Published online Feb 10, 2016. doi: 10.5317/wjog.v5.i1.39
Peer-review started: August 28, 2015
First decision: November 6, 2015
Revised: November 11, 2015
Accepted: December 8, 2015
Article in press: December 11, 2015
Published online: February 10, 2016
Processing time: 159 Days and 21.8 Hours
MUC16 (CA125) has remained the mainstay for ovarian cancer assessment and management since the early 1980’s. With the exception of HE4, it is the only reliable serum biomarker for ovarian cancer. MUC16 belongs to a family of high-molecular weight glycoproteins known as mucins. The mucin family is comprised of large secreted transmembrane proteins that includes MUC1, MUC4 and MUC16. These mucins are often overexpressed in a variety of malignancies. MUC1 and MUC4 have been shown to contribute to breast and pancreatic tumorigenesis. Recent studies have uncovered unique biological functions for MUC16 that go beyond its role as a biomarker for ovarian cancer. Here, we provide an overview of the literature to highlight the importance of MUC16 in ovarian cancer tumorigenesis. We focus on the growing literature describing the role of MUC16 in proliferation, migration, metastasis, tumorigenesis and drug resistance. Accumulating experimental evidence suggest that the C-terminal domain of MUC16 is critical to mediate theses effects. The importance of MUC16 in the pathogenesis of ovarian cancer emphasizes the need to fully understand the signaling capabilities of MUC16 C-terminal domain to develop more efficient strategies for the successful treatment of ovarian cancer.
Core tip: MUC16/CA125 has been a mainstay biomarker for ovarian cancer but its pathobiological role has remained mostly unknown. Recent literature has shown that MUC16 is much more than a biomarker. MUC16 has oncogenic properties and plays an important role in tumorigenesis. Here, we will review the current knowledge regarding the oncogenic role of MUC16.
- Citation: Piché A. Pathobiological role of MUC16 mucin (CA125) in ovarian cancer: Much more than a tumor biomarker. World J Obstet Gynecol 2016; 5(1): 39-49
- URL: https://www.wjgnet.com/2218-6220/full/v5/i1/39.htm
- DOI: https://dx.doi.org/10.5317/wjog.v5.i1.39
Cancer antigen 125 (CA125) is the most widely used and the best characterized biomarker for the management of epithelial ovarian cancer (EOC)[1-16]. CA125 was initially identified in 1981 as an antigen present in the serum of patients with EOC[17,18]. However, it was not until 2001 that the gene was cloned and identified as MUC16[19-22]. It was later realized that the CA125 antibody recognizes a repeating epitope located in the extracellular domain of MUC16[5,19,20]. For the purpose of this review, the term CA125 will be used to designate the cleaved extracellular domain that can be found in serum of EOC and the term MUC16 will designate the full length mucin glycoprotein. For CA125 to be detected in the serum of EOC patients, the MUC16 glycoprotein undergoes a proteolytic cleavage resulting in the release of most of the extracellular domain from the cell surface. Serum levels of CA125 are routinely used in clinic to monitor patients with EOC[2,7,9,10,12,13,15,18]. There is indeed a strong correlation between rising and falling levels of serum CA125 with the progression and regression of the disease. Although the MUC16 mucin may be expressed in other cancers, its use in clinic has been mostly limited to patients with EOC.
Despite the fact that serum CA125 has been the mainstay of EOC assessment and management since the early 1980’s, there are considerable gaps in our knowledge regarding the biological functions of MUC16 in normal physiology and particularly in oncogenesis. Not surprisingly, most of the early studies (before 2001) have focused on the clinical applicability of CA125 as a biomarker. However, the cloning of MUC16 gene and the characterization of its structure, which revealed that MUC16 belong to membrane-bound mucins, has encouraged studies aiming to elucidate its physiological normal function as well as its role in malignant conditions. Because other membrane-bound mucins, such as MUC1 and MUC4, have oncogenic properties, it has been hypothesized that MUC16 may possess oncogenic capabilities as well. However, the very large size of the molecule (MUC16 is the biggest mucin identified to date) has hampered the characterization of MUC16 functional domains leading to an incomplete understanding of the physiological and pathological roles of MUC16. Nonetheless, in the last few years, a number of studies have begun to unravel the biological functions of MUC16. In particular, these studies have demonstrated that MUC16 possesses oncogenic properties. Here, we will review the current knowledge regarding the oncologic role of MUC16 in ovarian cancer. Our discussion will focus on cancer cells signaling, invasion and metastasis, and on regulation of drug-induced apoptosis. This review will also highlight gaps in our knowledge to provide a framework for future research studies.
CA125 is the most widely studied serum biomarker for EOC. CA125 has been recognized as a tumor-associated antigen based on the characterization of the OC125 monoclonal antibody raised against the human ovarian cancer cell line OVCA433 in 1981[17,18]. This antibody detected a molecule that was named CA125 in the serum from patients with EOC[17]. It was later realized that CA125 is an epitope located on a repeated extracellular domain of MUC16 mucin[5,19,20]. Measurement of serum CA125 is an important part of the clinical management for EOC patients[2,10]. Its clinical utility has been evaluated as a screening test for the early detection of EOC, to distinguish benign diseases from malignant conditions, and to monitor EOC progression and regression following treatment.
The early detection of EOC remains a clinical challenge and minimal progress has been achieved to detect early diseases, which are often clinically asymptomatic, at a more curable stage. The use of serum CA125, as a single biomarker, for the early detection of EOC is tempered by several factors. First, MUC16 glycoprotein may be expressed in various malignant and non-malignant conditions. MUC16 is expressed at low levels in the normal airway epithelium and levels can increase in some chronic conditions such as cystic fibrosis[23-25] or vary with dexamethasone treatment[26]. MUC16 is expressed at the apical surface of the ocular and conjonctival epithelium where it is part of the glycocalyx protecting corneal cells from bacterial infections and dryness[27-30]. Immunohistochemistry of human tissues using the OC125 antibody detected MUC16 expression in fetal coelomic epithelia and its derivatives such as Müllerian duct, fallopian tube, endometrium, endocervix[8,14]. MUC16 is expressed by mesothelial cells of the peritoneum, the pleura and the pericardium[8,14]. Furthermore, CA125 serum levels can be elevated in various benign diseases including menstruation, first trimester pregnancy, endometriosis, adenomyosis, salpingitis, uterine fibroids, chronic renal failure or in inflammation of the pleura, peritoneum or pericardium[31-34]. However, MUC16 expression is not found in normal adult colon, rectum, cervix, small intestine, liver, pancreatic ducts, spleen, kidney, skin and ovaries[35]. Secondly, the expression of MUC16 in EOC tissues varies according to the histotype. MUC16 is expressed by 56% to 85% of serous, 65% of endometrioid, 40% of clear cell and 36% of undifferentiated adenocarcinomas of the ovary, but by only 12% of mucinous ovarian cancers[6,7,35]. Therefore, elevated levels of CA125 are most strongly associated with serous EOC subtypes and its expression is more limited in other subtypes. Consequently, the detection of serum CA125 is notably less sensitive for subtypes other than serous EOC. MUC16 is also detected in normal airways as well as in a small percentage of invasive breast carcinomas, lung carcinomas and pancreatic carcinomas[23,36-39]. Therefore, measurements of serum CA125 levels, as a single modality, have a limited utility for screening of EOC because of its lack of sensitivity and specificity.
The clinical utility of monitoring of serum CA125, as a screening test, has been evaluated by the ovarian component of The Prostate, Lung, Colorectal and Ovarian (PLCO) Cancer Screening Trial[40]. Results from this study showed that annual screening with CA125 and transvaginal ultrasound had no mortality benefit. Furthermore, the majority of the screen detected cancers presented in advanced stages. Another large ovarian cancer screening trial is currently ongoing, the UK Collaborative Trial of Ovarian Cancer Screening. The multimodal arm of this study utilizes the Risk of Ovarian Cancer Algorithm (ROCA) to predict the probability of EOC as a first-line screen[41]. The ROCA is based on the slope of serial serum MUC16 measurements. According to ROCA, patients are classified as low, intermediate and high risk of developing EOC. When the risk is intermediate or high, patients undergo a transvaginal ultrasound. Results of this trial are expected soon and it thus remains to be seen whether CA125 serum levels will be useful for screening of ovarian cancer in the general population.
Another potential application of CA125 serum monitoring, is its ability to distinguish benign gynecological conditions from EOC. However, as mentioned previously, the expression of MUC16 in a number of benign gynecological conditions limits the utilization of serum CA125 as a single biomarker. To overcome this limitation, a number of risk prediction scores have been developed to identify women that have high risk of EOC. Examples of risk prediction models include: (1) the Risk of Malignancy Index combines serum CA125 levels with ultrasound and menopausal status to identify women at high risk of having EOC[42]; (2) the Risk of Ovarian Malignancy Algorithm (ROMA), which uses menopausal status with CA125 and HE4 serum values[43-45]; and (3) OVA1, which uses CA125, β2-microglobulin, apolipoprotein A1, transthyretin and transferrin[46]. The use of ROMA added more accuracy for differentiating the benign and malignant conditions compared to CA125 alone. Other algorithms are being investigated and there is an intense effort to search for new potential biomarkers for EOC. Recently, the combination of serum CA125 with IL-6 ascites levels was evaluated for its ability to discriminate between benign conditions and EOC[47]. Despite encouraging data, further evaluation is needed to determine the clinical utility of these panels of markers.
The clinical utility of serum CA125 has been best validated for monitoring of therapy as well as detection of recurrence. Indeed, CA125 is the only biomarker currently recommended for the monitoring of therapy and for the detection of recurrent diseases. Elevated levels of serum CA125 are common in patients with advanced disease of serous histiotype (approximately 90%). It was shown by several groups that rising and falling levels of serum CA125 correlate with progression and regression of the disease and this formed the basis for monitoring CA125 serum levels for patient follow-up[14,15,18]. However, up to 20% of patients with advanced EOC have normal serum level of CA125. CA125 nadir serum values were associated with a significantly longer progression-free survival (PFS) and overall survival[9,13]. Pretreatment CA125 serum level is an independent predictor of PFS in patients with advanced EOC who received a standard chemotherapy regimen[16].
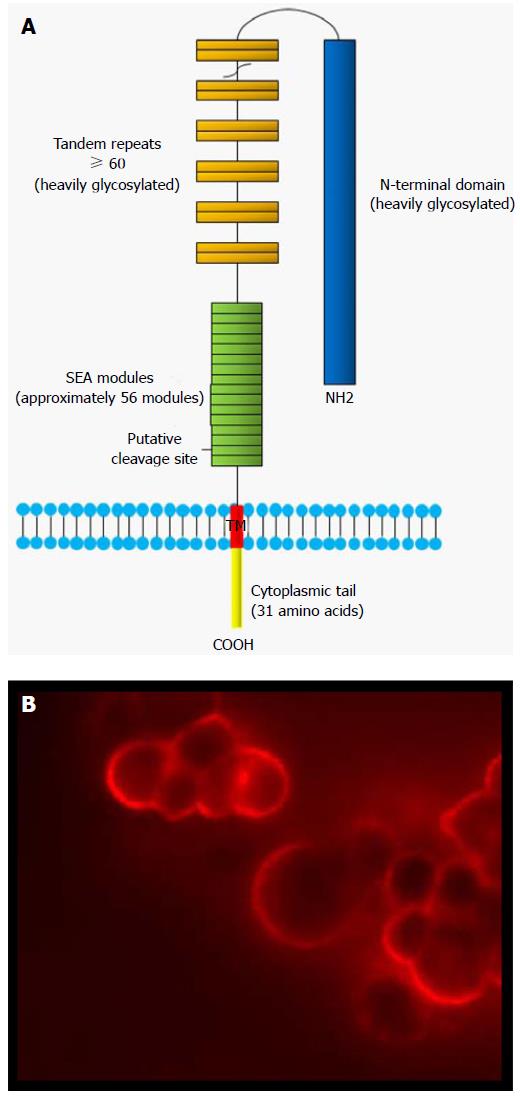
The cloning of the MUC16 gene in 2001 revealed that MUC16 is a tethered mucin[19-22]. The deduced amino acid sequence of MUC16 demonstrated that it shares common features with other membrane-bounded mucins with high serine, threonine and proline contents. MUC16 therefore belongs to the family of membrane-bound mucins. Other membrane-associated mucins include MUC1, MUC3A, MUC3B, MUC4, MUC12, MUC13, MUC15, MUC16, MUC17 and MUC20[48]. With a molecular weight of > 2 MDa, MUC16 is the largest membrane-bounded mucin known to date[21,22]. This glycoprotein is composed of a heavily glycosylated N-terminal domain, a large multiple repeat domain (up to 60 tandem repeats of 156 amino acids each), an ectodomain proximal to the putative cleavage site, a transmembrane domain (TM) and a small cytoplasmic domain of 31 amino acids[21] (Figure 1). MUC16 extracellular domain possesses much longer repeats (156 amino acids) as compared to MUC4 (16 amino acids) and MUC1 (20 amino acids). The N-terminal domain (12068 amino acids) and the repeat domain are heavily glycosylated with both O- and N-linked oligosaccharides[22]. In contrast to other membrane-bound mucins such as MUC1, which harbor a single sea urchin sperm protein, enterokinase and agrin (SEA) domain, MUC16 contains approximately 56 SEA domains (Figure 1)[49,50]. SEA domains consist of about 120 amino acids. Sequence analysis of MUC16 SEA modules showed that they display some sequence variability[50,51]. The second MUC16 SEA domain, proximal to the TM, however is relatively conserved and most closely resembles the SEA domain found in other mucins. Thus, MUC16 was speculated to contain a putative proteolytic cleavage site located in a SEA domain proximal to the membrane which could be involved in releasing the ectodomain[21]. Direct experimental validation has been lacking. However, a recent study has challenged this model. The investigators suggest that MUC16 cleavage occurs outside the proximal SEA domains and within the 12 extracellular amino acids proximal to the TM domain[52,53]. According to these authors, MUC16 cleavage takes place in the acidic pH of Golgi/post-Golgi compartments[54]. Notably, these authors did not found any evidence of intramembrane proteolysis with their constructs. The ectodomain of MUC16 has been reported to be released by metalloproteases [matrix metalloproteinase 7 (MMP-7), MMP-9, ZmpC] and neutrophil elastase[28,55]. ZmpC, a zinc metalloproteinase secreted by S. pneumoniae, selectively cleaves the ectodomain of MUC16 mucin from conjunctival and corneal epithelial cells[55]. The SEA domain is presumably the cleavage site for these proteases[28]. These findings contrast with that of Das et al[54] which showed that MUC16 cleavage was independent of neutrophil elastase and MMP-7. Post-translation modifications such as glycosylation can potentially regulate the release of MUC16 ectodomain and could explain these different findings. Intriguingly, mutation of Ser106Ala or Thr84/85Ala in the cytoplasmic tail did not affect the cleavage of MUC16[54]. According to these results, it would mean that there is no requirement for cytoplasmic cues for MUC16 extracellular cleavage. This is unexpected because previous studies showed that the phosphorylation of MUC16 cytoplasmic tail (Ser/Thr phosphorylation) has been associated with its secretion[56]. The secretion of MUC16 was also reported to be stimulated by epidermal growth factor (EGF) or tyrosine phosphatases[57]. Its shedding is decreased by glucocorticoids[26]. One limitation of the study by Das et al[54] is that, because of the lack of specific antibodies to MUC16 CT, they were not able to demonstrate cleavage of endogenous MUC16 in ovarian cancer cells to validate their observations, particularly in the context where most of the data were generated in non-ovarian cancer cell lines. Therefore, although MUC16 cleavage may transit in the Golgi before it reaches the cell surface, it is possible that its cleavage may also be triggered once the full length MUC16 is anchored in the cell membrane. The findings of Das et al[54], if confirmed, have important implications from a clinical standpoint. Since the cleavage of MUC16 appears to occur before it reaches the plasma membrane, antibodies raised against CA125 (extracellular portion of MUC16) may not be the most effective means to assess expression of MUC16 and could have been one of the reasons for the limited success in clinical studies of CA125 monoclonal antibodies[58-62].
Unlike MUC1 and MUC4, MUC16 lacks an EGF-like domain. Through their EGF-like motif located at C-terminal domain (extracellular portion), MUC1 and MUC4 bind to growth factor receptor tyrosine kinases (RTKs) such as erbB family and fibroblast growth factor receptor 3[63-66]. The formation of heterodimer with RTKs causes cross-phosphorylation of their respective cytoplasmic domain leading to the activation of various signaling pathways[67]. Because MUC16 lacks an RTK binding motif in its C-terminal domain, it is not clear whether MUC16-induced signaling is affected by RTKs although MUC16 extracellular domain shedding from the cell surface is stimulated by EGF. Consistent with the lack of an RTK binding motif, the intracellular interaction between MUC16 and β-catenin was not affected by EGF[68].
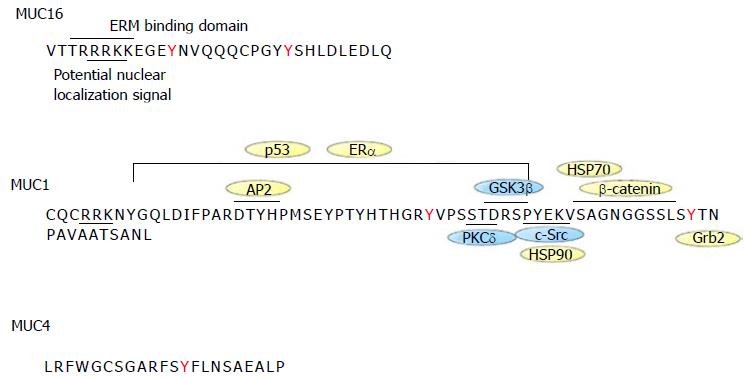
MUC16 31 amino acids long cytoplasmic tail (CT) contains serine/threonine/tyrosine residues that serve as potential phosphorylation sites. MUC16 CT also contains a polybasic sequence of amino acids (RRRKK), which is predicted to bind to the ezrin/radixin/moesin (ERM) family of proteins (Figure 2). This motif is not found in MUC1 and MUC4. The ERM proteins can interact with numerous membrane-associated proteins and the actin cytoskeleton. In line with this, MUC16 was recently shown to interact with E-cadherin and β-catenin, causing alterations in the actin cytoskeleton[68]. It remains unclear however whether MUC16/β-catenin and MUC16/E-cadherin interactions are mediated through the ERM motif of the MUC16 CT. Janus kinases (JAKs), which are non-receptor tyrosine kinases, contain an ERM domain. Previous studies showed that MUC16 co-immunoprecipitates with JAK2 in breast cancer cells resulting in the activation of STAT3, which may be involved in breast cancer development[69].
MUC1 cytoplasmic tail has been shown to bind to β-catenin and a serine-rich SXXXXXSSL motif in MUC1 CT is responsible for this interaction in vitro[70-72]. This motif is notably absent in MUC16. Interestingly however, the binding of MUC1 to β-catenin in cells was independent of the serine-rich motif[72]. These observations suggest that MUC16 interaction with β-catenin is mediated by an indirect mechanism, probably through another protein.
The positively charged R-K rich region of MUC16 CT also constitutes a putative nuclear localization motif[67]. Up until recently, evidence were lacking to support nuclear localization of MUC16 CT. However, a recent study demonstrated that MUC16 CT is cleaved from the cell surface and co-localized with JAK2 in the nucleus[52]. MUC16 CT-mediated increased nuclear JAK2 leads to up regulation of LMO2 and NANOG, two proteins shown to induce stem cell-like properties emphasizing the signaling capabilities of MUC16’s CT domain[52].
MUC16 is overexpressed by most serous EOC, by fallopian tube cancers and by cancers of the uterus[6,11,32,38]. Consistent with the role of MUC16 in tumorigenicity, overexpression of MUC16 mRNA and amplification of genomic regions encoding MUC16 DNA have been observed in The Cancer Genome Atlas (TCGA) ovarian cancer project that included 316 cases of serous EOC[73]. However, MUC16 expression was not correlated with overall survival or resistance. MUC16 is also aberrantly expressed by several other malignancies, including cancer of the lung, pancreas and breast. Along with MUC16, overexpression of other membrane-bound mucins, such as MUC1 and MUC4, is common in breast and pancreatic cancers[74]. Interestingly, both MUC1 and MUC4 mucins have been shown to possess oncogenic properties and promoted proliferation, invasion and metastasis in pancreatic cancer models[75-77]. These observations suggested that MUC16 could play a role in ovarian cancer progression as well as in other cancers.
Indeed, there are now several studies demonstrating that MUC16 C-terminal domain enhanced proliferation, migration, invasion and tumorigenicity of ovarian, breast and pancreatic cancer cell lines. The first study demonstrating the tumorigenic-enhancing properties of MUC16 was published by Thériault et al[78] in 2011. In this study, the knockdown of MUC16 in MUC16 overexpressing EOC cell line OVCAR3 decreased tumorigenicity as shown by the decreased number of colonies in soft agar and the reduction of in vivo tumor growth. In the same study, the enforced expression of MUC16 C-terminal domain (last 284 C-terminal residues) enhanced soft agar colony formation and tumor growth in nude mice. Consistently, intraperitoneal injection of MUC16 C-terminal domain-expressing tumor cells significantly reduced the survival of SCID mice compared to those injected with vector-expressing controls[78]. Subsequently, Lakshmanan et al[69] showed that MUC16 knockdown in overexpressing breast cancer cell lines decreased cell proliferation and in vivo tumor growth. Similarly, Reinartz et al[79] demonstrated that MUC16 knockdown in ovarian and breast cancer cells was associated with decreased migration and invasion. In line with these observations, more recent studies showed that ectopic expression of MUC16 C-terminal domain (last 114 C-terminal residues) enhanced proliferation, migration and in vivo tumor growth in pancreatic cancer cells whereas MUC16 knockdown reduced migration and invasion in these cancer cells[52,80]. Collectively, these studies clearly support a critical role for MUC16 in tumor growth and metastasis. Most importantly, expression of the C-terminal domain appears to be sufficient to mediate these effects.
The mechanism by which MUC16 enhances tumorigenicity of cancer cells remains poorly defined however. Although MUC16, as for MUC1 and MUC4, enhances tumorigenicity in cancer cells, the cytoplasmic tails of these three mucins are poorly conserved[50]. Size and amino acid sequence considerably vary suggesting a variety of functions in cell signaling. The ectopic expression of MUC16 C-terminal domain induces an epithelial to mesenchymal transition which has been associated with tumorigenesis and tumor progression[78]. Constitutive MUC16 C-terminal expression was associated with reduced E-cadherin expression which may enhance metastasis[81]. In addition, both Akt and ERK pathways may be activated by constitutive expression of MUC16 C-terminal domain[69,82]. The precise role these pathways was not however further investigated and therefore it is not clear whether their activation is essential to mediate MUC16 effects on tumor progression. However, Akt and ERK activation is frequently observed in serous OC and have been previously shown to be important for tumor progression[73,83,84].
Ectopic expression of MUC16 C-terminal domain in cancer cells has been associated with altered gene expression profile with increased expression of genes coding for proteins involved in invasion such as MMP2, MMP9, CXCL12 and CDH1[85]. MMPs are major proteolytic enzymes that are involved in cancer cell migration, invasion, and metastasis[86]. The CXCL12/CXCR7 axis plays an important role in ovarian cancer as it enhances cancer cell invasion by up-regulating MMP9 expression through the MAPK pathway[87]. MUC16 knockdown in breast cancer cells was associated with decreased expression of cyclin B1 and D1, both involved in regulation of cell cycle control[69]. The cytoplasmic domain of transmembrane mucins is usually embedded in the apical membrane of normal epithelial cells. However, upon transformation, the cytoplasmic tail of mucins such as MUC1 accumulates in the cytoplasm[88-90]. As mentioned above, there is evidence that MUC16 cytoplasmic tail gets cleaved and translocates in the nucleus[52]. However, whether this cleavage is required for MUC16 to enhance tumorigenicity remains to be determined.
Membrane-bound mucins such as MUC1 and MUC4 possess transforming properties[91,92]. For example, when the C-terminal portion of MUC1 was stably transfected into 3Y1 fibroblast cells, soft agar colonies and subcutaneous tumors in nude mice were readily obtained[91]. Interestingly, MUC16 is not expressed by normal ovarian surface epithelial (OSE) cell but its expression is induced in transformed OSE cells[93]. These observations suggest that MUC16 may play an important role at early stages in the development of EOC. Two recent studies indeed confirmed the transforming properties of MUC16. Giannakouros et al[82] stably expressed MUC16 C-terminal domain (last 284 C-terminal residues) and a similar construct lacking most of the cytoplasmic domain in immortalized NIH3T3 mouse fibroblast cells using lentiviral vectors. It was observed that MUC16 C-terminal domain-expressing cells displayed enhanced anchorage-dependent and -independent growth and enhanced the formation of sub-cutaneous tumor nodules in athymic nude mice[82]. Notably, these effects were abolished by the deletion of the cytoplasmic tail. In line with these observations, Rao et al[85], using constructs containing either the last 114 C-terminal amino acids or last 344 C-terminal amino acids, showed that both constructs significantly enhanced the number of soft agar colonies compared to vector only in NIH3T3 cells. In addition, both constructs formed larger sub-cutaneous tumors relative to control NIH3T3 cells with no significant differences suggesting that the proximal extracellular part of the molecule has little effect on the oncogenic effects of MUC16. These studies establish that MUC16 is an oncogenic glycoprotein much like MUC1 and MUC4.
There are, however, conflicting data regarding the minimal functional domain of the C-terminal domain required for transformation. Although, the SEA domain appears to be dispensable for tumorigenicity, the respective role of the ectodomain and the cytoplasmic tail are more controversial. Rao et al[85] showed that ectopic expression in NIH3T3 cells of a construct containing the ectodomain (58 amino acids), the TM (22 amino acids) and the first 6 amino acids of the CT had transforming effects mostly similar to a construct carrying the full length CT. In contrast, Thériault et al[78] and Giannakouros et al[82] showed that deletion of the CT in a construct consisting of the last 284 amino acids completely abolished the tumorigenic and transforming effects of MUC16 C-terminal domain. The reasons for this discrepancy are unclear but may reflect differences in integration of extracellular and intracellular signals with the different constructs that were used in these experiments.
Silencing MUC16 using single-chain antibodies increased the sensitivity of MUC16 overexpressing OVCAR3 cells to genotoxic agents such as cisplatin and doxorubicin but not to microtubule assembly inhibitors such paclitaxel and vinorelbin[94]. Conversely, expression of MUC16 C-terminal domain conferred increased resistance to cisplatin[94]. Consistent with the role of MUC16 in drug resistance, MUC1 has been shown to modulate the sensitivity of cancer cells to genotoxic agents. Indeed, expression of MUC1 C-terminal domain conferred resistance to cisplatin and etoposide in colon cancer cells[89,95]. The downregulation of MUC16 in OVCAR3 cells activates the PI3K/Akt pathway[68]. The authors also reported that MUC16 knockdown in these cells decreased FOXO3a nuclear localization. FOXO3a function is controlled in part by activation of the Akt pathway. Akt phosphorylates FOXO3a, resulting in binding of FOXO3a to 14-3-3 proteins and retention of FOXO3a in the cytoplasm. In contrast, dephosphorylation of FOXO3a induces its nuclear localization where it transactivates gene expression[96]. FOXO3a modulates the expression of several genes that regulate the cellular response to stress at the G2-M checkpoint. The growth arrest and DNA damage response gene Gadd54a is a target of FOXO3a that mediates part of FOXO3a’s effects on DNA repair[97]. Thus, preventing the nuclear localization of FOXO3a contributes to the apoptotic response to genotoxic drugs. It is therefore possible MUC16 knockdown sensitizes tumor cells to genotoxic drugs by activating Akt which in turn prevents FOXO3a nuclear localization. Although MUC16 expression may be associated with cisplatin resistance in vitro, no correlation was observed between MUC16 expression and resistance to chemotherapy in the TCGA ovarian cancer project[73]. Interestingly, however, a recent study suggested that the combination of serum CA125 and ascites leptin levels have a high discriminating potential to distinguish clinically resistant patients (intrinsic resistance) to first-line therapy from patients that responded to first-line chemotherapy[47].
Death receptor ligands such as tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) trigger rapid apoptosis in various tumor cell types[98-102]. TRAIL binds to death receptors, TRAIL-R1 (DR4) and -R2 (DR5), whose cytoplasmic death domain (DD) signals downstream caspase activation to mediate TRAIL-induced apoptosis[103]. Upon receptor activation, FADD and pro-caspase-8 are recruited to form a death-inducing signaling complex (DISC)[104]. When recruited to the DISC, pro-caspase-8 becomes activated and subsequently activates downstream effectors caspases-3, -6 and -7, leading to apoptosis. The cellular FLICE inhibitory protein (cFLIP) regulates both recruitment and processing of pro-caspase-8 within the DISC[105]. MUC16 ectopic expression, like MUC1, was shown to attenuate TRAIL-induced caspase-8 and mitochondria activation, resulting in decreased apoptosis[106]. Notably, MUC16 C-terminal expression was sufficient to attenuate TRAIL-induced apoptosis and signaling. MUC16-induced resistance to TRAIL was related to decreased TRAIL-R2 expression and recruitment at the DISC, and by increased cFLIP expression. MUC1-blockade of TRAIL-induced apoptosis has been attributed to localization of MUC1 C-terminal domain to the mitochondria or to the ability of MUC1 to directly bind caspase-8 and FADD, thereby inhibiting recruitment of caspase-8 at the DISC[107]. Thus, although both MUC1 and MUC16 can inhibit TRAIL-induced apoptosis, the mechanisms by which these mucins inhibit TRAIL signaling differ. This could be related to the amino acids composition of their cytoplasmic tail.
Since its discovery in the late 1970s, MUC16 glycoprotein has been recognized as a useful clinical biomarker in advance diseases. However, evidence now suggests that MUC16 is much more than a biomarker for disease progression. Recent studies have established that MUC16 can act as an oncogene. MUC16 expression leads to the transformation normal fibroblastic cells and it contributes to the pathogenesis and progression of EOC. These phenotypic effects are shared by other membrane-bounded mucins such as MUC1 and MUC4. However, the mechanisms by which these mucins exert their effects differ. The complex biochemical structure of MUC16 is a constant challenge to define functional domains. Although progress has been made toward understanding the role of MUC16 in tumorigenesis, many question remains. Further studies are required to gain a better understanding of the oncogenic role of MUC16. An area of critical importance is deciphering the mechanism by which MUC16 C-terminal domain enhances tumorigenesis and the downstream signaling associated with this effect. A better understanding of the contribution of MUC16 C-terminal domain to tumorigenesis may yield specific intracellular or extracellular signaling targets. Exploiting these targets could provide novel treatment for ovarian cancer which are urgently needed.
P- Reviewer: Sherbini MA, Wang PH, Yokoyama Y S- Editor: Wang JL L- Editor: A E- Editor: Wu HL
| 1. | Badgwell D, Bast RC. Early detection of ovarian cancer. Dis Markers. 2007;23:397-410. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 173] [Cited by in RCA: 187] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 2. | Duffy MJ, Bonfrer JM, Kulpa J, Rustin GJ, Soletormos G, Torre GC, Tuxen MK, Zwirner M. CA125 in ovarian cancer: European Group on Tumor Markers guidelines for clinical use. Int J Gynecol Cancer. 2005;15:679-691. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 198] [Cited by in RCA: 204] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 3. | Bast RC, Badgwell D, Lu Z, Marquez R, Rosen D, Liu J, Baggerly KA, Atkinson EN, Skates S, Zhang Z. New tumor markers: CA125 and beyond. Int J Gynecol Cancer. 2005;15 Suppl 3:274-281. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 283] [Cited by in RCA: 314] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 4. | Breitenecker G, Neunteufel W, Bieglmayer C, Kölbl H, Schieder K. Comparison between tissue and serum content of CA 125, CA 19-9, and carcinoembryonic antigen in ovarian tumors. Int J Gynecol Pathol. 1989;8:97-102. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 16] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 5. | Davis HM, Zurawski VR, Bast RC, Klug TL. Characterization of the CA 125 antigen associated with human epithelial ovarian carcinomas. Cancer Res. 1986;46:6143-6148. [PubMed] |
| 6. | de la Cuesta R, Maestro ML, Solana J, Vidart JA, Escudero M, Iglesias E, Valor R. Tissue quantification of CA 125 in epithelial ovarian cancer. Int J Biol Markers. 1999;14:106-114. [PubMed] |
| 7. | Høgdall EV, Christensen L, Kjaer SK, Blaakaer J, Kjaerbye-Thygesen A, Gayther S, Jacobs IJ, Høgdall CK. CA125 expression pattern, prognosis and correlation with serum CA125 in ovarian tumor patients. From The Danish “MALOVA” Ovarian Cancer Study. Gynecol Oncol. 2007;104:508-515. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 98] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 8. | Kabawat SE, Bast RC, Bhan AK, Welch WR, Knapp RC, Colvin RB. Tissue distribution of a coelomic-epithelium-related antigen recognized by the monoclonal antibody OC125. Int J Gynecol Pathol. 1983;2:275-285. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 413] [Cited by in RCA: 394] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 9. | Krivak TC, Tian C, Rose GS, Armstrong DK, Maxwell GL. A Gynecologic Oncology Group Study of serum CA-125 levels in patients with stage III optimally debulked ovarian cancer treated with intraperitoneal compared to intravenous chemotherapy: an analysis of patients enrolled in GOG 172. Gynecol Oncol. 2009;115:81-85. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 20] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 10. | Miller RE, Rustin GJ. How to follow-up patients with epithelial ovarian cancer. Curr Opin Oncol. 2010;22:498-502. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 14] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 11. | Nap M. Immunohistochemistry of CA 125. Unusual expression in normal tissues, distribution in the human fetus and questions around its application in diagnostic pathology. Int J Biol Markers. 1998;13:210-215. [PubMed] |
| 12. | Rustin GJ, Nelstrop AE, Tuxen MK, Lambert HE. Defining progression of ovarian carcinoma during follow-up according to CA 125: a North Thames Ovary Group Study. Ann Oncol. 1996;7:361-364. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 209] [Cited by in RCA: 213] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 13. | van Altena AM, Kolwijck E, Spanjer MJ, Hendriks JC, Massuger LF, de Hullu JA. CA125 nadir concentration is an independent predictor of tumor recurrence in patients with ovarian cancer: a population-based study. Gynecol Oncol. 2010;119:265-269. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 38] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 14. | Vasudev NS, Trigonis I, Cairns DA, Hall GD, Jackson DP, Broadhead T, Buxton J, Hutson R, Nugent D, Perren TJ. The prognostic and predictive value of CA-125 regression during neoadjuvant chemotherapy for advanced ovarian or primary peritoneal carcinoma. Arch Gynecol Obstet. 2011;284:221-227. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 24] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 15. | Vergote IB, Børmer OP, Abeler VM. Evaluation of serum CA 125 levels in the monitoring of ovarian cancer. Am J Obstet Gynecol. 1987;157:88-92. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 72] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 16. | Zorn KK, Tian C, McGuire WP, Hoskins WJ, Markman M, Muggia FM, Rose PG, Ozols RF, Spriggs D, Armstrong DK. The prognostic value of pretreatment CA 125 in patients with advanced ovarian carcinoma: a Gynecologic Oncology Group study. Cancer. 2009;115:1028-1035. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 76] [Cited by in RCA: 69] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 17. | Bast RC, Feeney M, Lazarus H, Nadler LM, Colvin RB, Knapp RC. Reactivity of a monoclonal antibody with human ovarian carcinoma. J Clin Invest. 1981;681:1331-1337. [RCA] [DOI] [Full Text] [Cited by in Crossref: 1132] [Cited by in RCA: 1129] [Article Influence: 25.7] [Reference Citation Analysis (0)] |
| 18. | Bast RC, Klug TL, St John E, Jenison E, Niloff JM, Lazarus H, Berkowitz RS, Leavitt T, Griffiths CT, Parker L. A radioimmunoassay using a monoclonal antibody to monitor the course of epithelial ovarian cancer. N Engl J Med. 1983;309:883-887. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1509] [Cited by in RCA: 1495] [Article Influence: 35.6] [Reference Citation Analysis (0)] |
| 19. | Yin BW, Lloyd KO. Molecular cloning of the CA125 ovarian cancer antigen: identification as a new mucin, MUC16. J Biol Chem. 2001;276:27371-27375. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 402] [Cited by in RCA: 447] [Article Influence: 18.6] [Reference Citation Analysis (0)] |
| 20. | Yin BW, Dnistrian A, Lloyd KO. Ovarian cancer antigen CA125 is encoded by the MUC16 mucin gene. Int J Cancer. 2002;98:737-740. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 167] [Cited by in RCA: 174] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 21. | O’Brien TJ, Beard JB, Underwood LJ, Dennis RA, Santin AD, York L. The CA 125 gene: an extracellular superstructure dominated by repeat sequences. Tumour Biol. 2001;22:348-366. [PubMed] |
| 22. | O’Brien TJ, Beard JB, Underwood LJ, Shigemasa K. The CA 125 gene: a newly discovered extension of the glycosylated N-terminal domain doubles the size of this extracellular superstructure. Tumour Biol. 2002;23:154-169. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 104] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 23. | Davies JR, Kirkham S, Svitacheva N, Thornton DJ, Carlstedt I. MUC16 is produced in tracheal surface epithelium and submucosal glands and is present in secretions from normal human airway and cultured bronchial epithelial cells. Int J Biochem Cell Biol. 2007;39:1943-1954. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 68] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 24. | Gronowitz E, Pitkänen S, Kjellmer I, Heikinheimo M, Strandvik B. Association between serum oncofetal antigens CA 19-9 and CA 125 and clinical status in patients with cystic fibrosis. Acta Paediatr. 2003;92:1267-1271. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 10] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 25. | Hattrup CL, Gendler SJ. Structure and function of the cell surface (tethered) mucins. Annu Rev Physiol. 2008;70:431-457. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 523] [Cited by in RCA: 580] [Article Influence: 34.1] [Reference Citation Analysis (0)] |
| 26. | Karlan BY, Amin W, Casper SE, Littlefield BA. Hormonal regulation of CA125 tumor marker expression in human ovarian carcinoma cells: inhibition by glucocorticoids. Cancer Res. 1988;48:3502-3506. [PubMed] |
| 27. | Argüeso P, Spurr-Michaud S, Russo CL, Tisdale A, Gipson IK. MUC16 mucin is expressed by the human ocular surface epithelia and carries the H185 carbohydrate epitope. Invest Ophthalmol Vis Sci. 2003;44:2487-2495. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 125] [Cited by in RCA: 150] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 28. | Blalock TD, Spurr-Michaud SJ, Tisdale AS, Gipson IK. Release of membrane-associated mucins from ocular surface epithelia. Invest Ophthalmol Vis Sci. 2008;49:1864-1871. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 90] [Cited by in RCA: 87] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 29. | Inatomi T, Spurr-Michaud S, Tisdale AS, Zhan Q, Feldman ST, Gipson IK. Expression of secretory mucin genes by human conjunctival epithelia. Invest Ophthalmol Vis Sci. 1996;37:1684-1692. [PubMed] |
| 30. | Jäger K, Wu G, Sel S, Garreis F, Bräuer L, Paulsen FP. MUC16 in the lacrimal apparatus. Histochem Cell Biol. 2007;127:433-438. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 26] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 31. | Ataseven H, Oztürk ZA, Arhan M, Yüksel O, Köklü S, Ibiş M, Başar O, Yilmaz FM, Yüksel I. Cancer antigen 125 levels in inflammatory bowel diseases. J Clin Lab Anal. 2009;23:244-248. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 11] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 32. | Miralles C, Orea M, España P, Provencio M, Sánchez A, Cantos B, Cubedo R, Carcereny E, Bonilla F, Gea T. Cancer antigen 125 associated with multiple benign and malignant pathologies. Ann Surg Oncol. 2003;10:150-154. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 100] [Cited by in RCA: 110] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 33. | Sikaris KA. CA125--a test with a change of heart. Heart Lung Circ. 2011;20:634-640. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 32] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 34. | Xiaofang Y, Yue Z, Xialian X, Zhibin Y. Serum tumour markers in patients with chronic kidney disease. Scand J Clin Lab Invest. 2007;67:661-667. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 28] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 35. | Dharma Rao T, Park KJ, Smith-Jones P, Iasonos A, Linkov I, Soslow RA, Spriggs DR. Novel monoclonal antibodies against the proximal (carboxy-terminal) portions of MUC16. Appl Immunohistochem Mol Morphol. 2010;18:462-472. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 42] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 36. | Ali M, Lillehoj EP, Park Y, Kyo Y, Kim KC. Analysis of the proteome of human airway epithelial secretions. Proteome Sci. 2011;9:4. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 46] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 37. | Kesimer M, Ehre C, Burns KA, Davis CW, Sheehan JK, Pickles RJ. Molecular organization of the mucins and glycocalyx underlying mucus transport over mucosal surfaces of the airways. Mucosal Immunol. 2013;6:379-392. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 151] [Cited by in RCA: 165] [Article Influence: 13.8] [Reference Citation Analysis (0)] |
| 38. | Moritani S, Ichihara S, Hasegawa M, Endo T, Oiwa M, Yoshikawa K, Sato Y, Aoyama H, Hayashi T, Kushima R. Serous papillary adenocarcinoma of the female genital organs and invasive micropapillary carcinoma of the breast. Are WT1, CA125, and GCDFP-15 useful in differential diagnosis? Hum Pathol. 2008;39:666-671. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 37] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 39. | Haridas D, Chakraborty S, Ponnusamy MP, Lakshmanan I, Rachagani S, Cruz E, Kumar S, Das S, Lele SM, Anderson JM. Pathobiological implications of MUC16 expression in pancreatic cancer. PLoS One. 2011;6:e26839. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 99] [Cited by in RCA: 108] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 40. | Buys SS, Partridge E, Black A, Johnson CC, Lamerato L, Isaacs C, Reding DJ, Greenlee RT, Yokochi LA, Kessel B. Effect of screening on ovarian cancer mortality: the Prostate, Lung, Colorectal and Ovarian (PLCO) Cancer Screening Randomized Controlled Trial. JAMA. 2011;305:2295-2303. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 870] [Cited by in RCA: 917] [Article Influence: 65.5] [Reference Citation Analysis (0)] |
| 41. | Menon U, Gentry-Maharaj A, Hallett R, Ryan A, Burnell M, Sharma A, Lewis S, Davies S, Philpott S, Lopes A. Sensitivity and specificity of multimodal and ultrasound screening for ovarian cancer, and stage distribution of detected cancers: results of the prevalence screen of the UK Collaborative Trial of Ovarian Cancer Screening (UKCTOCS). Lancet Oncol. 2009;10:327-340. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 640] [Cited by in RCA: 542] [Article Influence: 33.9] [Reference Citation Analysis (0)] |
| 42. | Jacobs I, Oram D, Fairbanks J, Turner J, Frost C, Grudzinskas JG. A risk of malignancy index incorporating CA 125, ultrasound and menopausal status for the accurate preoperative diagnosis of ovarian cancer. Br J Obstet Gynaecol. 1990;97:922-929. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 627] [Cited by in RCA: 593] [Article Influence: 16.9] [Reference Citation Analysis (0)] |
| 43. | Moore RG, Brown AK, Miller MC, Skates S, Allard WJ, Verch T, Steinhoff M, Messerlian G, DiSilvestro P, Granai CO. The use of multiple novel tumor biomarkers for the detection of ovarian carcinoma in patients with a pelvic mass. Gynecol Oncol. 2008;108:402-408. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 475] [Cited by in RCA: 477] [Article Influence: 28.1] [Reference Citation Analysis (0)] |
| 44. | Moore RG, McMeekin DS, Brown AK, DiSilvestro P, Miller MC, Allard WJ, Gajewski W, Kurman R, Bast RC, Skates SJ. A novel multiple marker bioassay utilizing HE4 and CA125 for the prediction of ovarian cancer in patients with a pelvic mass. Gynecol Oncol. 2009;112:40-46. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 538] [Cited by in RCA: 609] [Article Influence: 35.8] [Reference Citation Analysis (0)] |
| 45. | Moore RG, Miller MC, Disilvestro P, Landrum LM, Gajewski W, Ball JJ, Skates SJ. Evaluation of the diagnostic accuracy of the risk of ovarian malignancy algorithm in women with a pelvic mass. Obstet Gynecol. 2011;118:280-288. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 171] [Cited by in RCA: 198] [Article Influence: 14.1] [Reference Citation Analysis (0)] |
| 46. | Bast RC, Skates S, Lokshin A, Moore RG. Differential diagnosis of a pelvic mass: improved algorithms and novel biomarkers. Int J Gynecol Cancer. 2012;22 Suppl 1:S5-S8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 55] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 47. | Lane D, Matte I, Garde-Granger P, Laplante C, Carignan A, Rancourt C, Piché A. Inflammation-regulating factors in ascites as predictive biomarkers of drug resistance and progression-free survival in serous epithelial ovarian cancers. BMC Cancer. 2015;15:492. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 60] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 48. | Moniaux N, Escande F, Porchet N, Aubert JP, Batra SK. Structural organization and classification of the human mucin genes. Front Biosci. 2001;6:D1192-D1206. [PubMed] |
| 49. | Kui Wong N, Easton RL, Panico M, Sutton-Smith M, Morrison JC, Lattanzio FA, Morris HR, Clark GF, Dell A, Patankar MS. Characterization of the oligosaccharides associated with the human ovarian tumor marker CA125. J Biol Chem. 2003;278:28619-28634. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 178] [Cited by in RCA: 185] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 50. | Duraisamy S, Ramasamy S, Kharbanda S, Kufe D. Distinct evolution of the human carcinoma-associated transmembrane mucins, MUC1, MUC4 AND MUC16. Gene. 2006;373:28-34. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 85] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 51. | Jonckheere N, Skrypek N, Frénois F, Van Seuningen I. Membrane-bound mucin modular domains: from structure to function. Biochimie. 2013;95:1077-1086. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 59] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 52. | Das S, Rachagani S, Torres-Gonzalez MP, Lakshmanan I, Majhi PD, Smith LM, Wagner KU, Batra SK. Carboxyl-terminal domain of MUC16 imparts tumorigenic and metastatic functions through nuclear translocation of JAK2 to pancreatic cancer cells. Oncotarget. 2015;6:5772-5787. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 50] [Cited by in RCA: 69] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 53. | Berek JS. Immunotherapy of ovarian cancer with antibodies: a focus on oregovomab. Expert Opin Biol Ther. 2004;4:1159-1165. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 27] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 54. | Das S, Majhi PD, Al-Mugotir MH, Rachagani S, Sorgen P, Batra SK. Membrane proximal ectodomain cleavage of MUC16 occurs in the acidifying Golgi/post-Golgi compartments. Sci Rep. 2015;5:9759. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 41] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 55. | Govindarajan B, Menon BB, Spurr-Michaud S, Rastogi K, Gilmore MS, Argüeso P, Gipson IK. A metalloproteinase secreted by Streptococcus pneumoniae removes membrane mucin MUC16 from the epithelial glycocalyx barrier. PLoS One. 2012;7:e32418. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 73] [Cited by in RCA: 76] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 56. | Fendrick JL, Konishi I, Geary SM, Parmley TH, Quirk JG, O’Brien TJ. CA125 phosphorylation is associated with its secretion from the WISH human amnion cell line. Tumour Biol. 1997;18:278-289. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 30] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 57. | Konishi I, Fendrick JL, Parmley TH, Quirk JG, O’Brien TJ. Epidermal growth factor enhances secretion of the ovarian tumor-associated cancer antigen CA125 from the human amnion WISH cell line. J Soc Gynecol Investig. 1994;1:89-96. [PubMed] |
| 58. | Berek JS, Taylor PT, Gordon A, Cunningham MJ, Finkler N, Orr J, Rivkin S, Schultes BC, Whiteside TL, Nicodemus CF. Randomized, placebo-controlled study of oregovomab for consolidation of clinical remission in patients with advanced ovarian cancer. J Clin Oncol. 2004;22:3507-3516. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 143] [Cited by in RCA: 127] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 59. | Reinartz S, Köhler S, Schlebusch H, Krista K, Giffels P, Renke K, Huober J, Möbus V, Kreienberg R, DuBois A. Vaccination of patients with advanced ovarian carcinoma with the anti-idiotype ACA125: immunological response and survival (phase Ib/II). Clin Cancer Res. 2004;10:1580-1587. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 111] [Cited by in RCA: 97] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 60. | Reinartz S, Hombach A, Köhler S, Schlebusch H, Wallwiener D, Abken H, Wagner U. Interleukin-6 fused to an anti-idiotype antibody in a vaccine increases the specific humoral immune response against CA125+ (MUC-16) ovarian cancer. Cancer Res. 2003;63:3234-3240. [PubMed] |
| 61. | Sabbatini P, Dupont J, Aghajanian C, Derosa F, Poynor E, Anderson S, Hensley M, Livingston P, Iasonos A, Spriggs D. Phase I study of abagovomab in patients with epithelial ovarian, fallopian tube, or primary peritoneal cancer. Clin Cancer Res. 2006;12:5503-5510. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 52] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 62. | Sabbatini P, Harter P, Scambia G, Sehouli J, Meier W, Wimberger P, Baumann KH, Kurzeder C, Schmalfeldt B, Cibula D. Abagovomab as maintenance therapy in patients with epithelial ovarian cancer: a phase III trial of the AGO OVAR, COGI, GINECO, and GEICO--the MIMOSA study. J Clin Oncol. 2013;31:1554-1561. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 102] [Cited by in RCA: 110] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 63. | Li Y, Ren J, Yu W, Li Q, Kuwahara H, Yin L, Carraway KL, Kufe D. The epidermal growth factor receptor regulates interaction of the human DF3/MUC1 carcinoma antigen with c-Src and beta-catenin. J Biol Chem. 2001;276:35239-35242. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 192] [Cited by in RCA: 206] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 64. | Schroeder JA, Thompson MC, Gardner MM, Gendler SJ. Transgenic MUC1 interacts with epidermal growth factor receptor and correlates with mitogen-activated protein kinase activation in the mouse mammary gland. J Biol Chem. 2001;276:13057-13064. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 266] [Cited by in RCA: 279] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 65. | Pochampalli MR, el Bejjani RM, Schroeder JA. MUC1 is a novel regulator of ErbB1 receptor trafficking. Oncogene. 2007;26:1693-1701. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 87] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 66. | Ren J, Raina D, Chen W, Li G, Huang L, Kufe D. MUC1 oncoprotein functions in activation of fibroblast growth factor receptor signaling. Mol Cancer Res. 2006;4:873-883. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 66] [Cited by in RCA: 71] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 67. | Bafna S, Kaur S, Batra SK. Membrane-bound mucins: the mechanistic basis for alterations in the growth and survival of cancer cells. Oncogene. 2010;29:2893-2904. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 301] [Cited by in RCA: 290] [Article Influence: 19.3] [Reference Citation Analysis (0)] |
| 68. | Comamala M, Pinard M, Thériault C, Matte I, Albert A, Boivin M, Beaudin J, Piché A, Rancourt C. Downregulation of cell surface CA125/MUC16 induces epithelial-to-mesenchymal transition and restores EGFR signalling in NIH: OVCAR3 ovarian carcinoma cells. Br J Cancer. 2011;104:989-999. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 73] [Cited by in RCA: 88] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 69. | Lakshmanan I, Ponnusamy MP, Das S, Chakraborty S, Haridas D, Mukhopadhyay P, Lele SM, Batra SK. MUC16 induced rapid G2/M transition via interactions with JAK2 for increased proliferation and anti-apoptosis in breast cancer cells. Oncogene. 2012;31:805-817. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 121] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 70. | Yamamoto M, Bharti A, Li Y, Kufe D. Interaction of the DF3/MUC1 breast carcinoma-associated antigen and beta-catenin in cell adhesion. J Biol Chem. 1997;272:12492-12494. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 220] [Cited by in RCA: 239] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 71. | Wen Y, Caffrey TC, Wheelock MJ, Johnson KR, Hollingsworth MA. Nuclear association of the cytoplasmic tail of MUC1 and beta-catenin. J Biol Chem. 2003;278:38029-38039. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 138] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 72. | Huang L, Chen D, Liu D, Yin L, Kharbanda S, Kufe D. MUC1 oncoprotein blocks glycogen synthase kinase 3beta-mediated phosphorylation and degradation of beta-catenin. Cancer Res. 2005;65:10413-10422. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 169] [Cited by in RCA: 193] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 73. | Cancer Genome Atlas Research Network. Integrated genomic analyses of ovarian carcinoma. Nature. 2011;474:609-615. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6439] [Cited by in RCA: 6131] [Article Influence: 437.9] [Reference Citation Analysis (0)] |
| 74. | Haridas D, Ponnusamy MP, Chugh S, Lakshmanan I, Seshacharyulu P, Batra SK. MUC16: molecular analysis and its functional implications in benign and malignant conditions. FASEB J. 2014;28:4183-4199. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 85] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 75. | Kohlgraf KG, Gawron AJ, Higashi M, Meza JL, Burdick MD, Kitajima S, Kelly DL, Caffrey TC, Hollingsworth MA. Contribution of the MUC1 tandem repeat and cytoplasmic tail to invasive and metastatic properties of a pancreatic cancer cell line. Cancer Res. 2003;63:5011-5020. [PubMed] |
| 76. | Singh AP, Moniaux N, Chauhan SC, Meza JL, Batra SK. Inhibition of MUC4 expression suppresses pancreatic tumor cell growth and metastasis. Cancer Res. 2004;64:622-630. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 194] [Cited by in RCA: 200] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 77. | Chaturvedi P, Singh AP, Moniaux N, Senapati S, Chakraborty S, Meza JL, Batra SK. MUC4 mucin potentiates pancreatic tumor cell proliferation, survival, and invasive properties and interferes with its interaction to extracellular matrix proteins. Mol Cancer Res. 2007;5:309-320. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 115] [Cited by in RCA: 132] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 78. | Thériault C, Pinard M, Comamala M, Migneault M, Beaudin J, Matte I, Boivin M, Piché A, Rancourt C. MUC16 (CA125) regulates epithelial ovarian cancer cell growth, tumorigenesis and metastasis. Gynecol Oncol. 2011;121:434-443. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 137] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 79. | Reinartz S, Failer S, Schuell T, Wagner U. CA125 (MUC16) gene silencing suppresses growth properties of ovarian and breast cancer cells. Eur J Cancer. 2012;48:1558-1569. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 56] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 80. | Shukla SK, Gunda V, Abrego J, Haridas D, Mishra A, Souchek J, Chaika NV, Yu F, Sasson AR, Lazenby AJ. MUC16-mediated activation of mTOR and c-Myc reprograms pancreatic cancer metabolism. Oncotarget. 2015;6:19118-19131. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 55] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 81. | Giannakouros P, Comamala M, Matte I, Rancourt C, Piché A. MUC16 mucin (CA125) regulates the formation of multicellular aggregates by altering β-catenin signaling. Am J Cancer Res. 2015;5:219-230. [PubMed] |
| 82. | Giannakouros P, Matte I, Rancourt C, Piché A. Transformation of NIH3T3 mouse fibroblast cells by MUC16 mucin (CA125) is driven by its cytoplasmic tail. Int J Oncol. 2015;46:91-98. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 16] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 83. | Grille SJ, Bellacosa A, Upson J, Klein-Szanto AJ, van Roy F, Lee-Kwon W, Donowitz M, Tsichlis PN, Larue L. The protein kinase Akt induces epithelial mesenchymal transition and promotes enhanced motility and invasiveness of squamous cell carcinoma lines. Cancer Res. 2003;63:2172-2178. [PubMed] |
| 84. | Balmanno K, Cook SJ. Tumour cell survival signalling by the ERK1/2 pathway. Cell Death Differ. 2009;16:368-377. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 319] [Cited by in RCA: 396] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 85. | Rao TD, Tian H, Ma X, Yan X, Thapi S, Schultz N, Rosales N, Monette S, Wang A, Hyman DM. Expression of the Carboxy-Terminal Portion of MUC16/CA125 Induces Transformation and Tumor Invasion. PLoS One. 2015;10:e0126633. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 38] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 86. | Al-Alem L, Curry TE. Ovarian cancer: involvement of the matrix metalloproteinases. Reproduction. 2015;150:R55-R64. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 81] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 87. | Kulbe H, Chakravarty P, Leinster DA, Charles KA, Kwong J, Thompson RG, Coward JI, Schioppa T, Robinson SC, Gallagher WM. A dynamic inflammatory cytokine network in the human ovarian cancer microenvironment. Cancer Res. 2012;72:66-75. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 155] [Cited by in RCA: 177] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 88. | Carraway KL, Funes M, Workman HC, Sweeney C. Contribution of membrane mucins to tumor progression through modulation of cellular growth signaling pathways. Curr Top Dev Biol. 2007;78:1-22. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 65] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 89. | Ren J, Agata N, Chen D, Li Y, Yu WH, Huang L, Raina D, Chen W, Kharbanda S, Kufe D. Human MUC1 carcinoma-associated protein confers resistance to genotoxic anticancer agents. Cancer Cell. 2004;5:163-175. [PubMed] |
| 90. | Singh PK, Hollingsworth MA. Cell surface-associated mucins in signal transduction. Trends Cell Biol. 2006;16:467-476. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 298] [Cited by in RCA: 312] [Article Influence: 16.4] [Reference Citation Analysis (0)] |
| 91. | Li Y, Liu D, Chen D, Kharbanda S, Kufe D. Human DF3/MUC1 carcinoma-associated protein functions as an oncogene. Oncogene. 2003;22:6107-6110. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 154] [Cited by in RCA: 163] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 92. | Bafna S, Singh AP, Moniaux N, Eudy JD, Meza JL, Batra SK. MUC4, a multifunctional transmembrane glycoprotein, induces oncogenic transformation of NIH3T3 mouse fibroblast cells. Cancer Res. 2008;68:9231-9238. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 44] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 93. | Auersperg N, Wong AS, Choi KC, Kang SK, Leung PC. Ovarian surface epithelium: biology, endocrinology, and pathology. Endocr Rev. 2001;22:255-288. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 276] [Cited by in RCA: 227] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 94. | Boivin M, Lane D, Piché A, Rancourt C. CA125 (MUC16) tumor antigen selectively modulates the sensitivity of ovarian cancer cells to genotoxic drug-induced apoptosis. Gynecol Oncol. 2009;115:407-413. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 72] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 95. | Yin L, Huang L, Kufe D. MUC1 oncoprotein activates the FOXO3a transcription factor in a survival response to oxidative stress. J Biol Chem. 2004;279:45721-45727. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 86] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 96. | Nemoto S, Finkel T. Redox regulation of forkhead proteins through a p66shc-dependent signaling pathway. Science. 2002;295:2450-2452. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 682] [Cited by in RCA: 693] [Article Influence: 30.1] [Reference Citation Analysis (0)] |
| 97. | Tran H, Brunet A, Grenier JM, Datta SR, Fornace AJ, DiStefano PS, Chiang LW, Greenberg ME. DNA repair pathway stimulated by the forkhead transcription factor FOXO3a through the Gadd45 protein. Science. 2002;296:530-534. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 648] [Cited by in RCA: 655] [Article Influence: 28.5] [Reference Citation Analysis (0)] |
| 98. | Khaider NG, Lane D, Matte I, Rancourt C, Piché A. Targeted ovarian cancer treatment: the TRAILs of resistance. Am J Cancer Res. 2012;2:75-92. [PubMed] |
| 99. | Ashkenazi A, Dixit VM. Death receptors: signaling and modulation. Science. 1998;281:1305-1308. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4271] [Cited by in RCA: 4191] [Article Influence: 155.2] [Reference Citation Analysis (0)] |
| 100. | Ashkenazi A, Holland P, Eckhardt SG. Ligand-based targeting of apoptosis in cancer: the potential of recombinant human apoptosis ligand 2/Tumor necrosis factor-related apoptosis-inducing ligand (rhApo2L/TRAIL). J Clin Oncol. 2008;26:3621-3630. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 326] [Cited by in RCA: 331] [Article Influence: 19.5] [Reference Citation Analysis (0)] |
| 101. | Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M, Chin W, Jones J, Woodward A, Le T. Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat Med. 1999;5:157-163. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1924] [Cited by in RCA: 1927] [Article Influence: 74.1] [Reference Citation Analysis (0)] |
| 102. | Lane D, Cartier A, L’Espérance S, Côté M, Rancourt C, Piché A. Differential induction of apoptosis by tumor necrosis factor-related apoptosis-inducing ligand in human ovarian carcinoma cells. Gynecol Oncol. 2004;93:594-604. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 48] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 103. | Kischkel FC, Lawrence DA, Chuntharapai A, Schow P, Kim KJ, Ashkenazi A. Apo2L/TRAIL-dependent recruitment of endogenous FADD and caspase-8 to death receptors 4 and 5. Immunity. 2000;12:611-620. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 739] [Cited by in RCA: 753] [Article Influence: 30.1] [Reference Citation Analysis (0)] |
| 104. | Kischkel FC, Hellbardt S, Behrmann I, Germer M, Pawlita M, Krammer PH, Peter ME. Cytotoxicity-dependent APO-1 (Fas/CD95)-associated proteins form a death-inducing signaling complex (DISC) with the receptor. EMBO J. 1995;14:5579-5588. [PubMed] |
| 105. | Krueger A, Baumann S, Krammer PH, Kirchhoff S. FLICE-inhibitory proteins: regulators of death receptor-mediated apoptosis. Mol Cell Biol. 2001;21:8247-8254. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 403] [Cited by in RCA: 395] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 106. | Matte I, Lane D, Boivin M, Rancourt C, Piché A. MUC16 mucin (CA125) attenuates TRAIL-induced apoptosis by decreasing TRAIL receptor R2 expression and increasing c-FLIP expression. BMC Cancer. 2014;14:234. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 31] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 107. | Agata N, Ahmad R, Kawano T, Raina D, Kharbanda S, Kufe D. MUC1 oncoprotein blocks death receptor-mediated apoptosis by inhibiting recruitment of caspase-8. Cancer Res. 2008;68:6136-6144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 71] [Cited by in RCA: 72] [Article Influence: 4.2] [Reference Citation Analysis (0)] |