INTRODUCTION
The aqueous redox chemistry of the nucleosides and nucleotides has been extensively investigated for the last 40 years using pulse radiolysis, laser photolysis, electron spin resonance and other time resolved and steady state techniques[1]. More recently, theoretical methods have been employed in the study of redox damage of DNA[1]. This intense interest in the components of DNA is understandable since it carries our genetic code and if damaged can lead to mutations possibly resulting in cancer[2,3]. Furthermore, oxidative damage of DNA is implicated in aging[4] and bacterial drug resistance[5]. It is now understood that DNA damage initiated by ionising radiation elicits a complicated set of events engaging various signalling pathways in cells[6].
Given that cumulative cancer risk increases with the fourth power of age and is associated with an accumulation of DNA damage, oxidative DNA damage is of great interest regarding early tumorigenesis and eventually cancer. These redox damage mechanisms have a potential role in the initiation, promotion and malignant conversion stages of carcinogenesis[2]. Lesions such as 7,8-dihydro-8-oxoguanine (8OG) are established biomarkers of oxidative stress; coupled with their mutagenicity in mammalian cells, this has led to them being proposed as intermediate markers of cancer[2]. A more complete understanding of these oxidative damage processes in DNA is highly desirable in order to find new therapeutic strategies to battle this devastating disease.
BINDING SPECIFICITY ALTERATION OF THE GUANINE BASE
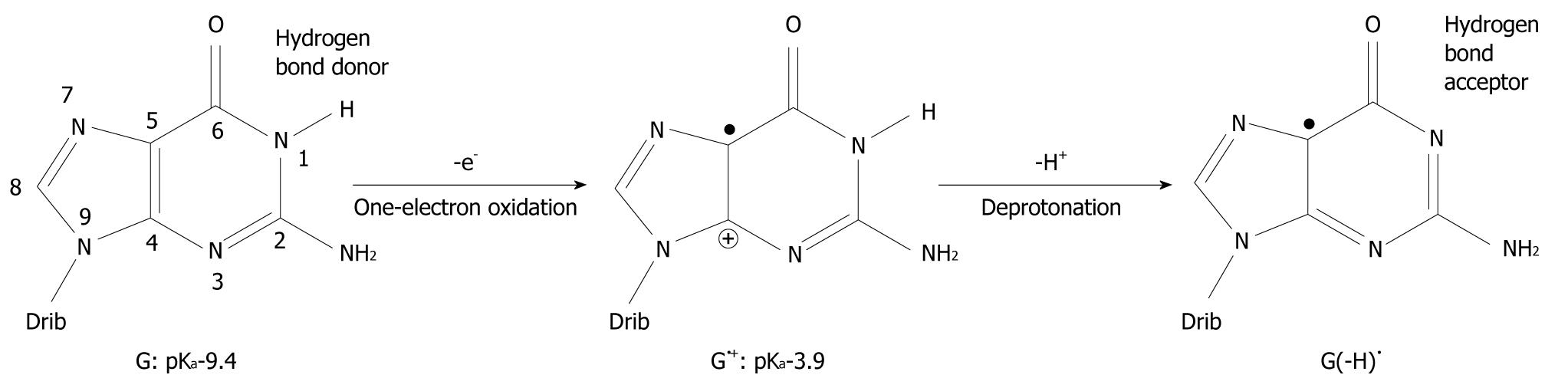
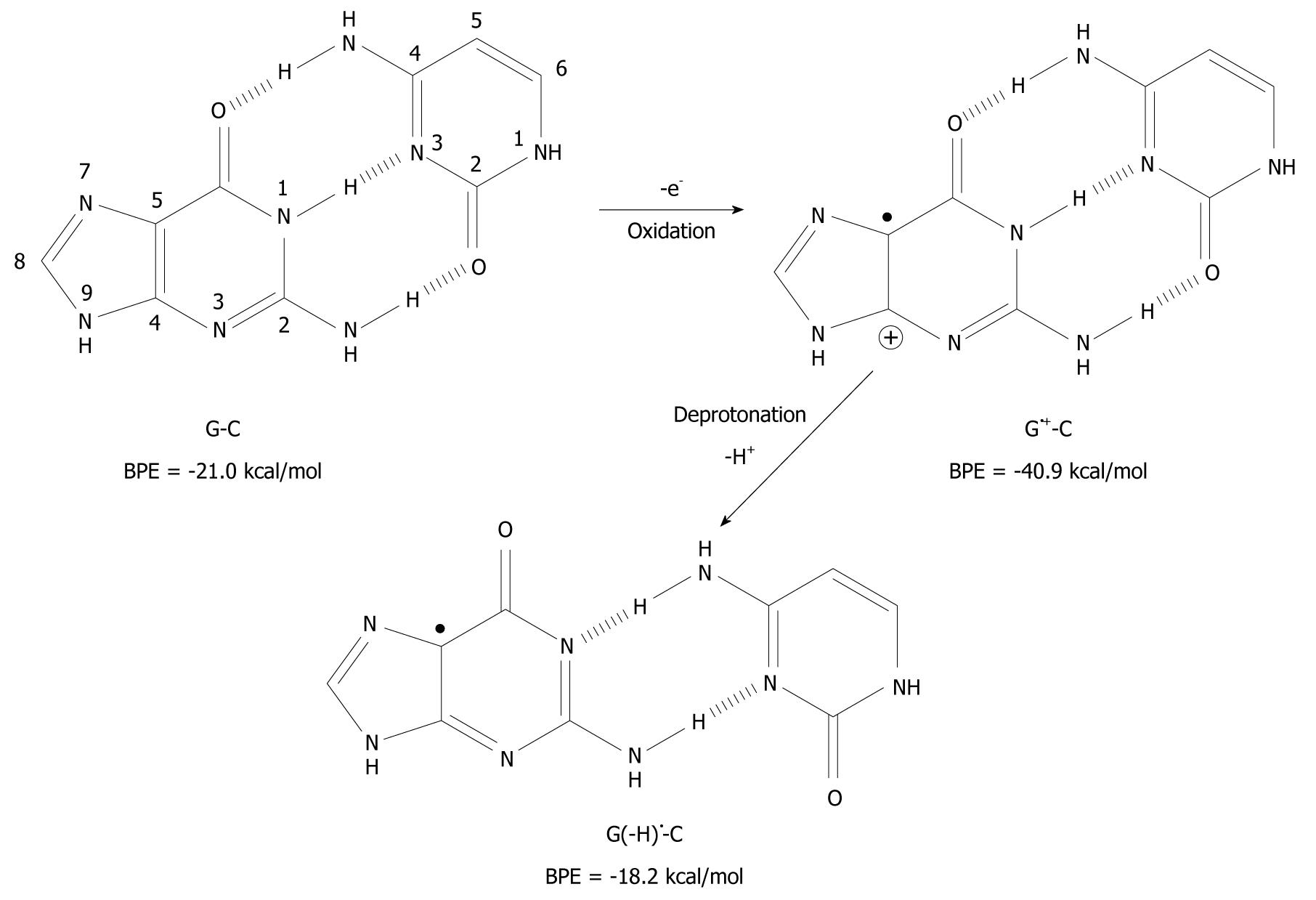
It has been found that when organic molecules are one-electron oxidized in the aqueous phase, a rapid deprotonation occurs from hydrogen bond donors undoubtedly driven by the massive solvation energy of the proton (∆Gaq = -263.9 kcal/mol)[7-9]. As an example, the pKa-value of cytosine (C) is lowered from 12.15 to between 2 and 4 when C is one-electron oxidized[8,10,11]. With respect to DNA, guanine (G) is its most easily oxidized component[12] and when the π-stack of double stranded DNA loses an electron, the positive charge migrates to G-C rich areas in the double strand[13-16] and the pKa-value of G is lowered significantly from 9.4 to 3.9 at the nitrogen-1 atom (N1), as depicted in Figure 1[10,17,18]. After departure of the proton from the N1-site, it becomes a hydrogen bond acceptor instead of a hydrogen bond donor. The question has emerged as to whether this event leads to a change in the pairing ability of the G moiety with other bases[19]? In fact, it is a common view that ligand hydrophobicity improves affinity, whereas hydrogen bonding improves specificity for interactions in biochemical systems[20]. Simulating one-electron oxidation and the consequent deprotonation of the central N1-proton for G-C, using the density functional theory (DFT)[21], a new slipped conformation of the base pair was formed as depicted in Figure 2[19]. This slipped configuration, G(-H)•-C, was later independently derived by Bera et al[22] using a systematic search for all possible hydrogen bonding configurations between G(-H)• and C. The predicted base pairing energy (BPE) was -18.2 kcal/mol for G(-H)•-C[19,23]. This lies between the BPE’s of the adenine-thymine base pair (A-T) at -13.0 kcal/mol and that of G-C at -21.0 kcal/mol[24,25].
Figure 1 Oxidation of guanine (G) and deprotonation of its radical cation (G•+).
The pKa-value of G is drastically lowered upon one-electron oxidation and subsequent deprotonation of the N1 proton changes it from a hydrogen bond donor to a hydrogen bond acceptor. The number of atoms constituting G is shown. Drib: 2’-deoxyribose moiety.
Figure 2 Deprotonation-induced structural change of the G-C base pair initiated by one-electron oxidation leading to the shifted base pair G(-H)• - C.
BPE: Base pairing energy.
DEPROTONATION OF OXIDIZED GUANINE IN DOUBLE STRANDED DNA
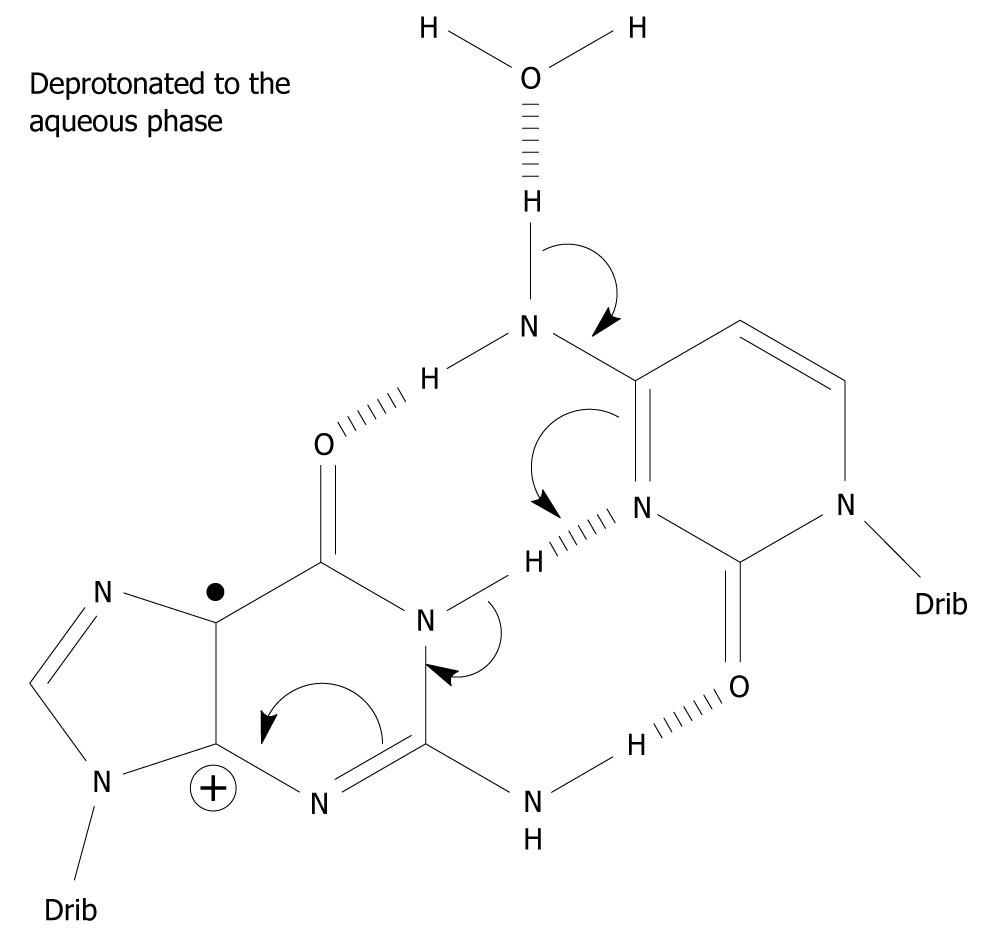
Under what circumstances can G•+-C in the DNA stack lose the central N1 proton making up one of the Watson-Crick hydrogen bonds? It does not have access to the aqueous phase since it is the central hydrogen bond and is flanked by base pairs on either side in the double stranded DNA helix. It is imperative that N1-H comes into contact with the water phase (water acting as a proton acceptor), i.e. within G-C, the G(N1-H)-C(N3) Watson-Crick hydrogen bond has to be broken for the N1 proton to be lost (Figure 2). The hydrogen bonds between the base pairs may be broken in three situations: First, the “swing-out” of the bases by concerted thermal motions of the DNA strand[26,27]. This mechanism is unlikely since it takes place on the milli- to micro-second time scale and is in competition with further charge migration in the DNA helix and/or with water addition to C8 of G•+, which are considerably faster. The rate of charge migration is estimated as 5 × 107/s and 6 × 104/s for the water addition, i.e. in the micro-nanosecond timescale[16,28]. Furthermore, the BPE of G•+-C is increased to -40.9 kcal/mol compared to -21.0 kcal/mol of its parent pair, inhibiting the frequency of the breathing motions of the base pair[24,29,30]. Second, when duplication of DNA occurs, the DNA strand is untwisted and the hydrogen bonds between the bases are broken to allow duplication of the strand. Third, during DNA transcription to messenger-RNA, it proceeds in a similar fashion to the duplication of DNA. In addition, it has been suggested that deprotonation occurs from the exocyclic amine group of C in G•+-C based on pulse-radiolysis and kinetic isotope experiments[31-33]. The proposed deprotonation mechanism is shown in Figure 3. This reaction cascade can lead to the G(-H)•-C slipped configuration[34].
Figure 3 A possible mechanism which involves the exocyclic amine moiety on C as the proton donor of the one-electron oxidized base pair in which the initial charge sits on G, i.
e. in the complementary strand. Spin-charge separation between G and C plays a crucial role in the reaction cascade. The depicted deprotonation can lead to the formation of G(-H)•-C[34].
PAIRING INFIDELITY OF THE DEPROTONATED GUANINE RADICAL
A related question has emerged as to whether it is possible to pair T, A and G itself to G(-H) •? This was investigated using the DFT method and the results are given in Figure 4[19].
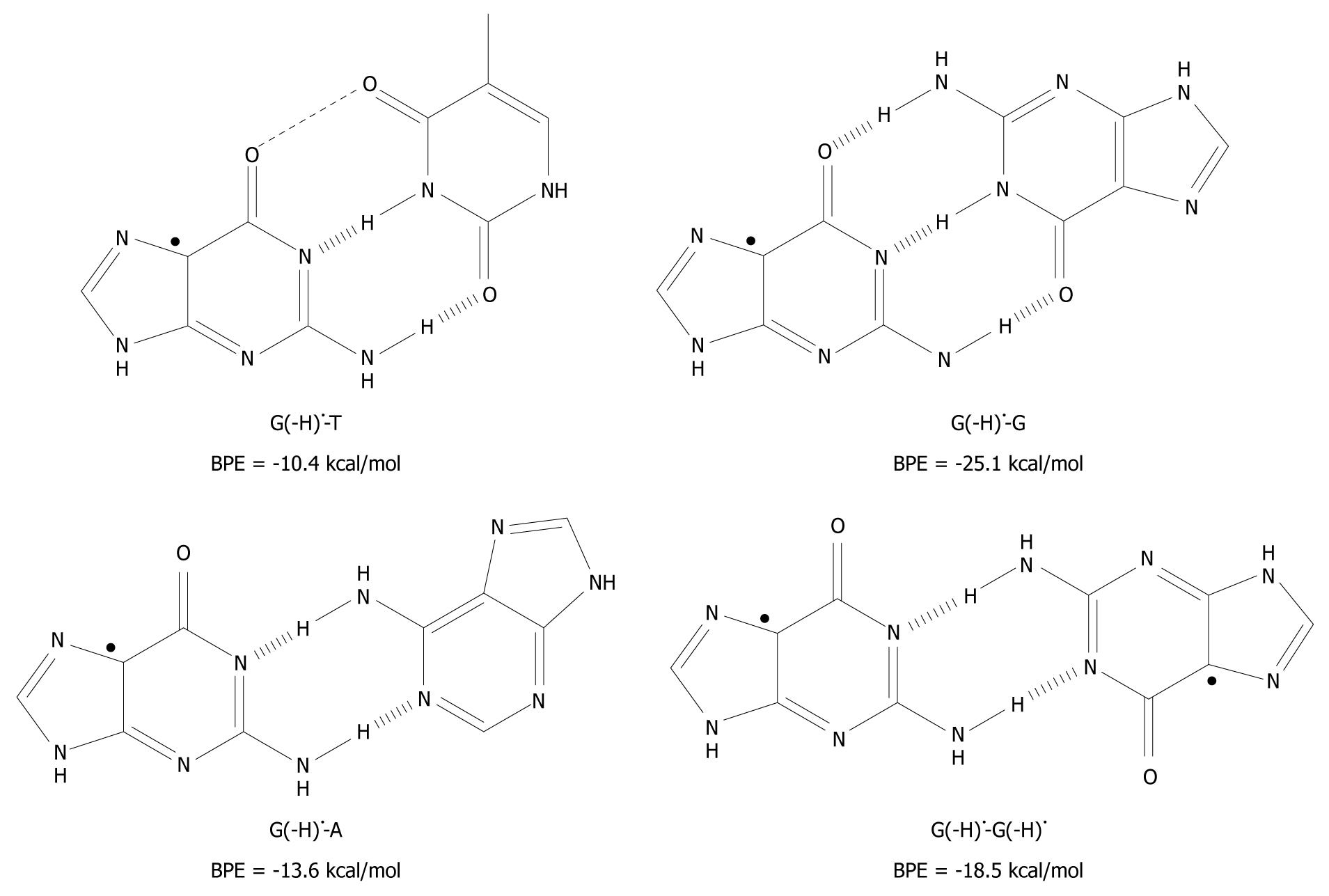
Figure 4 The unnatural base pairs between G(-H)• and the other bases[19].
The substantial base pairing energy (BPE) for the non-classical complexes depicted leads to the conclusion that G(-H)• does not have any specificity for C.
Armed with the knowledge that the G(-H)•-C base pair has only two hydrogen bonds, G(-H)• was paired to T and structurally optimized. The BPE was calculated to be -10.4 kcal/mol for G(-H)•-T, which is comparable to the A-T base pairing energy (-13.0 kcal/mol[24,25,29,35]). The relatively low energy can be explained in terms of the non-planarity of the bases with respect to each other. On the basis of the calculations, they are roughly 25º out of plane, measured at their carbonyl groups, O6 (G) and O4 (T). The distance between these oxygen atoms is 3.5 Å, which proximity leads to Coulombic repulsion and hence the non-planar conformation.
The calculated hydrogen bonding energy of the G(-H)•-A base pair is -13.6 kcal/mol, as shown in Figure 4. This binding is somewhat stronger than that for the natural A-T pairing (-13.0 kcal/mol)[24,25,29,35].
The hydrogen bond energy of G(-H)•-G (structure depicted in Figure 4) is similar to that of G-C[36]. This is not surprising as three hydrogen bonds are formed in both structures. A second type of G-G base pair is conceivable between two G(-H)• moieties (G(-H)•-G(-H)•) as shown in Figure 4. For this, the hydrogen bond energy is -18.5 kcal/mol, somewhat lower than for G(-H)•-G, since it has one less hydrogen bond. The Pt(II) electrophile coordinates at N7 of G. This acidifies the N1 proton, similar to the oxidation of G. With these Pt-G species, structures similar to G(-H)•-G and G(-H)•-G(-H)• were observed with 1H-NMR and X-ray crystallography[37], which provides experimental evidence of their existence.
ONE-ELECTRON OXIDATION DURING DNA DUPLICATION
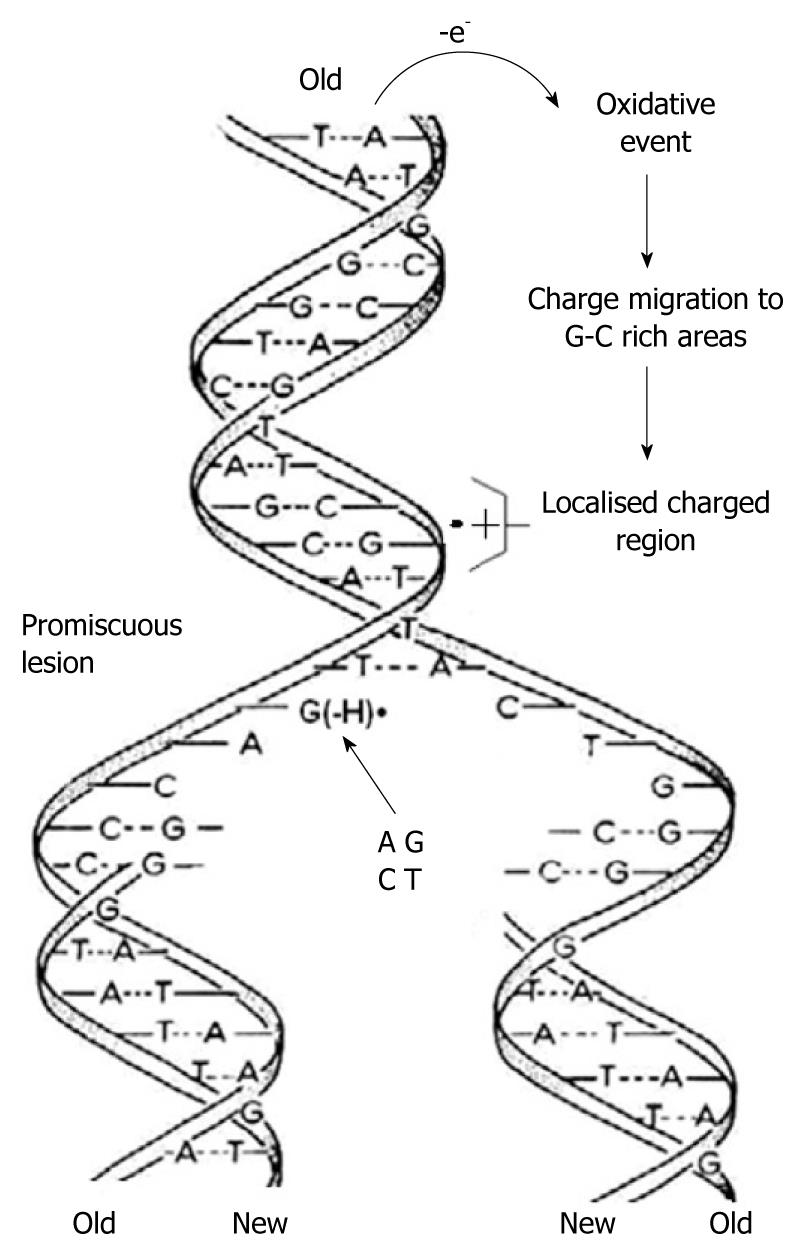
Using in-situ photolysis electron paramagnetic resonance (EPR), Hildenbrand and Schulte-Frohlinde, detected a long-lived radical (lifetime 5 s) which was produced only from double stranded DNA when ionised with < 220 nm light in an aqueous solution at pH 7[38]. This radical was assigned to G(-H)•. The rate of DNA duplication was measured to be between 5-500 nucleotides/s depending on the cell type, species and other factors[39,40]. Considering the long lifetime of G(-H)• in double stranded DNA and the rapid DNA duplication rate, it emerges that in the case of one-electron oxidation during mitosis-/-meiosis, G(-H)• is formed when the two strands unwind. As shown in Figure 4, it can form base pairs with all of the nucleotides with binding energies similar to the classical A-T and G-C Watson-Crick base pairs. This means that G(-H)• does not have specific affinity for C, i.e. it is completely promiscuous when it comes to base pairing. Therefore G(-H)• can pair with all of the nucleotides leading to mispairing. A depiction of this scenario is presented graphically in Figure 5.
Figure 5 As the two strands of the double helix unwind, each pairs up with the appropriate bases to form a new double helix.
The two new helices are identical to each other and to the original. This process is compromised by one-electron oxidation of the π-DNA stack, deprotonation from G•+ and the subsequent formation of G(-H)•, which is promiscuous with regard to base pairing.
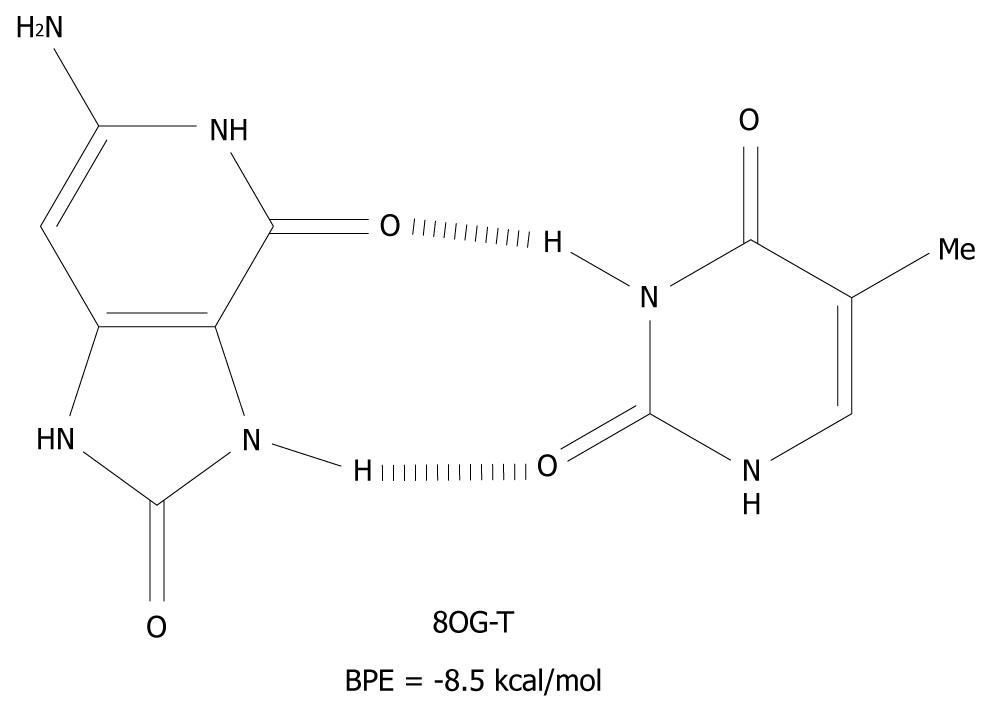
The mechanism presented here is new and an alternative to the scenario that mispairing of DNA bases is mostly caused by oxidative end products such as 8OG[41]. These products are closed shell, i.e. they are not radical species and therefore, have a much longer lifetime than G(-H)•. 8OG is one of the many redox products which is derived from the oxidation, and the subsequent water addition, of G[17,18,42]. It can form syn-anti base pairs[43], with all of the nucleotides and these have base pairing energies of -10 kcal/mol[44]. The 8OG-T base pair is depicted in Figure 6 as an example of syn-anti base pairs.
Figure 6 The syn-anti base pair of 8-oxoguanine-T.
BPE: Base pairing energy; 8OG: 8-oxoguanine.
So far, the role of DNA polymerase has not been considered and the DNA bases and base pairs have been treated as in vacuo as a model. The structure of DNA polymerase and its steric limitations within the active site are well documented[45-47]. The structure of the binding site in the replicating enzymes will undoubtedly have an effect on the proposed infidelity mechanism based on G(-H)•, e.g. the rate of duplication.
CONCLUSION
In this review, an alternative mechanism for promiscuous base pairing during DNA duplication, initiated by one-electron oxidation, is proposed based on theoretical calculations. Some experimental results exist which support the existence of the non-classical base pairs discussed, i.e. the slipped G(-H)•-C and the G(-H)•-G base pairs. Further experimental and theoretical work is needed to corroborate the mechanism proposed. In particular, experiments conducted with time resolved resonance Raman spectroscopy on model DNA duplication systems are pertinent as well as modelling studies on the effect of DNA polymerase.
Peer reviewer: Millie M Georgiadis, Associate Professor, Department of Biochemistry and Molecular Biology Indiana University School of Medicine, 635 Barnhill Dr. Indianapolis, IN 46202, United States
S- Editor Cheng JX L- Editor Webster JR E- Editor Ma WH