCR) of Mass-Like Breast Cancer after Neoadjuvant Therapy. J Oncol. 2019;2019:4731532. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 7] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 84. | Wu J, Cao G, Sun X, Lee J, Rubin DL, Napel S, Kurian AW, Daniel BL, Li R. Intratumoral Spatial Heterogeneity at Perfusion MR Imaging Predicts Recurrence-free Survival in Locally Advanced Breast Cancer Treated with Neoadjuvant Chemotherapy. Radiology. 2018;288:26-35. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 131] [Article Influence: 18.7] [Reference Citation Analysis (0)] |
| 85. | Fusco R, Granata V, Maio F, Sansone M, Petrillo A. Textural radiomic features and time-intensity curve data analysis by dynamic contrast-enhanced MRI for early prediction of breast cancer therapy response: preliminary data. Eur Radiol Exp. 2020;4:8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 13] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 86. | Mohammed ZM, McMillan DC, Edwards J, Mallon E, Doughty JC, Orange C, Going JJ. The relationship between lymphovascular invasion and angiogenesis, hormone receptors, cell proliferation and survival in patients with primary operable invasive ductal breast cancer. BMC Clin Pathol. 2013;13:31. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 37] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 87. | Braman NM, Etesami M, Prasanna P, Dubchuk C, Gilmore H, Tiwari P, Plecha D, Madabhushi A. Intratumoral and peritumoral radiomics for the pretreatment prediction of pathological complete response to neoadjuvant chemotherapy based on breast DCE-MRI. Breast Cancer Res. 2017;19:57. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 267] [Cited by in RCA: 439] [Article Influence: 54.9] [Reference Citation Analysis (0)] |
| 88. | Zhou J, Lu J, Gao C, Zeng J, Zhou C, Lai X, Cai W, Xu M. Predicting the response to neoadjuvant chemotherapy for breast cancer: wavelet transforming radiomics in MRI. BMC Cancer. 2020;20:100. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 78] [Article Influence: 15.6] [Reference Citation Analysis (0)] |
| 89. | Liu Z, Li Z, Qu J, Zhang R, Zhou X, Li L, Sun K, Tang Z, Jiang H, Li H, Xiong Q, Ding Y, Zhao X, Wang K, Liu Z, Tian J. Radiomics of Multiparametric MRI for Pretreatment Prediction of Pathologic Complete Response to Neoadjuvant Chemotherapy in Breast Cancer: A Multicenter Study. Clin Cancer Res. 2019;25:3538-3547. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 155] [Cited by in RCA: 316] [Article Influence: 52.7] [Reference Citation Analysis (0)] |
| 90. | Chen X, Chen X, Yang J, Li Y, Fan W, Yang Z. Combining Dynamic Contrast-Enhanced Magnetic Resonance Imaging and Apparent Diffusion Coefficient Maps for a Radiomics Nomogram to Predict Pathological Complete Response to Neoadjuvant Chemotherapy in Breast Cancer Patients. J Comput Assist Tomogr. 2020;44:275-283. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 33] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 91. | Hyun SJ, Kim EK, Moon HJ, Yoon JH, Kim MJ. Preoperative axillary lymph node evaluation in breast cancer patients by breast magnetic resonance imaging (MRI): Can breast MRI exclude advanced nodal disease? Eur Radiol. 2016;26:3865-3873. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 49] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 92. | Mattingly AE, Mooney B, Lin HY, Kiluk JV, Khakpour N, Hoover SJ, Laronga C, Lee MC. Magnetic Resonance Imaging for Axillary Breast Cancer Metastasis in the Neoadjuvant Setting: A Prospective Study. Clin Breast Cancer. 2017;17:180-187. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 15] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 93. | Ha SM, Cha JH, Kim HH, Shin HJ, Chae EY, Choi WJ. Diagnostic performance of breast ultrasonography and MRI in the prediction of lymph node status after neoadjuvant chemotherapy for breast cancer. Acta Radiol. 2017;58:1198-1205. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 13] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 94. | Ha R, Chang P, Karcich J, Mutasa S, Fardanesh R, Wynn RT, Liu MZ, Jambawalikar S. Axillary Lymph Node Evaluation Utilizing Convolutional Neural Networks Using MRI Dataset. J Digit Imaging. 2018;31:851-856. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 47] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 95. | Ha R, Chin C, Karcich J, Liu MZ, Chang P, Mutasa S, Pascual Van Sant E, Wynn RT, Connolly E, Jambawalikar S. Prior to Initiation of Chemotherapy, Can We Predict Breast Tumor Response? Deep Learning Convolutional Neural Networks Approach Using a Breast MRI Tumor Dataset. J Digit Imaging. 2019;32:693-701. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 74] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 96. | Ha R, Chang P, Karcich J, Mutasa S, Van Sant EP, Connolly E, Chin C, Taback B, Liu MZ, Jambawalikar S. Predicting Post Neoadjuvant Axillary Response Using a Novel Convolutional Neural Network Algorithm. Ann Surg Oncol. 2018;25:3037-3043. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 25] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 97. | Ashraf AB, Daye D, Gavenonis S, Mies C, Feldman M, Rosen M, Kontos D. Identification of intrinsic imaging phenotypes for breast cancer tumors: preliminary associations with gene expression profiles. Radiology. 2014;272:374-384. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 113] [Cited by in RCA: 114] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 98. | Ganeshan B, Panayiotou E, Burnand K, Dizdarevic S, Miles K. Tumour heterogeneity in non-small cell lung carcinoma assessed by CT texture analysis: a potential marker of survival. Eur Radiol. 2012;22:796-802. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 364] [Cited by in RCA: 393] [Article Influence: 28.1] [Reference Citation Analysis (0)] |
| 99. | Li H, Zhu Y, Burnside ES, Drukker K, Hoadley KA, Fan C, Conzen SD, Whitman GJ, Sutton EJ, Net JM, Ganott M, Huang E, Morris EA, Perou CM, Ji Y, Giger ML. MR Imaging Radiomics Signatures for Predicting the Risk of Breast Cancer Recurrence as Given by Research Versions of MammaPrint, Oncotype DX, and PAM50 Gene Assays. Radiology. 2016;281:382-391. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 290] [Cited by in RCA: 355] [Article Influence: 39.4] [Reference Citation Analysis (0)] |
| 100. | Ha R, Chang P, Mutasa S, Karcich J, Goodman S, Blum E, Kalinsky K, Liu MZ, Jambawalikar S. Convolutional Neural Network Using a Breast MRI Tumor Dataset Can Predict Oncotype Dx Recurrence Score. J Magn Reson Imaging. 2019;49:518-524. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 48] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 101. | Harris LN, Ismaila N, McShane LM, Andre F, Collyar DE, Gonzalez-Angulo AM, Hammond EH, Kuderer NM, Liu MC, Mennel RG, Van Poznak C, Bast RC, Hayes DF; American Society of Clinical Oncology. Use of Biomarkers to Guide Decisions on Adjuvant Systemic Therapy for Women With Early-Stage Invasive Breast Cancer: American Society of Clinical Oncology Clinical Practice Guideline. J Clin Oncol. 2016;34:1134-1150. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 594] [Cited by in RCA: 594] [Article Influence: 66.0] [Reference Citation Analysis (0)] |
| 102. | Saha A, Harowicz MR, Wang W, Mazurowski MA. A study of association of Oncotype DX recurrence score with DCE-MRI characteristics using multivariate machine learning models. J Cancer Res Clin Oncol. 2018;144:799-807. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 30] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 103. | Park H, Lim Y, Ko ES, Cho HH, Lee JE, Han BK, Ko EY, Choi JS, Park KW. Radiomics Signature on Magnetic Resonance Imaging: Association with Disease-Free Survival in Patients with Invasive Breast Cancer. Clin Cancer Res. 2018;24:4705-4714. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 178] [Article Influence: 25.4] [Reference Citation Analysis (0)] |
| 104. | Koh J, Lee E, Han K, Kim S, Kim DK, Kwak JY, Yoon JH, Moon HJ. Three-dimensional radiomics of triple-negative breast cancer: Prediction of systemic recurrence. Sci Rep. 2020;10:2976. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 105. | Kim S, Kim MJ, Kim EK, Yoon JH, Park VY. MRI Radiomic Features: Association with Disease-Free Survival in Patients with Triple-Negative Breast Cancer. Sci Rep. 2020;10:3750. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 25] [Article Influence: 5.0] [Reference Citation Analysis (0)] |