Published online Oct 28, 2021. doi: 10.35712/aig.v2.i5.124
Peer-review started: June 6, 2021
First decision: June 23, 2021
Revised: June 26, 2021
Accepted: September 1, 2021
Article in press: September 1, 2021
Published online: October 28, 2021
Processing time: 143 Days and 7.9 Hours
This minireview discusses the benefits and pitfalls of machine learning, and artificial intelligence in upper gastrointestinal endoscopy for the detection and characterization of neoplasms. We have reviewed the literature for relevant publications on the topic using PubMed, IEEE, Science Direct, and Google Scholar databases. We discussed the phases of machine learning and the importance of advanced imaging techniques in upper gastrointestinal endoscopy and its association with artificial intelligence.
Core Tip: This minireview aims to explore an important topic; the role of artificial intelligence in upper gastrointestinal (GI) endoscopy detection of cancer. We tried to delineate the most common obstacles encountered when trying to implement artificial intelligence in upper GI endoscopy for cancer detection and characterization. Moreover, we tried to outline the future prospects of this technique, along with its benefits, and uncertainties. This topic summarizes the wide scope for integration of artificial intelligence, between the practicing physicians and the computational engineers and how their collaboration could provide a better healthcare services.
- Citation: El-Nakeep S, El-Nakeep M. Artificial intelligence for cancer detection in upper gastrointestinal endoscopy, current status, and future aspirations. Artif Intell Gastroenterol 2021; 2(5): 124-132
- URL: https://www.wjgnet.com/2644-3236/full/v2/i5/124.htm
- DOI: https://dx.doi.org/10.35712/aig.v2.i5.124
Upper gastrointestinal (GI) cancers affecting the esophagus and stomach are responsible for more than one and half million annual deaths worldwide. Both are considered aggressive cancers, discovered mostly at an advanced stage, when curative measures are no longer applicable[1].
The current standard method for diagnosis of upper GI cancers is upper GI endoscopy and biopsy, using a white light endoscopy. The most common upper GI cancers encountered are esophageal adenocarcinoma, Barrett's esophagus (BE) and gastric cancer[2]. Artificial intelligence (AI) could add more accuracy to early cancerous and precancerous lesion detection in the upper GI during endoscopic evaluation[3].
Regardless of the great progress of AI in colonoscopy examinations, the integration of AI in upper GI endoscopy is still a new area of research with only a few pilot studies available, mostly due to unavailability of large datasets annotating upper GI cancers[2].
A recent meta-analysis examined the effect of AI in detecting Helicobacter pylori (H. pylori) infection during upper GI endoscopy, and found eight studies with pooled sensitivity of 87%, and specificity of 86%[4]. Moreover, another study combined the effect of neoplasm detection and H. pylori infection status, and found twenty-three studies with high pooled diagnostic accuracy in upper GI neoplasms; 96 in gastric cancer, 96% in BE, 88% in squamous esophagus and 92% in H. pylori detection[5].
The miss rate of detecting upper GI cancers reaches 11.3% according to a meta-analysis by Menon and Trudgill[6], and even higher rates could be observed in superficial neoplasms, reaching 75% (i.e., gastric superficial neoplasia)[7]. According to a recent meta-analysis by Arribas et al[3], using AI integrated upper GI endoscopy yielded pooled sensitivity of 90%, and specificity of 89% for detection of neoplastic lesions, independent of the type of neoplasia (whether esophageal adenocarcinoma, BE, or gastric adenocarcinoma).
Expert sensitivity and specificity criteria in detecting the upper GI tumors differ from the detection and characterization of colorectal polyps for a few reasons. First, due to over-specialization of certain types of upper GI cancers according to the geographical prevalence of the cancer, for example, in the gastroenterologist’s practice, resulting in limited training for the detection of non-prevalent types of cancers. AI integrated systems don’t suffer the same geographical bias, thus offering better detection independent of the prevalence of GI cancer types[3]. Colon cancer preva
The second reason, lesions that are minimal (in size or in depth) or hard to visualize by the inexperienced endoscopist, could be easily detected using the AI assistance[2]. Furthermore, gastric cancer lesions can be masked after eradication of H. pylori, this masking is due to regression of the mucosal elevation (decrease in its height) caused by the regression of chronic inflammatory process of H. pylori infection, or due to the coverage of the neoplastic area with atypical mucosa or even healthy columnar mucosa[8,9]. Advanced imaging techniques[10], when associated with AI, might help in detection of these masked neoplastic lesions.
Third reason being that training is not adequate in postgraduate courses, either because of insufficient interest (due to different cancer prevalence), or insufficient resources (especially for the computer simulation programs)[11]. However, an AI cumulative sensitivity of 91% for early-stage neoplasia proves that AI integrated systems will increase the efficacy of diagnostic upper GI endoscopy immensely. Thus, there is an urgent need for AI implementation in the clinical setting, even more urgent than the lower GI colonoscopy. Diagnostic settings have the issue of less experienced endoscopists compared to intervention intended settings, so early cancerous lesions tend to be easily overlooked (undetected)[3]. This of course will not eliminate the need for experienced endoscopists; however, integration with AI will have the best yield[12], considering that most of the upper GI lesions are non-polypoid which require higher level of skills for detection than colorectal cancer.
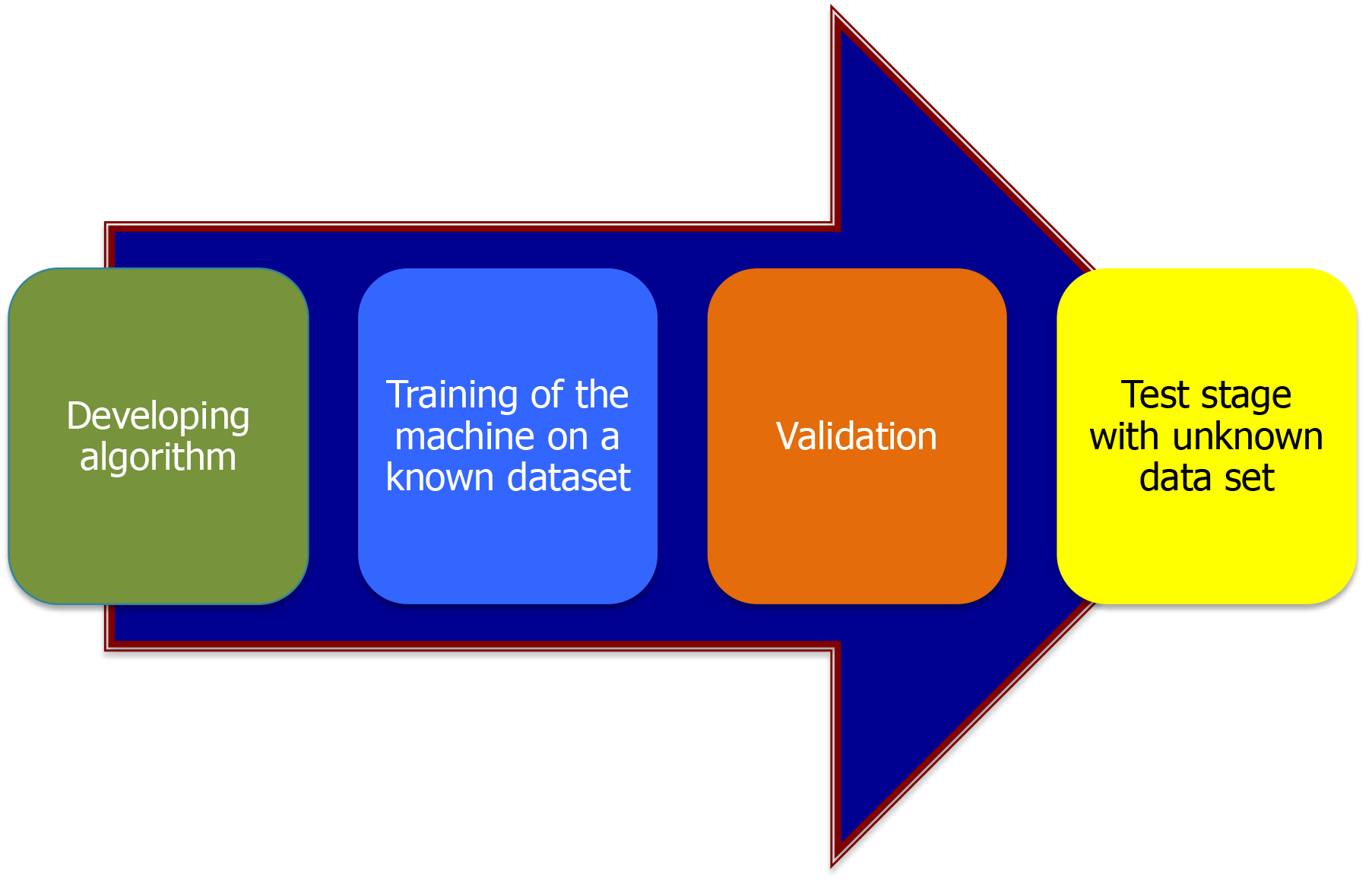
Machine learning (ML) must pass through multiple phases for validation in both training and testing (as shown in Figure 1). The AI used in endoscopy is ML, the most prevalent type of ML is deep learning (DL).
The first wave of AI was logic based handcrafted knowledge. In this logic-based handcrafted algorithms were developed separately for each task. This allowed the reasoning behind decisions of the first wave to be quite high, because every step of decision was handcrafted. However, the machine was unable to learn. The second wave of AI (the current wave) is the statistical ML in which the machines can learn from data, with an easily implemented learning algorithm, to generate a model used to carry out decisions. This eliminates the difficult part of designing and implementing a task-specific algorithm. While this raised the level of ML, it also caused a huge decline in the reasoning for the decisions. This means that the reasoning behind a wrong decision becomes hard to identify, rendering the algorithm a black box. The best way to avoid highly wrong decision rates is for provide a large amount of variable data to the machine to learn from[13].
ML passes through many phases. First phase is the training phase; where an annotated dataset is used to train the ML system, and then validated by determining the number of images it correctly identified. Second is the testing phase where a non-annotated dataset is given to the ML system to examine its diagnostic capabilities in comparison to experts in the field, and then using this ML system in a clinical setting, either in real time or in prospective trials to evaluate its performance in a real-world clinical setting.
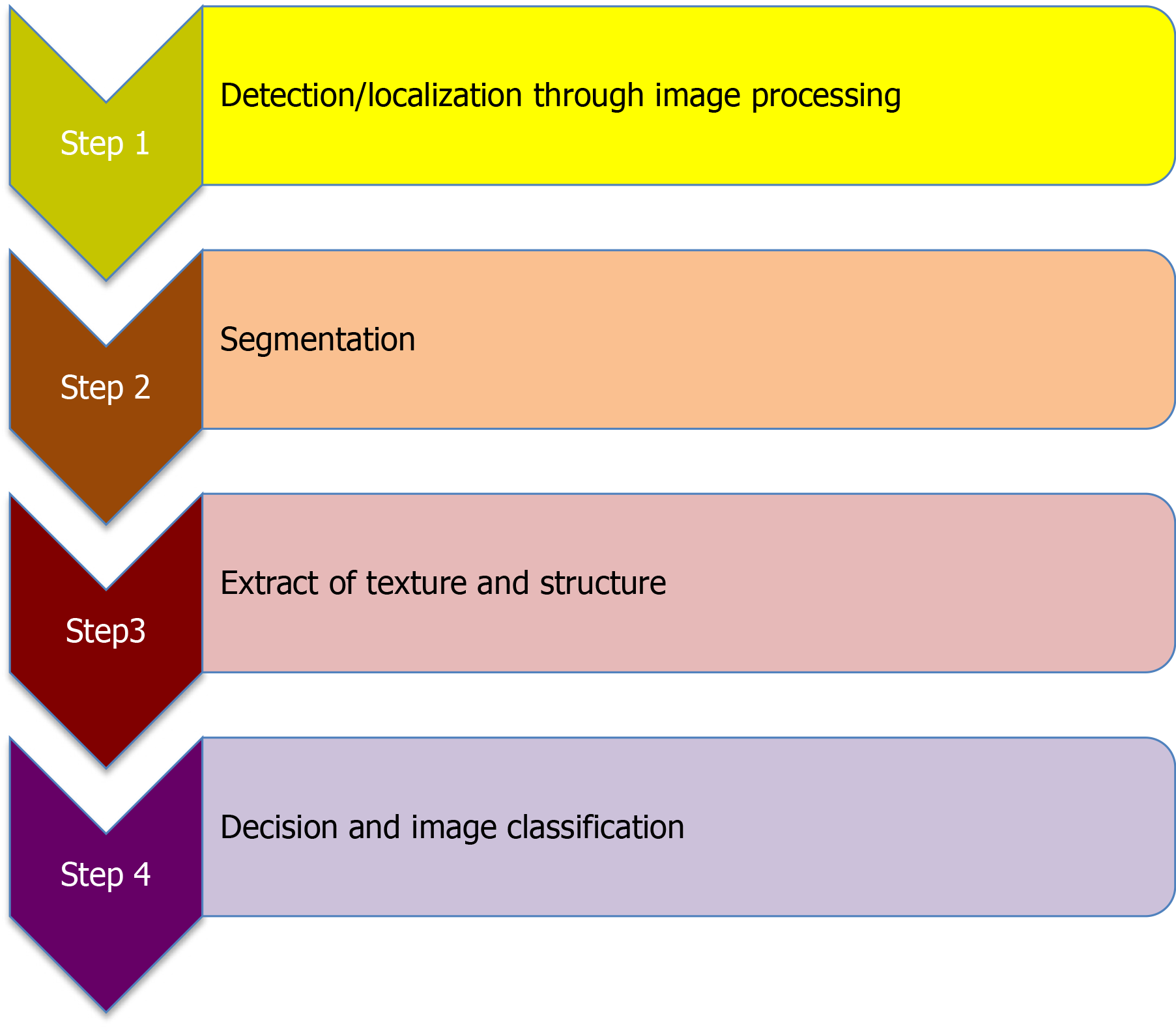
There are two types of gastric lesion examinations identified during upper GI endoscopy using AI: (as shown in Figure 2): (1) Lesion detection (to know whether it is present or absent) and localization (to know its exact location in the GI tract); and (2) Lesion characterization (to assess its histological prediction).
The first type uses images with low or moderate quality, but the second type uses advanced optical diagnostic tools including: Narrow band imaging (NBI), chromoendoscopy, endocytoscopy, optical magnification, among others[14,15]. All types use semi-automatic identification, where the endoscopist delineates the affected area and centers the polyp near the endoscope lens for better visualization[14]. Invasion depth has been successfully predicted (with 89% diagnostic accuracy) through coding systems that are not very complicated, a proposed implementation of automated DL models in gastric cancers. Furthermore, another proposed implementation of a modified version by the same author is faster by 13 min in the test stage on unknown data, but has a slightly lower accuracy of 82%, with similar performance to experts and higher than trainees[16].
If feasibility and usefulness of non-real time can be proved, then technical feasibility of real time is achievable, with an increased degree of sophistication of implementation and cost. Improvement of this real time feasibility could be accomplished through software programming of graphic processing unit (GPU) and central processing unit (CPU), along with implementation of specialized hardware systems.
Most implementations in AI use DL algorithms such as convolutional neural network (CNN). Wu et al[17] did the only randomized controlled trial (RCT) available on the topic. The team examined the diagnostic accuracy of their AI system using a deep convolution neural network. The system aimed to decrease the blind spots during upper GI endoscopy[17]. Unfortunately, they only examined images of benign and malignant lesions, not during a real time endoscopy performance.
Comparing white light alone vs linked color imaging showed that endoscopists had a lower miss rate with linked color imaging (30.7% vs 64.9%) in detecting early gastric cancers post-H. pylori eradication[18]. Linked color imaging is a technique that enhances the color range and brightness of images, developed by Fujifilm, Tokyo, Japan[19].
White light for detection of upper GI neoplasms is the most common and the standard technique. Other methods using advanced high-quality imaging are becoming increasingly available in most endoscopy centers. These advanced imaging techniques increase the sensitivity and specificity of diagnostic accuracy, especially in BE. There is a noticed "synergy" between AI integrated systems and advanced imaging techniques. On the other hand, the bias in having good quality images is apparent when identifying artifacts and lighting errors as cancerous lesions, or as called "spectrum bias" (this is a systematic error, where the data used do not represent the patients in question). This is equal in AI and humans[20,21].
While, dye-based imaging enhanced endoscopy (IEE) uses a dye to enhance detection of neoplastic lesions, this might not be helpful for examining a wide tract for lesions, nor for spraying the whole GI tract with dye. However, equipment enhanced IEE (eIEE) solves these problems. eIEE was originally classified into lightening-only techniques along with blue laser imaging (BLI), BLI-bright and NBI (Olympus), autofluorescence imaging (Olympus), and post-processing-only techniques such as: Flexible spectral image color enhancement - (Fujifilm) and iSCAN - (Pentax), all from Tokyo, Japan[22,23].
LCI merges the two techniques by low frequency intensity light, red color extraction, and variation enhancement in a red-green-blue color space digital image post-processing. The post-processing system has three modes of color enhancement (A, B and C) with varying grades. This yields enhancement of hemoglobin-related information and neoplastic lesion in C2 and C3 modes or enhancement of neoplastic structures in B7 and B8[10].
The visualization using a NBI was mostly used to detect the histological features in the research studies. NBI is an advanced imaging technology that uses digital optical methods to visualize more enhanced images than the standard white light[24]. NBI helps to examine the vascularity and abnormal histological features on site during colposcopy, thus adding AI to narrow band could improve the detection of the exact histology of polyps and saves time and effort waiting for histopathological assessment that may delay the intervention[25]. In addition, the NBI technique is easier than other more sophisticated techniques as chromoendoscopy[26].
Shin et al[27] used high resolution microendoscopy to detect esophageal cancer using AI integration, showing sensitivity of 93% and specificity of 92% in the training set and similar results, albeit slightly lower, in the test and independent sets.
Moreover, in other techniques like, capsule endoscopy, images taken couldn't be adjusted in position lightening or quality as they are dependent mainly on gut motility, plus their role in upper GI tract evaluation is still limited[28,29].
Online processing causes limitation on the acceptable latency requiring it to be low, so real time application mostly uses parallelization of the machine process. Current high-end GPU offer higher parallelization than current high-end CPUs, due to larger number of cores. An example for this issue appears when Nvidia Tensor RT, a software development kit SDK for highly parallel machine learning, marketed to reach up to 40 × performance speed than CPU only applications. Tensor RT runs only on CUDA (compute unified device architecture), which runs only on Nvidia graphic card. Furthermore, other libraries, as "Caffe", can be used either by CPU or GPU, through switching a flag in the source code[30,31].
Localized data sets and implementations, limited to specific institutions, will cause bias in methodological validation. Thus, public records of images and datasets are preferable to decrease this bias. On the other hand, implementation doesn't suffer the same urgency for public recording[32].
While latency in offline detection could reach days, this is not acceptable in online real time detection, as the latency during endoscopy procedures will cause missing of the lesions in vivo, but improvement is more beneficial, as the ideal scenario is no latency.
While some studies showed promising results in vitro, there is still work to do offline in order to get a real time implementation which can detect neoplasia during the endoscopy conduction in vivo[33]. However, of 36 included studies in a recent meta-analysis exploring the AI integration in all types of upper GI cancers[3], only three studies were in a clinical setting and one was RCT, but even the RCT was on images not real time, and the rest of studies were on stored images offline. Furthermore, very few studies included videos or live in vivo validation.
The first real time study for detection of gastric cancer was performed using an online AI system with Raman spectroscopy integrated to GI endoscopy in vivo. Total computation time ranged from 100-130 milliseconds for analysis, with diagnostic accuracy of 80%[34].
Ohmori et al[35] introduced a new AI system that could process 36 images per second, making it adequate for RT integration in upper GI endoscopy. One concern of the authors is that limiting their processing to high quality images could impair the RT usage at the time being.
A recent meta-analysis by Arribas et al[3], concluded that we need to focus more on real time AI systems in upper GI endoscopy, because due to small number of studies (only two were retrieved in this metaanalysis[36,37], we are still uncertain of the feasibility of integration of AI with the endoscopists in RT situations.
In Ebigbo et al[37], they used a live-stream camera, examining the classification and segmentation of 14 BE patients, with diagnostic accuracy of 89.9%. AI prediction takes 1.19 s with "ensembling" and 0.13 s without "ensembling".
Luo et al[36] performed the first aided AI RT implementation study in upper GI endoscopy. During a case-control study in six different hospitals in China, they developed a new AI system for RT examination named Gastrointestinal Artificial Intelligence Diagnosis System (GRAIDS), with latency of only 40 ms, and high diagnostic accuracy irrespective of the level of training of the endoscopists.
Imaging techniques, such as volumetric laser endomicroscopy, are used in BE to characterize different layers of the mucosa[38]. Characterization ideally includes the location, type and stage of neoplasia in the GI tract. A future prospect is the prognosis of this type.
"AI system is watching" is a statement that shows how endoscopists are more keen on clear videos and imaging when they know that an AI system will use those datasets[3,39].
The "blackbox" nature of CNN learning algorithms, means that we don’t know how the AI system reached its diagnosis, thus no human learning could be benefited from AI neoplasia recognition[40]. This is accompanied by the lack of training and lack of learning interest in postgraduates, mostly due to the cancer prevalence problems mentioned before. The story is different in colonoscopy, where in most studies, experts in the field usually beat the AI systems or show equal diagnostic efficacy, also where experts beat beginners or junior physicians[41-44].
Another solution presented by the AI implementation, is that only one system could be used in all types of upper GI endoscopy. In a multicenter study done by Luo et al[36], they used a new system called GRAIDS. This system allowed for the examination of all types of upper GI neoplasms including both esophageal and gastric in a single system. In addition, the system showed similar diagnostic accuracy when compared to experts[36].
AI implementation in upper GI endoscopy proceeds first from detection (the lesion is present or not), to segmentation (the lesion is differentiated from the surrounding normal tissue), and then to characterization (the lesion is histologically predicted). A quality assessment tool for diagnostic accuracy studies called QUDAS score and its modified version are used for quality assessment of these diagnostic accuracy trials[45].
One of the most promising findings was the early detection of precancerous lesions with chronic inflammatory background (chronic atrophic gastritis) with high specificity of all grades (mild, moderate and severe)[46]. This research might offer a solution to the hypothetical problem of background inflammatory state confusion with cancer. However, future validation is needed to reach our goal.
Accumulation of datasets, with the help of experts in annotating the pictures and videos of lesions in the upper GI endoscopy and linking them to the histopathological findings is mandatory for the progress of the AI in upper GI endoscopy. And public datasets will allow researchers to conduct their algorithm freely, without limitation to geographical regions or expert specialization in certain types of cancers.
Using a single system for detection of pan GI neoplasms with acceptable diagnostic accuracy for all GI regions is the ultimate goal, in addition to resolving the real time delay for image processing, which is still only scarcely examined in upper GI endoscopy.
Using AI integration with upper GI endoscopy could benefit trainees and general practitioners. Building a dataset library that is accessible to the researchers, with upper GI lesions apparent irrespective of the geographical area could be of great benefit to even experts in the fields with limited knowledge of the non-prevalent cancers in their area of practice.
Dr. Anup Kasi [Assistant Professor of Oncology in University of Kansas School of Medicine, Kansas City, KS (Medical Oncology), Unites States of America] edited the final version of the manuscript for English language flow and grammar.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: Egypt
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Wang G, Yasuda T, Yu H S-Editor: Liu M L-Editor: A P-Editor: Li JH
| 1. | Arnold M, Abnet CC, Neale RE, Vignat J, Giovannucci EL, McGlynn KA, Bray F. Global Burden of 5 Major Types of Gastrointestinal Cancer. Gastroenterology. 2020;159:335-349.e15. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 857] [Cited by in RCA: 1232] [Article Influence: 246.4] [Reference Citation Analysis (0)] |
| 2. | Yu H, Singh R, Shin SH, Ho KY. Artificial intelligence in upper GI endoscopy - current status, challenges and future promise. J Gastroenterol Hepatol. 2021;36:20-24. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 7] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 3. | Arribas J, Antonelli G, Frazzoni L, Fuccio L, Ebigbo A, van der Sommen F, Ghatwary N, Palm C, Coimbra M, Renna F, Bergman JJGHM, Sharma P, Messmann H, Hassan C, Dinis-Ribeiro MJ. Standalone performance of artificial intelligence for upper GI neoplasia: a meta-analysis. Gut. 2020 epub ahead of print. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 46] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 4. | Bang CS, Lee JJ, Baik GH. Artificial Intelligence for the Prediction of Helicobacter Pylori Infection in Endoscopic Images: Systematic Review and Meta-Analysis Of Diagnostic Test Accuracy. J Med Internet Res. 2020;22:e21983. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 64] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 5. | Lui TKL, Tsui VWM, Leung WK. Accuracy of artificial intelligence-assisted detection of upper GI lesions: a systematic review and meta-analysis. Gastrointest Endosc. 2020;92:821-830.e9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 77] [Cited by in RCA: 61] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 6. | Menon S, Trudgill N. How commonly is upper gastrointestinal cancer missed at endoscopy? Endosc Int Open. 2014;2:E46-E50. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 176] [Cited by in RCA: 230] [Article Influence: 20.9] [Reference Citation Analysis (0)] |
| 7. | Sekiguchi M, Oda I. High miss rate for gastric superficial cancers at endoscopy: what is necessary for gastric cancer screening and surveillance using endoscopy? Endosc Int Open. 2017;5:E727-E728. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 6] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 8. | Kitamura Y, Ito M, Matsuo T, Boda T, Oka S, Yoshihara M, Tanaka S, Chayama K. Characteristic epithelium with low-grade atypia appears on the surface of gastric cancer after successful Helicobacter pylori eradication therapy. Helicobacter. 2014;19:289-295. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 47] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 9. | Ito M, Tanaka S, Takata S, Oka S, Imagawa S, Ueda H, Egi Y, Kitadai Y, Yasui W, Yoshihara M, Haruma K, Chayama K. Morphological changes in human gastric tumours after eradication therapy of Helicobacter pylori in a short-term follow-up. Aliment Pharmacol Ther. 2005;21: 559-566. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 78] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 10. | Yasuda T, Yagi N, Omatsu T, Hayashi S, Nakahata Y, Yasuda Y, Obora A, Kojima T, Naito Y, Itoh Y. Benefits of linked color imaging for recognition of early differentiated-type gastric cancer: in comparison with indigo carmine contrast method and blue laser imaging. Surg Endosc. 2021;35:2750-2758. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 13] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 11. | Pawa R, Chuttani R. Benefits and limitations of simulation in endoscopic training. Tech Gastrointest Endosc. 2011;13:191-198.. [RCA] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 12. | Niu PH, Zhao LL, Wu HL, Zhao DB, Chen YT. Artificial intelligence in gastric cancer: Application and future perspectives. World J Gastroenterol. 2020;26:5408-5419. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 56] [Cited by in RCA: 69] [Article Influence: 13.8] [Reference Citation Analysis (1)] |
| 13. | Chang M. Artificial Intelligence for Drug Development, Precision Medicine, and Healthcare. 1st ed. Boca Raton: Chapman and Hall/CRC, 2020. [DOI] [Full Text] |
| 14. | Sánchez-Montes C, Bernal J, García-Rodríguez A, Córdova H, Fernández-Esparrach G. Review of computational methods for the detection and classification of polyps in colonoscopy imaging. Gastroenterol Hepatol. 2020;43:222-232. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 4] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 15. | Zhao Z, Yin Z, Wang S, Wang J, Bai B, Qiu Z, Zhao Q. Meta-analysis: The diagnostic efficacy of chromoendoscopy for early gastric cancer and premalignant gastric lesions. J Gastroenterol Hepatol. 2016;31:1539-1545. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 49] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 16. | Bang CS, Lim H, Jeong HM, Hwang SH. Use of Endoscopic Images in the Prediction of Submucosal Invasion of Gastric Neoplasms: Automated Deep Learning Model Development and Usability Study. J Med Internet Res. 2021;23:e25167. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 22] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 17. | Wu L, Zhou W, Wan X, Zhang J, Shen L, Hu S, Ding Q, Mu G, Yin A, Huang X, Liu J, Jiang X, Wang Z, Deng Y, Liu M, Lin R, Ling T, Li P, Wu Q, Jin P, Chen J, Yu H. A deep neural network improves endoscopic detection of early gastric cancer without blind spots. Endoscopy. 2019;51:522-531. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 194] [Cited by in RCA: 156] [Article Influence: 26.0] [Reference Citation Analysis (0)] |
| 18. | Kitagawa Y, Suzuki T, Nankinzan R, Ishigaki A, Furukawa K, Sugita O, Hara T, Yamaguchi T. Comparison of endoscopic visibility and miss rate for early gastric cancers after Helicobacter pylori eradication with white-light imaging versus linked color imaging. Dig Endosc. 2020;32: 769-777. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 26] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 19. | Fukuda H, Miura Y, Osawa H, Takezawa T, Ino Y, Okada M, Khurelbaatar T, Lefor AK, Yamamoto H. Linked color imaging can enhance recognition of early gastric cancer by high color contrast to surrounding gastric intestinal metaplasia. J Gastroenterol. 2019;54:396-406. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 56] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 20. | Yang YJ, Bang CS. Application of artificial intelligence in gastroenterology. World J Gastroenterol. 2019;25:1666-1683. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 211] [Cited by in RCA: 160] [Article Influence: 26.7] [Reference Citation Analysis (5)] |
| 21. | Willis BH. Spectrum bias--why clinicians need to be cautious when applying diagnostic test studies. Fam Pract. 2008;25:390-396. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 113] [Cited by in RCA: 105] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 22. |
van der Laan JJH, van der Waaij AM, Gabriëls RY, Festen EAM, Dijkstra G, Nagengast WB.
Endoscopic imaging in inflammatory bowel disease: current developments and emerging strategies |
| 23. | Shinozaki S, Osawa H, Hayashi Y, Lefor AK, Yamamoto H. Linked color imaging for the detection of early gastrointestinal neoplasms. Therap Adv Gastroenterol. 2019;12:1756284819885246. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 41] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 24. | Barbeiro S, Libânio D, Castro R, Dinis-Ribeiro M, Pimentel-Nunes P. Narrow-Band Imaging: Clinical Application in Gastrointestinal Endoscopy. GE Port J Gastroenterol. 2018;26:40-53. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 53] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 25. | Song EM, Park B, Ha CA, Hwang SW, Park SH, Yang DH, Ye BD, Myung SJ, Yang SK, Kim N, Byeon JS. Endoscopic diagnosis and treatment planning for colorectal polyps using a deep-learning model. Sci Rep. 2020;10:30. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 65] [Cited by in RCA: 64] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 26. | Mori Y, Neumann H, Misawa M, Kudo SE, Bretthauer M. Artificial intelligence in colonoscopy - Now on the market. What's next? J Gastroenterol Hepatol. 2021;36:7-11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 51] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 27. | Shin D, Protano MA, Polydorides AD, Dawsey SM, Pierce MC, Kim MK, Schwarz RA, Quang T, Parikh N, Bhutani MS, Zhang F, Wang G, Xue L, Wang X, Xu H, Anandasabapathy S, Richards-Kortum RR. Quantitative analysis of high-resolution microendoscopic images for diagnosis of esophageal squamous cell carcinoma. Clin Gastroenterol Hepatol. 2015;13:272-279.e2. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 66] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 28. | Saurin JC, Beneche N, Chambon C, Pioche M. Challenges and Future of Wireless Capsule Endoscopy. Clin Endosc. 2016;49:26-29. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 14] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 29. | Nadler M, Eliakim R. The role of capsule endoscopy in acute gastrointestinal bleeding. Therap Adv Gastroenterol. 2014;7:87-92. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 13] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 30. | Lequan Yu, Hao Chen, Qi Dou, Jing Qin, Pheng Ann Heng. Integrating Online and Offline Three-Dimensional Deep Learning for Automated Polyp Detection in Colonoscopy Videos. IEEE J Biomed Health Inform. 2017;21: 65-75. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 138] [Cited by in RCA: 106] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 31. | Akbari M, Mohrekesh M, Rafiei S, Reza Soroushmehr SM, Karimi N, Samavi S, Najarian K. Classification of Informative Frames in Colonoscopy Videos Using Convolutional Neural Networks with Binarized Weights. Annu Int Conf IEEE Eng Med Biol Soc. 2018;2018:65-68. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 11] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 32. | Chen PJ, Lin MC, Lai MJ, Lin JC, Lu HH, Tseng VS. Accurate Classification of Diminutive Colorectal Polyps Using Computer-Aided Analysis. Gastroenterology. 2018;154:568-575. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 250] [Cited by in RCA: 276] [Article Influence: 39.4] [Reference Citation Analysis (0)] |
| 33. | Mori Y, Kudo SE, Mohmed HEN, Misawa M, Ogata N, Itoh H, Oda M, Mori K. Artificial intelligence and upper gastrointestinal endoscopy: Current status and future perspective. Dig Endosc. 2019;31:378-388. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 87] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 34. | Duraipandian S, Sylvest Bergholt M, Zheng W, Yu Ho K, Teh M, Guan Yeoh K, Bok Yan So J, Shabbir A, Huang Z. Real-time Raman spectroscopy for in vivo, online gastric cancer diagnosis during clinical endoscopic examination. J Biomed Opt. 2012;17:081418. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 88] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 35. | Ohmori M, Ishihara R, Aoyama K, Nakagawa K, Iwagami H, Matsuura N, Shichijo S, Yamamoto K, Nagaike K, Nakahara M, Inoue T, Aoi K, Okada H, Tada T. Endoscopic detection and differentiation of esophageal lesions using a deep neural network. Gastrointest Endosc. 2020;91:301-309.e1. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 100] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 36. | Luo H, Xu G, Li C, He L, Luo L, Wang Z, Jing B, Deng Y, Jin Y, Li Y, Li B, Tan W, He C, Seeruttun SR, Wu Q, Huang J, Huang DW, Chen B, Lin SB, Chen QM, Yuan CM, Chen HX, Pu HY, Zhou F, He Y, Xu RH. Real-time artificial intelligence for detection of upper gastrointestinal cancer by endoscopy: a multicentre, case-control, diagnostic study. Lancet Oncol. 2019;20:1645-1654. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 155] [Cited by in RCA: 253] [Article Influence: 42.2] [Reference Citation Analysis (0)] |
| 37. | Ebigbo A, Mendel R, Probst A, Manzeneder J, Prinz F, de Souza LA Jr, Papa J, Palm C, Messmann H. Real-time use of artificial intelligence in the evaluation of cancer in Barrett's oesophagus. Gut. 2020;69:615-616. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 84] [Cited by in RCA: 124] [Article Influence: 24.8] [Reference Citation Analysis (0)] |
| 38. | Wolfsen HC. Volumetric Laser Endomicroscopy in Patients With Barrett Esophagus. Gastroenterol Hepatol (N Y). 2016;12: 719-722. [PubMed] |
| 39. | Bohr A, Memarzadeh K. The rise of artificial intelligence in healthcare applications. Artif Intell Healthcare. 2020;25-60. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 87] [Cited by in RCA: 377] [Article Influence: 75.4] [Reference Citation Analysis (1)] |
| 40. | Choi J, Shin K, Jung J, Bae HJ, Kim DH, Byeon JS, Kim N. Convolutional Neural Network Technology in Endoscopic Imaging: Artificial Intelligence for Endoscopy. Clin Endosc. 2020;53:117-126. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 35] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 41. | Hassan C, Spadaccini M, Iannone A, Maselli R, Jovani M, Chandrasekar VT, Antonelli G, Yu H, Areia M, Dinis-Ribeiro M, Bhandari P, Sharma P, Rex DK, Rösch T, Wallace M, Repici A. Performance of artificial intelligence in colonoscopy for adenoma and polyp detection: a systematic review and meta-analysis. Gastrointest Endosc. 2021;93:77-85.e6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 361] [Cited by in RCA: 308] [Article Influence: 77.0] [Reference Citation Analysis (1)] |
| 42. | Lui TKL, Guo CG, Leung WK. Accuracy of artificial intelligence on histology prediction and detection of colorectal polyps: a systematic review and meta-analysis. Gastrointest Endosc. 2020;92:11-22.e6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 80] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 43. | Barua I, Vinsard DG, Jodal HC, Løberg M, Kalager M, Holme Ø, Misawa M, Bretthauer M, Mori Y. Artificial intelligence for polyp detection during colonoscopy: a systematic review and meta-analysis. Endoscopy. 2021;53:277-284. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 158] [Article Influence: 39.5] [Reference Citation Analysis (1)] |
| 44. | Aziz M, Fatima R, Dong C, Lee-Smith W, Nawras A. The impact of deep convolutional neural network-based artificial intelligence on colonoscopy outcomes: A systematic review with meta-analysis. J Gastroenterol Hepatol. 2020;35:1676-1683. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 50] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 45. | Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, Leeflang MM, Sterne JA, Bossuyt PM; QUADAS-2 Group. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155:529-536. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6953] [Cited by in RCA: 9589] [Article Influence: 684.9] [Reference Citation Analysis (0)] |
| 46. | Zhang Y, Li F, Yuan F, Zhang K, Huo L, Dong Z, Lang Y, Zhang Y, Wang M, Gao Z, Qin Z, Shen L. Diagnosing chronic atrophic gastritis by gastroscopy using artificial intelligence. Dig Liver Dis. 2020;52:566-572. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 79] [Article Influence: 15.8] [Reference Citation Analysis (0)] |