Published online Aug 14, 2017. doi: 10.3748/wjg.v23.i30.5508
Peer-review started: February 28, 2017
First decision: April 16, 2017
Revised: May 18, 2017
Accepted: July 4, 2017
Article in press: July 4, 2017
Published online: August 14, 2017
To investigate the microRNA expression profile in esophageal neosquamous epithelium from patients who had undergone ablation of Barrett’s esophagus.
High throughput screening using TaqMan® Array Human MicroRNA quantitative PCR was used to determine expression levels of 754 microRNAs in distal esophageal mucosa (1 cm above the gastro-esophageal junction) from 16 patients who had undergone ablation of non-dysplastic Barrett’s esophagus using argon plasma coagulation vs pretreatment mucosa, post-treatment proximal normal non-treated esophageal mucosa, and esophageal mucosal biopsies from 10 controls without Barrett’s esophagus. Biopsies of squamous mucosa were also taken from 5 cm above the pre-ablation squamo-columnar junction. Predicted mRNA target pathway analysis was used to investigate the functional involvement of differentially expressed microRNAs.
Forty-four microRNAs were differentially expressed between control squamous mucosa vs post-ablation neosquamous mucosa. Nineteen microRNAs were differentially expressed between post-ablation neosquamous and post-ablation squamous mucosa obtained from the more proximal non-treated esophageal segment. Twelve microRNAs were differentially expressed in both neosquamous vs matched proximal squamous mucosa and neosquamous vs squamous mucosa from healthy patients. Nine microRNAs (miR-424-5p, miR-127-3p, miR-98-5p, miR-187-3p, miR-495-3p, miR-34c-5p, miR-223-5p, miR-539-5p, miR-376a-3p, miR-409-3p) were expressed at higher levels in post-ablation neosquamous mucosa than in matched proximal squamous and healthy squamous mucosa. These microRNAs were also more highly expressed in Barrett’s esophagus mucosa than matched proximal squamous and squamous mucosa from controls. Target prediction and pathway analysis suggests that these microRNAs may be involved in the regulation of cell survival signalling pathways. Three microRNAs (miR-187-3p, miR-135b-5p and miR-31-5p) were expressed at higher levels in post-ablation neosquamous mucosa than in matched proximal squamous and healthy squamous mucosa. These miRNAs were expressed at similar levels in pre-ablation Barrett’s esophagus mucosa, matched proximal squamous and squamous mucosa from controls. Target prediction and pathway analysis suggests that these microRNAs may be involved in regulating the expression of proteins that contribute to barrier function.
Neosquamous mucosa arising after ablation of Barrett’s esophagus expresses microRNAs that may contribute to decreased barrier function and microRNAs that may be involved in the regulation of survival signaling pathways.
Core tip: We report that the microRNA profile of esophageal neosquamous mucosa developing after ablation of Barrett’s esophagus is different to normal squamous epithelium, and that the differentially expressed microRNAs in neosquamous mucosa may regulate survival signalling pathways and contribute to decreased barrier function in the esophagus.
- Citation: Sreedharan L, Mayne GC, Watson DI, Bright T, Lord RV, Ansar A, Wang T, Kist J, Astill DS, Hussey DJ. MicroRNA profile in neosquamous esophageal mucosa following ablation of Barrett’s esophagus. World J Gastroenterol 2017; 23(30): 5508-5518
- URL: https://www.wjgnet.com/1007-9327/full/v23/i30/5508.htm
- DOI: https://dx.doi.org/10.3748/wjg.v23.i30.5508
The incidence of esophageal adenocarcinoma has increased rapidly in the western world over recent decades, with overall 5-year survival rates of approximately 15%[1]. A strategy to improve survival outcome is early detection of cancer, or detection at the pre-malignant stage - high grade dysplasia. Barrett’s esophagus is the precursor to adenocarcinoma[2], and results from a metaplastic change of normal esophageal squamous epithelium to columnar epithelium with intestinal differentiation[3,4]. This is a consequence of chronic gastro-esophageal reflux, and it can be identified in 1%-2% of individuals aged over 60[4]. Barrett’s esophagus progresses to cancer in a sequential manner through low and then high grade dysplasia[2]. The risk of progression of non-dysplastic Barrett’s esophagus to adenocarcinoma has been reported to be 0.2%-0.5% per patient year for patients enrolled in surveillance programs[1].
Endoscopic surveillance remains the mainstay of cancer prevention in individuals with Barrett’s esophagus, and definitive management by surgery or endoscopy is reserved for individuals who develop high grade dysplasia or cancer[5]. Several endoscopic treatments are widely used for the treatment of high grade dysplasia or early cancer in Barrett’s esophagus, including radiofrequency ablation, argon plasma coagulation, and endoscopic mucosal resection.
Generally, endoscopic therapy for Barrett’s esophagus aims to completely eradicate any columnar mucosa, although persistent genomic alterations at tumour suppressor loci have been found after ablation[6]. Even though endoscopic removal of Barrett’s esophagus by ablative therapies is possible in the majority of patients[7], there is still a risk of recurrence and progression to adenocarcinoma following complete eradication of Barrett’s esophagus. As a result endoscopically treated patients are maintained under surveillance[4].
There have also been concerns about the risk of recurrence associated with residual sub-squamous glandular tissue which is not visible with a white light endoscope. However, Basu et al[8] reported that sub-squamous glandular tissue was not associated with recurrence of Barrett’s esophagus in patients with effective acid suppression after argon plasma coagulation ablation, and newer ablation modalities such as radio frequency ablation appear to have largely eliminated this problem[9], due to increased control of depth and uniformity of tissue ablation[10]. There have been several reports of patients progressing to cancer in whom Barrett’s esophagus has been completely eradicated[11-13], and this has brought into question the “normality” of the regenerated neosquamous epithelium. In a previous study we observed that the expression levels of cytokeratins CK-8 and CK-14, and microRNA-205, are similar in post-ablation neosquamous epithelium and more proximal normal squamous epithelium from patients with Barrett’s esophagus. However, microRNA-143 expression, which is elevated in Barrett’s esophagus[14], was elevated in post-ablation neosquamous mucosa and in squamous mucosa above the metaplastic segment compared to squamous epithelium from healthy patients, suggesting that the regenerated neosquamous mucosa might not be “normal”[15].
MicroRNAs are short (about 22 nucleotides) non-coding RNA molecules that regulate gene expression. Because a single microRNA can target several mRNAs, dysregulated microRNA expression can impact on key biological pathways and contribute to cancer development[16]. Dysregulated microRNA expression along the squamous - Barrett’s - dysplasia - adenocarcinoma pathway has been reported by several groups[3,14,17-19]. In the current study we profiled global microRNA expression in esophageal mucosa before and after ablation of Barrett’s esophagus using Argon plasma coagulation. Our aim was to further investigate differences in microRNA expression between neosquamous and normal squamous mucosa, and to investigate how these differences might contribute to altered biology using predicted targets and pathway analysis.
The methods used for the tissue collection and processing are described in our previous study[15], and reproduced here for completion.
Esophageal mucosal tissue was collected before and after Argon plasma coagulation ablation of Barrett’s esophagus from 16 individuals of median age 54.2 years (range 28.9-68.1) who were enrolled in the treatment arms of previously reported randomised controlled trials of Barrett’s esophagus ablation vs endoscopic surveillance[20]. Barrett’s esophagus was defined as columnar epithelium in the distal esophagus with histological confirmation of the presence of intestinal metaplasia. All patients were free of reflux symptoms following treatment of gastro-esophageal reflux, by either high dose proton pump inhibitors (n = 8) or a laparoscopic fundoplication (n = 8); before enrolment in the trial, at pre and post-treatment sample collection, and at Barrett’s esophagus ablation.
All patients underwent baseline endoscopy and biopsies from the distal esophageal mucosa were collected as described below. Biopsies were assessed by standard histopathological techniques. The presence of intestinal metaplasia and the absence of dysplasia within the Barrett’s esophagus segment were confirmed in all patients. The pre-ablation length of Barrett’s esophagus ranged from 1-10 cm in length (median 3 cm). Patients underwent endoscopic Argon plasma coagulation ablation following baseline endoscopy. The details of the ablation protocol have been described in detail previously[20]. Of the patients contributing tissues to the current study, complete ablation of the Barrett’s esophagus was achieved in 13 of 16 (82%). In the other 3, 95%, 99% and 95% ablation was achieved. Patient-matched post ablation neosquamous and proximal squamous samples were included in the study.
Four quadrant esophageal biopsies were taken commencing from 1 cm above the gastro-esophageal junction and then every 2 cm proximally for the length of the Barrett’s segment and sent for histopathology. An additional three biopsies were collected for research purposes from each sampled level of the Barrett’s esophagus, and stored in RNAlater® (Ambion, Austin, Texas, United States) as per the manufacturer’s protocol.
Repeat endoscopy was performed at a median of 6 wk (inter quartile range 4.96-6.5 wk) after the last ablation treatment and biopsies were collected from the post-ablation neosquamous esophageal mucosa using the same biopsy collection protocol. Additional biopsies were collected 5 cm above the proximal margin of the pre-ablation Barrett’s mucosa, and the corresponding site post-ablation for use as patient-matched non-regenerated squamous esophageal mucosa.
Biopsies collected from patients who had undergone Barrett’s esophagus ablation were selected for analysis from the following sites: (1) pre-ablation Barrett’s esophagus mucosa (columnar mucosa with intestinal metaplasia), 1 cm above the gastro-esophageal junction; (2) post-ablation neosquamous mucosa, 1 cm above the gastro-esophageal junction; and (3) post-ablation squamous mucosa, 5 cm above the level of the pre-ablation squamo-columnar junction.
Endoscopic esophageal mucosal biopsies were also collected at endoscopy from fourteen control individuals of median age 51.9 (range 24.1-71.0) with no known esophageal disease. These biopsies were taken from the distal esophagus, at an equivalent distance from the gastro-esophageal junction to the neosquamous mucosal biopsies, to allow direct comparison of the biopsies from normal squamous mucosa from the control patients to the biopsies of neosquamous mucosa from the patients who underwent ablation. The inclusion criteria for the control patients were: (1) no reflux symptoms; (2) endoscopy was not undertaken for the investigation of reflux; (3) no macroscopic esophagitis seen at endoscopy; (4) gastro-esophageal junction closed when viewed from within the esophagus; and (5) gastro-esophageal junction snug around the endoscope when viewed with the retroflexed endoscope.
All biopsy samples for the study were immediately stored in RNAlater® (Ambion, Austin, Texas, United States) as per the manufacturer’s protocol at -20 °C until required. When required, the samples were thawed and RNAlater removed. Twenty-five precent of each tissue sample was fixed in formalin and embedded in paraffin for histopathology to confirm that the sample contained the required epithelium. This protocol has been described in detail previously[21]. There were no buried sub-squamous columnar glands detected by histopathology in any of the biopsies of neosquamous mucosa used in this study. The remaining tissue was used for gene expression analysis. RNA was extracted using Trizol (Invitrogen, Carlsbad, CA, United States), and RNA concentration was determined using a Biophotometer (Eppendorf AG, Hamburg, Germany).
TaqMan® Array Human MicroRNA Card Set (A and B) v3.0 was used to profile the expression of 754 microRNAs[22]. The extracted RNA was reverse transcribed using Megaplex™ RT Primers Pool A and B. Each reverse transcription reaction had a final volume of 7.5 μL, and contained 45 ng of total RNA in 3 μL, and 4.5 μL of RT reaction mix containing reverse transcriptase, Megaplex™ RT Primers Pool A and B, and other reverse transcription agents. The recommended RT thermal cycling conditions were used: (16 °C, 2 min; 42 °C, 1 min; 50 °C, 1 s for 40 cycles); 85 °C, 5 min; 4 °C hold. 2.5 μL of RT product (cDNA) was added to 22.5 μL of PreAmp Reaction mix, containing Megaplex™ RT Primers Pool A or B and TaqMan® PreAmp Master Mix to increase the quantity of cDNA prior to PCR on the Taqman® Open Array® Micro-RNA Panels. The final volume (25 μL) of preamplification reaction mix underwent the following thermo cycling conditions: 95 °C 10 min; 55 °C, 2 min; 72 °C, 2 min; (95 °C, 15 s, 60 °C, 4 min for 12 cycles), 99.9 °C, 10 min; 4 °C hold. Four microlitres of each preamplified product was diluted in ultrapure water (156 μL) to give a final dilution of 1:40 as per recommended protocol. For the real-time PCR 22.5 μL of Taqman® Open Array® Real-Time PCR Master Mix was added to 22.5 μL of preamplification product to give a total volume of 45 μL. Five microlitres of each PCR Reaction mix was added to 8 wells on an OpenArray® 384-Well-sample plate. The Taqman® Open Array® Micro-RNA Panel was loaded with the samples from the 384-Well-sample plate using the standard AccuFill™ method. Open Array® Real-Time qPCR was performed on the loaded Taqman® Open Array® Micro-RNA Panel using an Open Array® Real-Time PCR instrument and recommended software.
Raw fluorescence data was exported from the OpenArray® Real-Time qPCR Analysis Software (BioTroveTM, version 1.0.4) to a comma delimited text file. A Ct (cycle threshold) value was determined for each individual qPCR assay by using the statistical software R (version 3.0.2) to fit a 3-parameter logistic curve, assuming an amplification efficiency of 2, to the raw fluorescence data of each microRNA, and the Ct of each qPCR was determined using the second derivative maximum of the fitted logistic curve. PCR reactions that did not amplify were assigned a Ct value of 40.
To investigate whether any of the samples had low quality data we used the “detector profiling across samples” module in the RealTime PCR Statminer® software analysis program (v4.5, Integromics) to examine the correlations of Ct values between samples from the same epithelial tissue type across all of the amplified microRNAs. Samples that had multiple outliers were excluded from further analysis.
For normalisation of the OpenArray® microRNA expression data we selected 14 microRNAs using the following criteria: (1) they were expressed in all samples and at high levels (median Ct < 30); (2) they were not statistically different in epithelial tissue type comparisons (Welch's t-test, P > 0.1); and (3) they were the least variable miRNAs (coefficient of variation < 1.0 for relative levels in each epithelial tissue type). The values for these selection criteria for each of the 14 microRNAs used for normalisation, plus mature nucleic acid sequences and Accession numbers, are presented in Supplementary Table 1.
The relative levels of the microRNAs were determined using the formula 2(40-Ct), and were normalized using the geometric mean of the relative levels of the 14 House Keeping Genes. The data was pre-filtered using the following criteria to include microRNAs that were more likely to be informative: (1) each microRNA had to have at least 50% of samples amplified in one of the comparison groups; and (2) the differential expression between groups had to be greater than 1.4 fold. Mann Whitney U tests were then used to discover differentially expressed microRNAs in control squamous mucosa vs post-ablation neosquamous mucosa, and in post-ablation squamous mucosa vs post-ablation neosquamous mucosa. False discovery rates (the proportions of false positives) were estimated for each epithelial tissue type comparison. MicroRNAs that had P < 0.05 in both of these epithelial tissue type comparisons were termed “overlapping miRNAs”. Subset analyses were subsequently performed for these overlapping microRNAs in 2 sub-groups: (1) patients who were treated either medically or surgically for reflux; and (2) patients in whom complete ablation was achieved vs all patients. This was done by averaging the differential expression and the Mann Whitney U test P values in each patient subgroup for the overlapping microRNAs. Differences between the groups in (1) differential expression; and (2) Mann Whitney U test P values were tested using Welch’s t-test. The statistical methods of this study were reviewed by Professor Richard Woodman from Flinders University.
The overlapping miRNAs were further investigated to compare the direction of differential expression of these microRNAs in post-ablation neosquamous mucosa vs control squamous mucosa and post-ablation squamous mucosa, to the direction of differential expression in Barrett’s esophagus mucosa vs control squamous mucosa and post-ablation squamous mucosa. The potential roles of the overlapping miRNAs in regulating cellular processes were investigated using biological pathway enrichment analysis (described in next section).
To identify highly predicted mRNA targets of the differentially expressed microRNAs in neosquamous mucosa, we used the Predicted Target Module of miRWalk v2[23] (http://zmf.umm.uni-heidelberg.de/apps/zmf/mirwalk2/). To generate the putative target genes list we used a minimum seed length of 7 and/or P value < 0.05, from position 1 of the 3’ UTR, and included extra databases: RNA22 (https://cm.jefferson.edu/rna22/), miRanda (http://www.microrna.org/microrna/home.do) and Targetscan (http://www.targetscan.org/vert_71/). The predicted lists for each microRNA were then screened to identify mRNAs that were predicted to be the targets of at least two different microRNAs.
To identify pathways containing a statistically significant number of predicted targets, we used a publicly available, manually curated signalling pathway database[24] (http://www.innatedb.com/redirect.do?go=batchPw). The target list was subjected to a Pathway Enrichment Analysis which groups target genes according to function, and identifies further components and associated networks. The target list (in REFSeq ID format) was analysed using InnateDB, which identifies statistically enriched pathways by testing for over-representation using the Hypergeometric distribution (by default; other distributions are available), and by using the Benjamini Hochberg correction for multiple tests (by default). InnateDB uses multiple curated databases for the pathway analysis: Reactome (http://www.reactome.org/), KEGG (http://www.genome.jp/kegg/), PID Biocarta and PID NCI (http://www.home.ndexbio.org/), NetPath (http://www.netpath.org/), INOH (only available within InnateDB).
Forty-four microRNAs were differentially expressed at P < 0.05 between control squamous mucosa and post-ablation neosquamous mucosa (Supplementary Table 2). Thirty-three of these microRNAs had higher expression in post-ablation neosquamous mucosa vs control squamous mucosa, and 25 of these had a fold difference greater than 2. There were 11 microRNAs that had lower expression in post-ablation neosquamous mucosa vs control squamous mucosa, although only 2 of these were expressed at levels of 50% or less.
Nineteen microRNAs were differentially expressed at P < 0.05 between post-ablation neosquamous and post-ablation squamous mucosa (Supplementary Table 3). Fourteen microRNAs had higher expression and 5 microRNAs had lower expression in post-ablation neosquamous mucosa compared with post-ablation squamous mucosa.
Due to the large number of microRNAs that were assayed it is possible that some differentially expressed microRNAs occurred by chance alone and are thus false positives. We therefore estimated the false discovery rate (FDR) in each epithelial tissue type comparison: in control squamous vs post-ablation neosquamous mucosa the FDR was 11%, and in the post-ablation squamous vs post-ablation neosquamous mucosa the FDR was 19%. The post-ablation squamous vs post-ablation neosquamous comparison could also identify microRNAs associated with differential expression along the length of the esophagus[25]. In order to address these issues we investigated whether there were microRNAs that were differentially expressed in both epitheilial tissue type comparisons. We reasoned that because the control squamous mucosa samples were obtained from different patients to the post-ablation squamous mucosa, any differentially expressed microRNAs found in both mucosal comparisons were much less likely to be due to chance alone. This approach identified 12 microRNAs that were present in both control squamous vs post-ablation neosquamous mucosa and the post-ablation squamous vs post-ablation neosquamous mucosa groups (Table 1). Scatter plots for the 12 overlapping microRNAs are in Supplementary Figure 2. OpenArray assay identifiers, miRBase names and accession numbers, and mature nucleotide sequences for these overlapping microRNAs are in Supplementary Table 4. All of these microRNAs were more highly expressed in post-ablation neosquamous tissues, and 10 of the 12 overlapping microRNAs had similar fold differences in the two groups. In both comparisons miR-424-5p was the most significantly differentially expressed microRNA (Table 1). We further investigated these overlapping miRNAs in subsets of the data (patients with complete ablation vs all patients, and patients who were medically treated vs surgically treated for reflux) and did not find significant differences in differential expression between these subset groups (Supplementary Tables 7 and 8).
| Mature miRNA | Post-NS/control-S | P value | Post-NS/ Post-S | P value | Higher in pre-BE vs post-NS |
| miR-424-5p | 485.2 | 0.00002 | 233.7 | 0.00053 | Yes |
| miR-135b-5p | 2.0 | 0.00071 | 1.7 | 0.00363 | No |
| miR-376c-3p | 6.4 | 0.00145 | 3.1 | 0.02673 | Yes |
| miR-135a-5p | 3.0 | 0.00224 | 2.5 | 0.00821 | Yes |
| miR-187-3p | 208.1 | 0.00414 | 2.7 | 0.01196 | No |
| miR-409-3p | 6.8 | 0.00502 | 4.6 | 0.00272 | Yes |
| miR-214-5p | 47.0 | 0.00502 | 46.1 | 0.02673 | Yes |
| miR-31-5p | 1.44 | 0.00869 | 1.5 | 0.00632 | No |
| miR-199a-5p | 396.3 | 0.01223 | 512.9 | 0.04478 | Yes |
| miR-223-5p | 230.9 | 0.02306 | 204.5 | 0.01350 | Yes |
| miR-127-3p | 4.8 | 0.02675 | 3.8 | 0.03305 | Yes |
| miR-136-3p | 7.2 | 0.02675 | 201.2 | 0.03305 | Yes |
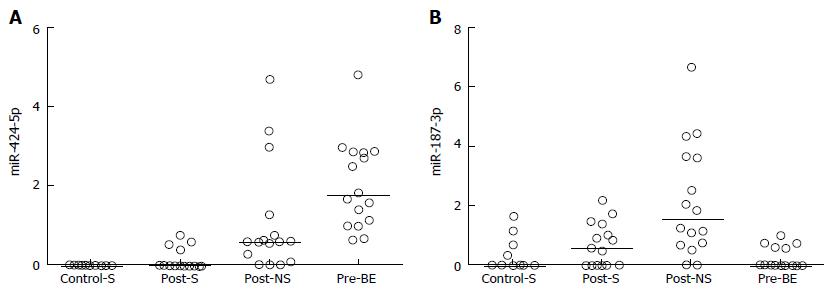
The 12 overlapping microRNAs were further investigated to determine their levels of expression in pre-ablation Barrett’s esophagus mucosa. The expression levels of 9 of the 12 overlapping microRNAs were higher in Barrett’s esophagus mucosa than in the squamous mucosa, and for all of these 9 microRNAs their expression in post-ablation neosquamous mucosa was in the same direction (i.e., higher) as in the Barrett’s esophagus mucosa (Table 1; Figure 1A for a representative example). For the remaining 3 microRNAs the levels in the Barrett’s esophagus mucosa were not different to the non-neosquamous mucosa (Figure 1B for a representative example).
For the 3 overlapping microRNAs that were found not to be increased in Barrrett’s esophagus mucosa relative to non-neosquamous mucosa mirWalk predicted 1566 mRNA targets, and 163 mRNAs with 2 or more microRNAs targeting them (Supplementary Table 5). Pathway analysis using InnateDB indicated that the predicted target mRNAs are involved in active membrane transport and in calcium signalling (Table 2). For the 9 microRNAs that are increased in both neosquamous and Barrett’s esophagus mucosa relative to squamous mucosa mirWalk predicted 3297 mRNA targets, and 839 mRNAs with 2 or more microRNAs targeting them (Supplementary Table 6). Pathway analysis using InnateDB indicated that the predicted target mRNAs are involved in hemostasis and in cell survival pathways (Table 3).
| Pathway name | Pathway uploaded gene count | Genes in InnateDB for this entity | Pathway P value | Pathway P value (corrected) |
| Ion transport by P-type ATPases | 5 | 43 | 1.57E-05 | 0.005 |
| Transmembrane transport of small molecules | 14 | 606 | 1.31E-04 | 0.022 |
| Ion channel transport | 7 | 169 | 2.56E-04 | 0.028 |
| Calcium signaling pathway | 7 | 183 | 4.15E-04 | 0.035 |
| Pathway name | Pathway uploaded gene count | Genes in InnateDB for this entity | Pathway P value | Pathway P value (corrected) |
| JAK STAT pathway and regulation | 28 | 273 | 5.41E-06 | 0.005 |
| Hemostasis | 41 | 508 | 1.68E-05 | 0.005 |
| Regulation of bad phosphorylation | 7 | 23 | 2.36E-05 | 0.006 |
Over the last decade, endoscopic treatment of high grade dysplasia and early cancer arising in Barrett’s esophagus has largely superseded surgical resection, because of the perception of reduced morbidity, and the lower risk of lymph node metastases when cancer stage is limited to no worse than stage T1a[26,27]. Consensus guidelines for endoscopic therapy suggest that complete eradication of all Barrett’s esophagus mucosa is required to eliminate the risk of metachronous and covert synchronous neoplasia[28,29]. However, increasing evidence suggests that complete eradication of Barrett’s esophagus might not eliminate the risk of cancer in some patients. For example, Templeton et al[11] (2014) reported three patients who progressed to invasive adenocarcinoma despite prior complete eradication of Barrett’s esophagus using endoscopic therapy. Even after re-treatment and complete endoscopic eradication of the post-ablation recurrences of Barrett’s esophagus, Guthikonda et al[12] (2016) reported a progression rate to invasive cancer of 2.1% per year, and predicted that 5.1% (Kaplan Meier model estimate; 95%CI: 0.0-11.3) of re-treated patients would experience invasive cancer progression by 5 years after complete eradication of Barrett’s esophagus[12]. In a meta-analysis of 21 radiofrequency ablation studies that reported 603 cases of Barrett’s esophagus recurrence from 3186 patients, pooled incidence ratios (IR’s) of recurrent Barrett’s esophagus, dysplastic Barrett’s esophagus, and HGD/EAC were 9.5% (95%CI: 6.7-12.3), 2.0% (95%CI: 1.3-2.7), and 1.2% (95%CI: 0.8-1.6) per patient-year, respectively[13]. Ongoing endoscopic surveillance has therefore been recommended after eradication of Barrett’s esophagus to monitor for recurrence and disease progression.
It has been suggested that residual sub-squamous glandular tissue after ablation may contribute to the progression of Barrett’s esophagus to high grade dysplasia and adenocarcinoma[30,31]. For example, it has been reported that after argon plasma coagulation ablation buried glandular tissue underneath neosquamous mucosa had higher levels of cancer associated biomarkers (Ki67, COX-2, BCL-2) than normal esophageal epithelium[32]. However, Basu et al[8] (2002) reported that the presence of buried Barrett’s glands was not associated with recurrence in patients with effective acid suppression after argon plasma coagulation ablation. Furthermore, neosquamous epithelium has been reported not to contain genetic abnormalities after radio frequency ablation in patients who had pre-ablation Barrett’s esophagus containing early cancer or high-grade dysplasia[9].
In an earlier study we investigated neo-squamous mucosa in patients who had undergone argon plasma coagulation ablation of non-dysplastic Barrett’s esophagus, and observed that expression levels of miR-143 were elevated in neosquamous epithelium and biopsies from squamous epithelium above the metaplastic segment, compared to squamous epithelium from controls without Barrett’s esophagus[15]. miR-143 has also been shown to have increased expression in Barrett’s esophagus mucosa. This study suggested that post-ablation neosquamous epithelium might not be normal, and might express persistent molecular markers consistent with the original Barrett’s esophagus.
In our current study we used post-ablation mucosa from patients who did not have dysplasia or early cancer to further investigate the biology of the neosquamous epithelium, and it is unlikely that these biopsies would have contained residual dysplasia associated biomarkers[32], or be effected by DNA mutations that are commonly found in dysplastic tissues[6].
We sought to strengthen our approach by restricting the list of microRNAs to those that are differentially expressed in both neosqamous vs independent squamous mucosa from control individuals, and in neosquamous vs paired proximal squamous mucosa comparisons. This approach should minimise the issues associated with inter-individual variation, epithelial repair, and tissue proximity to the gastro-esophageal junction as possible causes of differences in microRNA expression. This approach produced 12 microRNAs that were differentially expressed in both of the squamous mucosa comparisons.
To investigate the potential biological effects of the differential expression of these 12 microRNAs we utilized miRWalk to predict their mRNA targets, and InnateDB to assess which potential signalling pathways these mRNAs are involved in. This approach identified signalling pathways which the differentially expressed miRNAs might be regulating.
Nine microRNAs were expressed at higher levels in both neosquamous and pre-ablation Barrett’s mucosa, vs both squamous mucosa from controls and proximal squamous mucosa collected after ablation from the ablation patients. The predicted mRNA targets of these microRNAs are involved in the JAK-STAT signalling pathway, and in the regulation of the anti-apoptotic family member, Bad. Three of the discovered microRNAs have been reported to have targets involved in hemostasis[33-35].
The JAK-STAT signalling pathway transmits information from extracellular signals directly to the cell nucleus, and results in expression of genes involved in proliferation, differentiation, apoptosis and oncogenesis. The JAK-STAT3 pathway is activated by IL-6 in Barrett’s esophagus, and this promotes survival of the metaplastic intestinal cells[36,37]. However, activated JAK-STAT has been reported to be undetectable in normal esophageal squamous mucosa[38], so the effect of altered regulation of this pathway in neosquamous mucosa is not clear.
Three microRNAs were increased in neosquamous, but not pre-ablation Barrett’s esophagus mucosa, vs both healthy squamous and post-squamous mucosa. The predicted mRNA targets of these microRNAs are potentially involved in regulating pathways involved in transmembrane transport of small molecules, in ion channel transport, and in ion and lipid transport by P-type ATPases (which includes the calcium pump, Ca2+-ATPase). Jovov et al[39] (2013) found that post-ablation neosquamous epithelium has decreased barrier function, measured as persistent paracellular permeability to ions and uncharged molecules[39], so these active transporter targets are likely to be involved in the regulation of barrier protection. This is an important consideration because acid reflux injury has been implicated in the development of intestinal metaplasia, and decreased barrier function may therefore contribute to recurrence in patients with uncontrolled reflux following successful ablation.
The mucosal biopsies used in this study were from patients treated with Argon Plasma Coagulation ablation. These were collected as part of clinical trials previously established in our institution and were therefore readily available to us[20]. The use of biopsies from patients treated with argon plasma coagulation ablation is a potential limitation of our study, since radiofrequency ablation is now the standard technique for ablation of Barrett’s esophagus due to its ease of use and consistent depth of tissue destruction[40]. However, some patients in whom complete eradication has been achieved following radiofrequency ablation have still progressed to invasive cancer, which suggests that post radiofrequency ablation neosquamous epithelium may also not be normal. It is worth noting that the above described report of decreased barrier function in neosquamous mucosa was associated with radiofrequency ablation[39]. Future studies of neosquamous epithelium miRNA profile should therefore include other endoscopic methods such as radiofrequency ablation and endomucosal resection, and validation in independent cohorts.
The observed decrease in barrier function in neosquamous mucosa is consistent with reports of an association between recurrence after ablation and persistent reflux, and with reduced proton pump inhibitor dosing. Kahaleh et al[41] (2002) found that persistence of acid reflux and greater length of diseased segment were the major factors associated with a recurrence after successful initial reversal with argon plasma coagulation ablation. Basu et al[8] (2002) reported that patients who reduced their dose of the proton pump inhibitor omeprazole to 20 mg once daily or less after argon plasma coagulation ablation had significantly greater recurrence of intestinal metaplasia[8]. Conversely, in patients with complete squamous regeneration after argon plasma coagulation ablation who took a high dose of omeprazole (40 mg three times a day) there were no relapses or evidence of dysplasia under continuous acid suppression during a median follow-up of 12 mo (range 2 to 51 mo)[42].
Three (miR-424-5p, miR-223-5p, miR-409-3p) of the microRNAs that are increased in neosquamous mucosa relative to post-squamous and control-squamous mucosa have been reported to be up-regulated in esophageal adenocarcinoma relative to Barrett’s esophagus and normal squamous tissues. Wu et al[18] (2013) observed progressively increased expression of miR-424-5p, miR-223-5p and miR-409-3p from normal squamous epithelium to Barrett’s to adenocarcinoma. MiR-223-5p and miR-409-3p have also been reported to be overexpressed in serum exosomes from patients with esophageal adenocarcinoma[43].
In conclusion, this study demonstrates that the miRNA expression profile in neosquamous mucosa, following argon plasma coagulation ablation of Barrett’s esophagus, is significantly different from normal squamous mucosa. The main strength of this study is that the mucosal biopsies were not from patients who were treated for dysplasia or cancer. Our results suggest that altered miRNA expression may contribute to the previously reported defective barrier function in neosquamous epithelium, and this may place the mucosa at increased risk of disease progression relative to normal esophageal squamous mucosa. Further research to explore the roles of miRNAs in the response to ablation of Barrett’s esophagus, and the long term behaviour of neosquamous epithelium may lead to improvements in clinical management of this condition.
We thank the SA Pathology anatomical pathology laboratory for assistance with preparation of histopathology slides.
Overall 5 year survival of patients with esophageal adenocarcinoma is around 15%, but survival can be improved via early detection using endoscopic surveillance of patients with Barrett’s esophagus, and endoscopic ablative therapy for the treatment of early stage disease.
Some patients still experience disease progression after complete ablation of Barrett’s esophagus, which suggests that the neosquamous mucosa formed after ablation may be at increased risk of disease progression. Improved understanding of the biological properties of post-ablation neosquamous mucosa might improve the clinical management of patients who undergo ablative therapy.
Previous studies have investigated whether post ablation mucosa has genetic alterations in, or increased expression of cancer associated genes. Neither of these approaches directly addresses whether the non-cancer associated biology of neosquamous mucosa is different. Previous studies have reported reduced barrier function in neosquamous mucosa, and this may have implications for clinical management.
Altered microRNA expression in neosquamous mucosa might result in reduced barrier function, thereby placing the mucosa at increased susceptibility to reflux induced disease. MicroRNAs might therefore have the potential to be developed into a biomarker with clinical utility to improve the management of patients who have been treated endoscopically for early stage disease.
Post-ablation neosquamous mucosa: post-treatment regenerated esophageal squamous mucosa that was Barrett’s intestinal metaplasia prior to treatment, with biopsies taken 1 cm above the gastro-esophageal junction. Post-ablation squamous mucosa: post-treatment proximal normal non-treated esophageal mucosa, with biopsies collected 5 cm above the proximal margin of the pre-ablation Barrett’s mucosa. Healthy squamous mucosa: squamous mucosa from patients without esophageal disease, with biopsies taken from the same level relative to the gastro-esophageal junction as the post-ablation neosquamous mucosa.
This manuscript evaluates a topic of real interest. The analysis of the subject has been done in an appropriate way. The background of the problem was evaluated in a comprehensive way, the hypothesis was clearly stated, and the materials, methods and results are presented in an understandable way (I will add minor comments).
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: Australia
Peer-review report classification
Grade A (Excellent): A
Grade B (Very good): B
Grade C (Good): 0
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Durand L, Garcia-Olmo D S- Editor: Qi Y L- Editor: A E- Editor: Li D
| 1. | Chisholm JA, Mayne GC, Hussey DJ, Watson DI. Molecular biomarkers and ablative therapies for Barrett’s esophagus. Expert Rev Gastroenterol Hepatol. 2012;6:567-581. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3] [Cited by in F6Publishing: 3] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 2. | Phillips WA, Lord RV, Nancarrow DJ, Watson DI, Whiteman DC. Barrett’s esophagus. J Gastroenterol Hepatol. 2011;26:639-648. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 43] [Cited by in F6Publishing: 41] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 3. | Fassan M, Volinia S, Palatini J, Pizzi M, Baffa R, De Bernard M, Battaglia G, Parente P, Croce CM, Zaninotto G. MicroRNA expression profiling in human Barrett’s carcinogenesis. Int J Cancer. 2011;129:1661-1670. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 82] [Cited by in F6Publishing: 88] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 4. | Shaheen NJ, Richter JE. Barrett’s oesophagus. Lancet. 2009;373:850-861. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 234] [Cited by in F6Publishing: 225] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 5. | Haidry RJ, Dunn JM, Butt MA, Burnell MG, Gupta A, Green S, Miah H, Smart HL, Bhandari P, Smith LA. Radiofrequency ablation and endoscopic mucosal resection for dysplastic barrett’s esophagus and early esophageal adenocarcinoma: outcomes of the UK National Halo RFA Registry. Gastroenterology. 2013;145:87-95. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 173] [Cited by in F6Publishing: 160] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 6. | Hage M, Siersema PD, Vissers KJ, Dinjens WN, Steyerberg EW, Haringsma J, Kuipers EJ, van Dekken H. Genomic analysis of Barrett’s esophagus after ablative therapy: persistence of genetic alterations at tumor suppressor loci. Int J Cancer. 2006;118:155-160. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 29] [Cited by in F6Publishing: 29] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 7. | Hage M, Siersema PD, Vissers KJ, Steyerberg EW, Haringsma J, Kuipers EJ, van Dekken H. Molecular evaluation of ablative therapy of Barrett’s oesophagus. J Pathol. 2005;205:57-64. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 39] [Cited by in F6Publishing: 42] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 8. | Basu KK, Pick B, Bale R, West KP, de Caestecker JS. Efficacy and one year follow up of argon plasma coagulation therapy for ablation of Barrett’s oesophagus: factors determining persistence and recurrence of Barrett’s epithelium. Gut. 2002;51:776-780. [PubMed] [Cited in This Article: ] |
| 9. | Pouw RE, Gondrie JJ, Rygiel AM, Sondermeijer CM, ten Kate FJ, Odze RD, Vieth M, Krishnadath KK, Bergman JJ. Properties of the neosquamous epithelium after radiofrequency ablation of Barrett’s esophagus containing neoplasia. Am J Gastroenterol. 2009;104:1366-1373. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 91] [Cited by in F6Publishing: 85] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 10. | Ganz RA, Utley DS, Stern RA, Jackson J, Batts KP, Termin P. Complete ablation of esophageal epithelium with a balloon-based bipolar electrode: a phased evaluation in the porcine and in the human esophagus. Gastrointest Endosc. 2004;60:1002-1010. [PubMed] [Cited in This Article: ] |
| 11. | Templeton A, Bodnar A, Gan SI, Irani S, Ross A, Low D. Occurrence of invasive cancer after endoscopic treatment of Barrett’s esophagus with high-grade dysplasia and intramucosal cancer in physiologically fit patients: time for a review of surveillance and treatment guidelines. Gastrointest Endosc. 2014;79:839-844. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 5] [Cited by in F6Publishing: 5] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 12. | Guthikonda A, Cotton CC, Madanick RD, Spacek MB, Moist SE, Ferrell K, Dellon ES, Shaheen NJ. Clinical Outcomes Following Recurrence of Intestinal Metaplasia After Successful Treatment of Barrett’s Esophagus With Radiofrequency Ablation. Am J Gastroenterol. 2017;112:87-94. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 43] [Cited by in F6Publishing: 40] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 13. | Krishnamoorthi R, Singh S, Ragunathan K, A Katzka D, K Wang K, G Iyer P. Risk of recurrence of Barrett’s esophagus after successful endoscopic therapy. Gastrointest Endosc. 2016;83:1090-1106.e3. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 67] [Cited by in F6Publishing: 72] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 14. | Wijnhoven BP, Hussey DJ, Watson DI, Tsykin A, Smith CM, Michael MZ; South Australian Oesophageal Research Group. MicroRNA profiling of Barrett’s oesophagus and oesophageal adenocarcinoma. Br J Surg. 2010;97:853-861. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 98] [Cited by in F6Publishing: 109] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 15. | Dijckmeester WA, Wijnhoven BP, Watson DI, Leong MP, Michael MZ, Mayne GC, Bright T, Astill D, Hussey DJ. MicroRNA-143 and -205 expression in neosquamous esophageal epithelium following Argon plasma ablation of Barrett’s esophagus. J Gastrointest Surg. 2009;13:846-853. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 27] [Cited by in F6Publishing: 29] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 16. | Calin GA, Croce CM. MicroRNA-cancer connection: the beginning of a new tale. Cancer Res. 2006;66:7390-7394. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 787] [Cited by in F6Publishing: 844] [Article Influence: 46.9] [Reference Citation Analysis (0)] |
| 17. | Smith CM, Watson DI, Michael MZ, Hussey DJ. MicroRNAs, development of Barrett’s esophagus, and progression to esophageal adenocarcinoma. World J Gastroenterol. 2010;16:531-537. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 34] [Cited by in F6Publishing: 36] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 18. | Wu X, Ajani JA, Gu J, Chang DW, Tan W, Hildebrandt MA, Huang M, Wang KK, Hawk E. MicroRNA expression signatures during malignant progression from Barrett’s esophagus to esophageal adenocarcinoma. Cancer Prev Res (Phila). 2013;6:196-205. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 75] [Cited by in F6Publishing: 84] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 19. | Feber A, Xi L, Luketich JD, Pennathur A, Landreneau RJ, Wu M, Swanson SJ, Godfrey TE, Litle VR. MicroRNA expression profiles of esophageal cancer. J Thorac Cardiovasc Surg. 2008;135:255-260; discussion 260. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 282] [Cited by in F6Publishing: 301] [Article Influence: 17.7] [Reference Citation Analysis (0)] |
| 20. | Sie C, Bright T, Schoeman M, Game P, Tam W, Devitt P, Watson D. Argon plasma coagulation ablation versus endoscopic surveillance of Barrett’s esophagus: late outcomes from two randomized trials. Endoscopy. 2013;45:859-865. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 24] [Cited by in F6Publishing: 25] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 21. | Beck P, Mayne GC, Astill D, Irvine T, Watson DI, Dijckmeester WA, Wijnhoven BP, Hussey DJ. Accuracy of identification of tissue types in endoscopic esophageal mucosal biopsies used for molecular biology studies. Clin Exp Gastroenterol. 2009;2:1-7. [PubMed] [Cited in This Article: ] |
| 22. | McAlinden A, Varghese N, Wirthlin L, Chang LW. Differentially expressed microRNAs in chondrocytes from distinct regions of developing human cartilage. PLoS One. 2013;8:e75012. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 44] [Cited by in F6Publishing: 40] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 23. | Dweep H, Gretz N. miRWalk2.0: a comprehensive atlas of microRNA-target interactions. Nat Methods. 2015;12:697. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 905] [Cited by in F6Publishing: 983] [Article Influence: 109.2] [Reference Citation Analysis (0)] |
| 24. | Breuer K, Foroushani AK, Laird MR, Chen C, Sribnaia A, Lo R, Winsor GL, Hancock RE, Brinkman FS, Lynn DJ. InnateDB: systems biology of innate immunity and beyond--recent updates and continuing curation. Nucleic Acids Res. 2013;41:D1228-D1233. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 745] [Cited by in F6Publishing: 796] [Article Influence: 66.3] [Reference Citation Analysis (0)] |
| 25. | Ali I, Rafiee P, Hogan WJ, Jacob HJ, Komorowski RA, Haasler GB, Shaker R. Dickkopf homologs in squamous mucosa of esophagitis patients are overexpressed compared with Barrett’s patients and healthy controls. Am J Gastroenterol. 2006;101:1437-1448. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 17] [Cited by in F6Publishing: 16] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 26. | Dunbar KB, Spechler SJ. The risk of lymph-node metastases in patients with high-grade dysplasia or intramucosal carcinoma in Barrett’s esophagus: a systematic review. Am J Gastroenterol. 2012;107:850-862; quiz 863. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 172] [Cited by in F6Publishing: 168] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 27. | Fuchs HF, Harnsberger CR, Broderick RC, Chang DC, Sandler BJ, Jacobsen GR, Bouvet M, Horgan S. Mortality after esophagectomy is heavily impacted by center volume: retrospective analysis of the Nationwide Inpatient Sample. Surg Endosc. 2017;31:2491-2497. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 71] [Cited by in F6Publishing: 58] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 28. | Fitzgerald RC, di Pietro M, Ragunath K, Ang Y, Kang JY, Watson P, Trudgill N, Patel P, Kaye PV, Sanders S. British Society of Gastroenterology guidelines on the diagnosis and management of Barrett’s oesophagus. Gut. 2014;63:7-42. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 858] [Cited by in F6Publishing: 808] [Article Influence: 80.8] [Reference Citation Analysis (0)] |
| 29. | Spechler SJ, Sharma P, Souza RF, Inadomi JM, Shaheen NJ; American Gastroenterological Association. American Gastroenterological Association technical review on the management of Barrett’s esophagus. Gastroenterology. 2011;140:e18-e52; quiz e13. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 840] [Cited by in F6Publishing: 773] [Article Influence: 59.5] [Reference Citation Analysis (0)] |
| 30. | Titi M, Overhiser A, Ulusarac O, Falk GW, Chak A, Wang K, Sharma P. Development of subsquamous high-grade dysplasia and adenocarcinoma after successful radiofrequency ablation of Barrett’s esophagus. Gastroenterology. 2012;143:564-566.e1. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 99] [Cited by in F6Publishing: 93] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 31. | Lee JK, Cameron RG, Binmoeller KF, Shah JN, Shergill A, Garcia-Kennedy R, Bhat YM. Recurrence of subsquamous dysplasia and carcinoma after successful endoscopic and radiofrequency ablation therapy for dysplastic Barrett’s esophagus. Endoscopy. 2013;45:571-574. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 25] [Cited by in F6Publishing: 24] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 32. | Lewis CJ, Thrumurthy SG, Pritchard S, Armstrong G, Attwood SE. Comparison of COX-2, Ki-67, and BCL-2 expression in normal esophageal mucosa, Barrett’s esophagus, dysplasia, and adenocarcinoma with postablation mucosa and implications for ablative therapies. Surg Endosc. 2011;25:2564-2569. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 12] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 33. | Fort A, Borel C, Migliavacca E, Antonarakis SE, Fish RJ, Neerman-Arbez M. Regulation of fibrinogen production by microRNAs. Blood. 2010;116:2608-2615. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 76] [Cited by in F6Publishing: 83] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 34. | Landry P, Plante I, Ouellet DL, Perron MP, Rousseau G, Provost P. Existence of a microRNA pathway in anucleate platelets. Nat Struct Mol Biol. 2009;16:961-966. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 350] [Cited by in F6Publishing: 364] [Article Influence: 24.3] [Reference Citation Analysis (0)] |
| 35. | Edelstein LC, Simon LM, Montoya RT, Holinstat M, Chen ES, Bergeron A, Kong X, Nagalla S, Mohandas N, Cohen DE. Racial differences in human platelet PAR4 reactivity reflect expression of PCTP and miR-376c. Nat Med. 2013;19:1609-1616. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 156] [Cited by in F6Publishing: 163] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 36. | Zhang HY, Zhang Q, Zhang X, Yu C, Huo X, Cheng E, Wang DH, Spechler SJ, Souza RF. Cancer-related inflammation and Barrett’s carcinogenesis: interleukin-6 and STAT3 mediate apoptotic resistance in transformed Barrett’s cells. Am J Physiol Gastrointest Liver Physiol. 2011;300:G454-G460. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 46] [Cited by in F6Publishing: 53] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 37. | Dvorak K, Chavarria M, Payne CM, Ramsey L, Crowley-Weber C, Dvorakova B, Dvorak B, Bernstein H, Holubec H, Sampliner RE. Activation of the interleukin-6/STAT3 antiapoptotic pathway in esophageal cells by bile acids and low pH: relevance to barrett’s esophagus. Clin Cancer Res. 2007;13:5305-5313. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 68] [Cited by in F6Publishing: 70] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 38. | You Z, Xu D, Ji J, Guo W, Zhu W, He J. JAK/STAT signal pathway activation promotes progression and survival of human oesophageal squamous cell carcinoma. Clin Transl Oncol. 2012;14:143-149. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 34] [Cited by in F6Publishing: 35] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 39. | Jovov B, Shaheen NJ, Orlando GS, Djukic Z, Orlando RC. Defective barrier function in neosquamous epithelium. Am J Gastroenterol. 2013;108:386-391. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 23] [Cited by in F6Publishing: 18] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 40. | Haidry R, Lovat L, Sharma P. Radiofrequency ablation for Barrett’s dysplasia: past, present and the future? Curr Gastroenterol Rep. 2015;17:13. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 5] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 41. | Kahaleh M, Van Laethem JL, Nagy N, Cremer M, Devière J. Long-term follow-up and factors predictive of recurrence in Barrett’s esophagus treated by argon plasma coagulation and acid suppression. Endoscopy. 2002;34:950-955. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 76] [Cited by in F6Publishing: 81] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 42. | Schulz H, Miehlke S, Antos D, Schentke KU, Vieth M, Stolte M, Bayerdörffer E. Ablation of Barrett’s epithelium by endoscopic argon plasma coagulation in combination with high-dose omeprazole. Gastrointest Endosc. 2000;51:659-663. [PubMed] [Cited in This Article: ] |
| 43. | Warnecke-Eberz U, Chon SH, Hölscher AH, Drebber U, Bollschweiler E. Exosomal onco-miRs from serum of patients with adenocarcinoma of the esophagus: comparison of miRNA profiles of exosomes and matching tumor. Tumour Biol. 2015;36:4643-4653. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 79] [Cited by in F6Publishing: 94] [Article Influence: 10.4] [Reference Citation Analysis (0)] |