Published online Jan 14, 2014. doi: 10.3748/wjg.v20.i2.333
Revised: January 1, 2014
Accepted: January 5, 2014
Published online: January 14, 2014
Hepatocellular carcinoma (HCC), the predominant form of adult liver malignancies, is a global health concern. Its dismal prognosis has prompted recent significant advances in the understanding of its etiology and pathogenesis. The deregulation of epigenetic mechanisms, which maintain heritable gene expression changes and chromatin organization, is implicated in the development of multiple cancers, including HCC. This review summarizes the current knowledge of epigenetic mechanisms in the pathogenesis of HCC, with an emphasis on HCC mediated by chronic hepatitis B virus infection. This review also discusses the encouraging outcomes and lessons learnt from epigenetic therapies for hematological and other solid cancers, and highlights the future potential of similar therapies in the treatment of HCC.
Core tip: Hepatocellular carcinoma (HCC) is a global health concern; molecularly targeted therapeutics remains limited to sorafenib. New targets and drugs are urgently needed to broaden the limited treatment options for HCC. Many lines of evidence suggest that epigenetics is associated with the initiation and development of HCC. Here, we review the current state of knowledge on epigenetic deregulation in HCC, and potential therapies that can be exploited for interventions.
- Citation: Ma L, Chua MS, Andrisani O, So S. Epigenetics in hepatocellular carcinoma: An update and future therapy perspectives. World J Gastroenterol 2014; 20(2): 333-345
- URL: https://www.wjgnet.com/1007-9327/full/v20/i2/333.htm
- DOI: https://dx.doi.org/10.3748/wjg.v20.i2.333
Liver cancer is a global health concern. It is the fifth most frequently diagnosed cancer and second most frequent cause of cancer death in men worldwide. Liver cancer is endemic particularly in East Asia and Southeast Asia, where more than half of total cases globally are diagnosed. In addition, Middle Africa, West Africa and South Africa are among the top five regions affected[1]. In the United States, the incidence rate of hepatocellular carcinoma (HCC) (the most common histological subtype of liver cancer) has tripled in the past decade, with the rise being related to increased hepatitis C virus (HCV) infection[2,3]. Risk factors for HCC are well characterized, including chronic hepatitis B virus (HBV) infection, HCV infection, excessive alcohol consumption, diabetes, non-alcoholic fatty liver disease, and dietary exposure to aflatoxin[4].
Epigenetics is defined as heritable states of gene expression without altering DNA sequences. Epigenetic mechanisms encompass genomic DNA modifications (methylation of DNA cytosine bases), chemical modifications of histone tails, and non-coding miRNA regulation. During cell division, these epigenetic modifications are passed down faithfully to daughter cells to maintain “cellular memory”[5]. DNA methyltransferases (DNMTs) catalyze the addition of methyl groups (CH3) to the 5’ cytosine nucleotides. Mechanistically, DNA methylation leads to transcriptional gene silencing in two ways. First, methylation at CpG sites sterically hinders accessibility of transcription factors to their cognate binding sites on respective gene promoters[6]. The second mechanism involves direct binding of methyl CpG binding domain (MBD)-containing proteins to the methylated DNA, causing transcription repression[7]. Gene silencing mediated by DNA methylation is observed in many cancer types. Cancers often present with features of global hypomethylation; by contrast, promoters of tumor suppressor genes, in particular, are hypermethylated[8]. Several lines of evidence suggest that changes in the epigenome are associated with liver cancer initiation and progression[9].
Thus far, three independent genome-wide methylation profiling studies demonstrated that HCC tumors display differential DNA methylome patterns compared to the respective adjacent normal liver tissues[10-12]. Hernandez-Vargas et al[10] used a bead array to analyze 1505 CpG sites in 30 patients with either HBV- or HCV-associated HCC, and observed that HCC tumors exhibit specific DNA methylation signatures that are correlated with major risk factors and tumor progression stage, implying potential clinical applications in early diagnosis and prognosis. Specifically, a panel of hypermethylated gene promoters (APC, RASSFIA, CDKN2A and FZD7) were able to discriminate HCC tumors from paired surrounding non-tumor liver tissues. Another set of hypermethylated genes (e.g., NAT2, CSPG2 and DCC) were exclusively associated with HBV-related HCC[10]. In particular, promoter methylation of DNMT1 was found to significantly correlate with poor tumor differentiation. By using the latest Illumina Methylation450 BeadChip, which allowed measurement of DNA methylation levels at 485577 loci across 99% of RefSeq genes (including 96% of the known CpG islands), Song et al[11] measured DNA methylation levels in HCC tissues or adjacent normal liver tissues from 27 HCC patients, and found significant enrichment of promoter CpG island DNA methylation loci in the signaling networks of cellular development, gene expression, cell death, and cancer. The genes BMP4, CDKN2A, GSTP1, and NFATC1 were among the top of the gene list. Shen et al[12] carried out a genome-wide methylation study using plasma DNA from a cohort consisting predominantly of HBV+ (79%) HCC patients, and found that the top five hypermethylated genes were DAB2IP, BMP4, ZFP41, SPDY1 and CDKN2A, whereas the top five hypomethylated genes were CCL20, ATK3, SCGB1D1, WFDC6 and PAX4.
Functional consequences of methylation-mediated silencing of tumor suppressor genes (TSGs) were addressed by recent studies by Nishida et al[13] and Revill et al [14]. Using combined genome-wide methylation profiling and “epigenetic unmasking” approaches in primary HCC and gene re-expression in cell lines, Revill et al[14] narrowed down 13 relevant candidate TSGs to SMPD3 and NEFH. Overexpression of SMPD3 and NEFH led to repression of cell proliferation, with NEFH causing a lesser effect. Nishida et al[13] identified a panel of eight TSGs that are highly predictive of progression from chronic HCV infection to HCC. However, the biological functions of these TSGs may vary depending on the cellular context, and their roles in HCC remain to be validated in HCC arising from other risk factors. DNA methylation is therefore a significant mechanism in the silencing of these TSGs.
The above studies provide strong evidence that aberrant gene methylation is closely associated with disease stage and clinical outcome in HCC, and suggest that methylation profiling (in particular, using patient plasma) may be a feasible approach for early diagnosis and prognosis of HCC. However, specific gene methylation signatures remain to be validated.
Within the chromosome, DNA is packaged into chromatin where the DNA coils around an octamer of histones. One hundred and forty-five base pairs of DNA are wrapped around the histone octamer, comprising H2A, H2B, H3 and H4, forming the repeating unit of chromatin, the nucleosome[15]. Histone tails protruding out of the nucleosome are targets of post-translational modifications, including acetylation and methylation of lysine (K) and arginine (R) residues, phosphorylation of serine (S) and threonine (T) residues, and ubiquitination of lysine residues[16]. These modifications can turn transcription of genes on or off, and are therefore key players in establishing the gene expression patterns of cells by adjusting the tightness of DNA bound to histones, thereby affecting accessibility of transcription factors[17].
Histone acetylation is controlled by two families of enzymes: histone acetyltransferases (HATs) that “write” the acetyl mark. Acetylation counteracts the positive charge of histones, thereby loosening the tight interaction between histones and DNA. Conversely, histone deacetylases (HDACs) “erase” the acetyl group, resulting in tight coiling of DNA around the histones, leading to the transcriptionally inactive or closed chromatin state[18]. In contrast, histone methylation is associated with either transcriptionally active or closed chromatin, depending on which histone or which lysine residue is modified. For example, histone 3 lysine 27 trimethylation (H3K27me3) is associated with transcriptional repression, whereas trimethylation of lysine 4 of histone 3 is indicative of gene activation[19,20].
Evaluation of histone methylation status in HCC remains limited to correlative studies with clinicopathlogical features of HCC, using semi-quantitative methods of protein detection such as immunohistochemistry or Western blotting. High levels of trimethylated histone H3 lysine 4 (H3K4me3) were correlated with reduced overall survival and poor prognosis in HCC[21]. Another study showed that high levels of H3K27me3 predicted worse prognosis, and were additionally closely correlated with aggressive tumor features, including vascular invasion, large tumor size, multiplicity of tumors, and poor differentiation[22]. Further studies using more precise detection methods, such as ChIP-sequencing, will be required to analyze these specific DNA-protein modifications in order to fully understand their roles in HCC.
Polycomb-group proteins are chromatin-modifying complexes mediating heritable gene silencing. Polycomb repressive complexes (PRCs) function in the maintenance of cell lineage commitment and stem cell pluripotency. The PRC1 complex comprises the core protein BMI, and RING1A and RING1B, which work as ubiquitin ligases for H2AK119. The other associated protein CBX7 binds to H3K27 via its chromodomain[23]. The polycomb repressive complex 2 (PRC2) complex consists of SUZ12, EZH1/2, EED1, and RbAp48. EZH2 is a methyltransferase that mediates gene silencing by trimethylating H3K27[24]. During embryonic stem cell development, the Suz–/– and Ezh2–/– cells exhibit distinct defects during gastrulation. Loss of Suz12 destabilizes Ezh2, causing a global loss of H3K27me3[25].
Elevated expression of EZH2 has been reported in breast[26,27] and prostate[28] cancers. EZH2 mRNA transcript[29] and protein[30,31] levels were consistently elevated in HCC in comparison to non-tumor liver tissues. Specifically, clinicopathological analysis of paired resected tumor and non-tumor tissues showed that high levels of EZH2 were strongly associated with aggressive and metastatic features (including portal vein invasion and lack of tumor encapsulation)[29], and with poor prognosis[30], although no significant differences were observed in either disease free survival[29] or cumulative survival rate between high and low EZH2 expression groups[31].
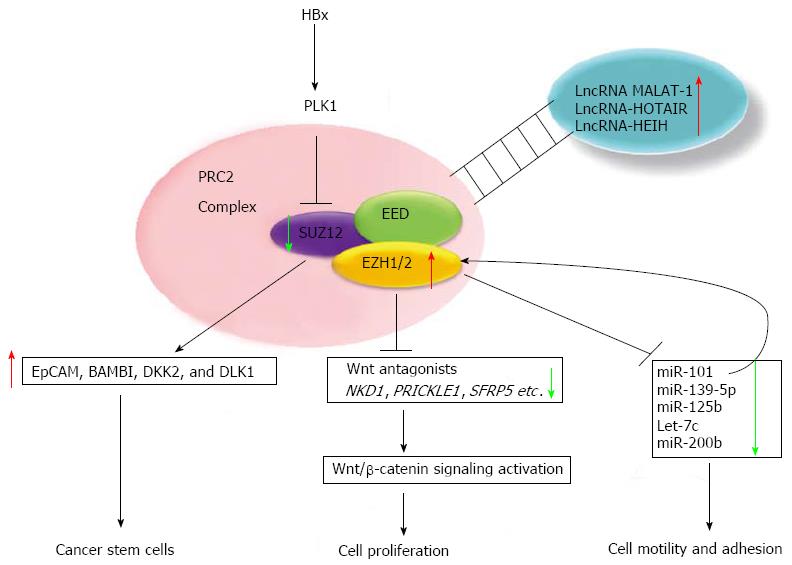
Detailed mechanistic studies have further elucidated the biological roles of EZH2 in HCC pathogenesis, which support the above clinical correlations. For example, EZH2 was shown to silence WNT antagonists, thereby activating Wnt/β-catenin signaling to promote cancer progression[32]. In contrast, knockdown of EZH2 in liver cancer cell lines reduces the repressive H3K27me3 marker, leading to re-expression of a distinct subpopulation of tumor suppressor miRNAs (miR-139-5p, miR-125b, miR-101, let-7c, and miR-200b), which control motility and adhesion[33]. Another study showed that knockdown of EZH2 profoundly inhibited proliferation of Dlk+ hepatic progenitor cells, promoting their differentiation into hepatocytes[34]. Additionally, Wang et al[35] reported that c-Myc together with EZH2 silences the tumor suppressive miRNA-101, which in turn targets the PRC2 complex in a double negative feedback loop fashion to account for the overexpression of EZH2 in HCC. Similarly, overexpression of EZH2 resulting from aberrant genomic loss of miR-101 was also reported in prostate cancer[36]. Taken together, these findings show the combined regulatory effects of chromatin-modifying activities and miRNA expression in promoting HCC progression, and provide evidence for an essential role of EZH2 in hepatic progenitor cell homeostasis.
In another study, the role of the viral HBx encoded by HBV was studied for its contribution to hepatocyte transformation. HBx is weakly oncogenic, and essential in the HBV life cycle[37,38]. HBx activates mitogenic pathways and increases polyploid cells (> 4 N), which causes genetic instability[39]. Moreover, HBx activates mitotic polo-like kinase (PLK) 1[40], which likely downregulates the PRC2 component Suz12 via phosphorylation. Elevated PLK1 and reduced protein levels of Znf198 and Suz12 are also observed in human HCC cell lines, as well as in liver tumors from X/c-myc bi-transgenic mice and woodchucks infected with the woodchuck hepatitis virus[41]. Importantly, loss of Suz12 results in de-repression of a subset of PRC2 target genes, specifically those with elevated expression in hepatic cancer stem cells, including EpCAM, BAMBI, DKK2, and DLK1[41,42]. These findings suggest that chronic HBV infection may give rise to a small population of cells with hepatic cancer stem cell properties, which ultimately could contribute to the proliferation and progression of HCC.
To summarize, PRC2 subunits SUZ12 and EZH2 have distinct roles during HCC pathogenesis. Over-expression of EZH2 is consistently found in advanced HCC. This elevated expression was associated with late stage features such as invasion and metastasis. In contrast, in the setting of chronic HBV infection, the HBx protein modulates SUZ12 protein levels, thereby maintaining the “stemness” of a subpopulation of hepatocyte stem/progenitor cells. Depending on how the risk factors interact with the host DNA and epigenetic players, each specific epigenetic modifier component may play a distinct role at different HCC stages.
MicroRNAs (miRNAs) are non-coding small RNA (ncRNA) molecules that are 20-23 nucleotides in length. They play important regulatory roles in plants and animals by targeting mRNAs for cleavage or translational repression. More than 1000 miRNAs have been identified to date. Through their roles in post-transcriptional gene regulation, miRNAs regulate diverse cellular functions including proliferation, differentiation, apoptosis, cell fate, and plasticity[43]. Pri-miRNAs are transcribed by RNA polymerase II either from their own gene or are located in introns of protein-coding genes. The pri-miRNA is then cleaved in the nucleus to form an approximately 60-70 nt stem loop intermediate, known as the miRNA precursor, or the pre-miRNA, which is further exported from the nucleus to the cytoplasm by Ran-GTP and the export receptor exportin-5. In the cytoplasm, further processing by Dicer, another RNase III endonuclease, generates the 5’ phosphate and approximately 2 nt 3’ overhang characteristic of an RNase III and produces an siRNA-like imperfect duplex that comprises the mature miRNA. One strand of the duplex is incorporated into the RNA-induced silencing complex, forming a complementary complex with the 3’-untranslated region of the target mRNA, and resulting in mRNA degradation or inhibition of mRNA translation, and hence gene expression silencing[44].
It has been increasingly recognized that aberrant miRNA expression profiles are linked to liver cancer development and progression[45]. MiRNAs play a role in virus-host interaction, and provide an anti-viral defense mechanism, such as against the HCV and the primate foamy virus type 1[46,47]. In a miRNA library screen, miR-141 was shown to repress HBV expression and replication in HepG2 cells, via direct suppression of the nuclear receptor peroxisome proliferator-activated receptor (PPAR)-α. PPAR-α regulates HBV gene expression through interactions with HBV promoter regulatory elements[48]. Another miRNA of interest is miR-122, the most abundant miRNA expressed in the liver, making up as much as 72% of total liver miRNAs[49]. Interestingly, miR-122 exerts opposite functions in HBV and HCV replication[50,51]. It can bind directly to a region of the HBV pre-genomic RNA and negatively regulates HBV replication. An inverse correlation was observed between miR-122 and HBV genome copies in peripheral blood mononuclear cells obtained from HBV+ samples[50]. On the other hand, miR-122 is essential for the stability, propagation and replication of HCV RNA[51]. Indeed, an antisense oligonucleotide (Miravirsen) with locked nucleic acid (LNA)-modified DNA that sequesters mature miR-122 has shown high efficacy in reducing HCV RNA levels in human clinical trials[52]. However, miR-122 is frequently downregulated in HCC[53,54], suggesting its tumor suppressor role[55]; therefore, attempts to restore miR-122 expression may inhibit HCC, or theoretically promote HCV replication and HCC, and have to be carefully considered depending on the viral status of HCC patients.
In contrast to miR-122, miR-1 over-expression increases HBV replication via induction of the hepatic nuclear receptor farnesoid X receptor, which then enhances transcription of the HBV core protein. This is accompanied by cell cycle arrest in G1 phase and differentiation of hepatocytes, thereby providing a favorable environment for HBV replication[56]. Furthermore, in cultured liver cancer cells, the HBx protein represses p53-mediated expression of miRNA-148a, which in turn down-regulates the hematopoietic pre-B cell leukemia transcription factor-interacting protein (HPIP). In HBV-mediated HCC, expression of miR-148a is reduced, whereas that of HPIP is elevated. These observations support that down-regulation of miR-148a has a role in liver cancer pathogenesis[57]. Other miRNAs with reported roles in HCC are listed in Table 1, and are discussed in other comprehensive reviews[58,59].
| Role | miRNA | Characteristics | Ref. |
| Viral replication | miR-141 | Represses HBV expression and replication | [48] |
| miR-122 | Inhibits HBV replication | [50] | |
| miR-122 | HCV RNA stabilization, propagation and replication | [51] | |
| miR-1 | Increases HBV replication | [56] | |
| Tumor supressor | miR-138 | Down-regulated in HCC tissues. miR-138 can directly target cyclin D3 | [61] |
| miR-26a and miR-26b | Down-regulated in tumors compared to paired non-tumor tissues | [64] | |
| miR-125b | Arrests cell cycle progression, and inhibits migration and invasion by directly targeting the oncogene LIN28B2 | [65] | |
| miR-140-5p | Suppresses tumor growth and metastasis by targeting TGFBR1 and FGF9 | [66] | |
| miR-122a | Mice lacking the gene encoding miR-122a are viable but develop temporally controlled steatohepatitis, fibrosis, and HCC | [68] | |
| Target tumor initiating cells | miR-181 | Maintains the stemness of liver cancer stem cells, target liver differentiation transcription factors CDX2 and GATA6 | [78] |
| miR-150 | Overexpression led to reduction of CD133+ cells | [80] | |
| miR-548c-5p | Ectopic overexpression inhibited proliferation, migration, and invasion of CD90+ HepG2 cells by down-regulating the expressions of β-catenin, Bcl-2, Tg737, Bcl-XL, and caspase 3 | [81] |
Profiling of differentially expressed miRNAs and their targets at different disease stages also suggests that miRNAs are associated with disease pathogenesis. In HBV-associated HCC, using TaqMan low-density miRNA arrays, Wang et al[60] found that miR-138 and miR-199a-5p expression was deregulated. In several studies, miR-138 levels were down-regulated in HCC tissues. MiR-138 exerts tumor suppressor function by directly targeting cyclin D3 and resulting in cell cycle arrest. It was also demonstrated that the use of a miR-138 mimic significantly reduced xenograft tumor growth in nude mice[61]. Differential expression levels of miR-199a-5p were also reported in HCC tissues and cell lines[62,63]. In particular, miR-199a-5p inhibits cell invasion by targeting discoidin domain receptor 1, a tyrosine kinase involved in signaling pathways that mediate cell invasion[62].
In a cohort of HCC patients characterized with a background of HBV infection (approximately 90%), miR-26a and miR-26b expression in non-liver tissues was higher in women than in men, suggesting that they have a protective role[64]. Moreover, miR-26a and miR-26b expression was down-regulated in tumors compared to paired non-tumor tissues. Patients with low miR-26 expression were associated with shorter survival, but were more likely to respond to interferon α therapy, making it an ideal candidate for predicting therapy response[64]. Additional miRNAs with a role in HCC are described below. Another miRNA that is suppressed in human liver cancer is miR-125b, which possesses tumor-suppressive functions such as arresting cell cycle progression, and inhibiting migration and invasion by directly targeting the oncogene LIN28B2[65]. MiR-140-5p expression is also decreased in HCC as well as six HCC cell lines. MiR-140-5p suppresses tumor growth and metastasis by targeting TGFBR1 and FGF9[66]. Genome-wide miRNA and mRNA profiling during mouse liver development implicates miR-302b and miR-20a in repressing transforming growth factor-β signaling[67]. MiR-122, in addition to its anti-HBV role, also functions as a tumor suppressor. Mice lacking the gene encoding miR-122a are viable but develop temporally controlled steatohepatitis, fibrosis, and HCC[68]. Restoration of miR-122 in a hepatic cell line reverses its migration and invasion properties[69].
The hypothesis of liver cancer stem cells (LCSCs) is supported by the identification of subpopulations of cancer cells with antigenic markers, which contribute to cancer origin and chemoresistance[70,71]. To date, the three main LCSC markers are epithelial cell adhesion molecule (EpCAM)[72,73], CD133[74,75] and CD90[76,77]. Cells bearing these markers demonstrate cancer stem cell properties including: (1) ability to form tumorspheres in anchorage-independent assays; (2) ability to initiate tumor growth in vitro and in vivo; and (3) ability to self-renew. MiR-181 is highly expressed in EpCAM-positive HCC cells isolated from α-fetoprotein-positive tumors, and is important in maintaining the stemness of LCSCs by directly targeting hepatic differentiation transcription factors CDX2 and GATA6[78]. MiR-181 levels are transcriptionally induced by the Wnt/beta-catenin signaling pathway[79]. In addition, over-expression of miR-150 leads to significant reduction of CD133+ cells, as well as inhibition of cell proliferation and tumorsphere formation[80]. MiR-548c-5p, miR-198, miR-375, and miR-874 levels are decreased, whereas miR-155, miR-198, and miR-1289 levels are increased in CD90+ in comparison to CD90– HepG2 cells. Transfection with exogenous miR-548c-5p inhibited proliferation, migration, and invasion of CD90+ HepG2 cells by down-regulating the expressions of β-catenin, Bcl-2, Tg737, Bcl-XL, and caspase 3[81].
Long non-coding RNAs (lncRNA) are transcripts longer than 200 nt, constituting a subpopulation of ncRNAs. They exert molecular regulatory functions via diverse modes of mechanisms[82]. Accumulating evidence indicates that lncRNAs are implicated in many cellular functions and play a role in carcinogenesis of multiple cancer types[83]. It has been shown that 20% of lncRNAs are associated with PRC2, through which they recruit and guide chromatin modifying complexes to specific genomic regions to regulate gene expression[84]. They behave like transcription co-activators/repressors by directly binding with various interaction partners; for example, lncRNA TERRA can directly bind to human telomerase and inhibit telomerase activity[85]. Alternatively, they act as decoys competing for miRNAs to modulate the expression of target genes[86].
LncRNA HULC (highly up-regulated in liver cancer) was the first lncRNA with highly specific up-regulation detected in the blood of HCC patients[87]. Du et al[88] further revealed that HBx could regulate the promoter of HULC to promote hepatoma cell proliferation via down-regulating the tumor suppressor p18. HBx was also found to downregulate an lncRNA termed lncRNA-Dreh, which can inhibit HCC growth and metastasis in vitro and in vivo, and acts as a tumor suppressor in the development of HBV-HCC[89]. The downregulation of lncRNA-Dreh additionally correlated with poor survival of HCC patients.
LncRNA MALAT-1 (metastasis-associated lung adenocarcinoma transcript 1) was initially reported to be closely associated with non-small cell lung cancer (NSCLC) metastasis[90,91]. Likewise, lncRNA MALAT-1[92] and lncRNA HOTAIR (HOX antisense intergenic RNA)[93] have been shown to be upregulated in large cohorts of HCC patients. In particular, lncRNA HOTAIR was implicated as a prognostic biomarker for tumor recurrence after liver transplantation. Both studies demonstrated that siRNA-mediated reduction of lncRNA MALAT-1 and lncRNA HOTAIR suppressed cell viability and cell invasion, sensitized tumor necrosis factor (TNF)-α induced apoptosis, and increased the chemotherapeutic sensitivity of cancer cells to cisplatin and doxorubicin[93]. LncRNA MVIH (microvascular invasion in HCC) is elevated in HCC, and as its name suggests, was associated with microvascular invasion in HCC[94]. Another lncRNA overexpressed in HCC, lncRNA HEIH (high expression in HCC), was found to be significantly associated with recurrence and is an independent prognostic factor for survival[95]. Additionally, lncRNA-HEIH was found to physically interact with EZH2 - an interaction which is fundamental for repressing target genes such as p16 (lncRNA-HEIH can enhance the binding of EZH2 and H3K27me3 levels across the p16 promoters)[95].
Given the key regulatory functions of lncRNAs in cancers including HCC, and the evidence of dysregulated lncRNA expression in HCC, the targeting of lncRNAs offers a novel exciting opportunity to treat HCC. In principle, targeting of lncRNA can be achieved using the following approaches: (1) siRNA-mediated silencing; (2) functional block using small molecules or oligonucleotide inhibitors to prevent interactions of lncRNAs with proteins such as PRC2; and (3) structure disruption using small molecules or oligonucleotide inhibitors to change or mimic their secondary structure to compete for their binding partners. Since targeting lcnRNAs is still in its infancy, no investigational agents are currently available[96]. The complex epigenetic regulatory networks in HCC are summarized in Figure 1.
HCC is a heterogeneous disease whose management requires a multidisciplinary approach. One of the major clinical challenges is the inability to detect HCC at its early stages; patients are often diagnosed at advanced stages, which limits therapeutic options and leads to poor prognosis and unfavorable outcome[97]. Current treatment strategies for HCC include: (1) surgical removal of the tumor and liver transplantation; (2) minimal invasive surgery with application of radiofrequency ablation or cryoablation; and (3) chemoembolization by injecting drugs directly into the liver[98]. However, these approaches are limited by shortage of organ donors, small percentage of patients suitable for surgical removal, high post-operative recurrence rate, and underlying complications such as cirrhosis, HBV and HCV infections[99].
To date, only a number of molecularly-based therapeutics are available in the clinical management of HCC. In 2007, sorafenib (Nexavar) was approved by the United States Food and Drug Administration (FDA) for treatment of advanced primary HCC. This multi-tyrosine kinase inhibitor works by interfering with vascular endothelial growth factor signaling pathways in tumor angiogenesis. Clinical trials showed modest prolonged median survival and time to progression of 3 mo[100]. Similarly, in a cohort of patients from the Asia-Pacific region, the median overall survival increased from 4.2 to 6.5 mo[101]. Ongoing clinical trials are underway to test the efficacy of sorafenib in combination with other drugs[102]. Studies are also in progress for other drugs targeting the epidermal growth factor receptor, hepatocyte growth factor/c-Met, platelet-derived growth factor receptor, and mammalian target of rapamycin, all involved in molecular pathways of growth[103]. These drugs all show variable outcomes in the treatment of HCC.
In contrast to conventional or molecularly-targeted therapies for inhibiting dysregulated genes or signaling pathways, epigenetic drugs provide an alternative approach by reversing the methylation status and histone modifications of aberrantly expressed genes[104]. There are currently four FDA-approved epigenetic drugs, including two DNMT inhibitors, 5-azacytidine and decitabine, and two HDAC inhibitors, vorinostat and valporic acid. These drugs have been successful in treating hematological cancers, specifically myelodysplastic syndrome, a blood cancer characterized by inability to generate blood cells in the bone marrow[105]. Interestingly, low doses of 5-azacytidine and decitabine show anti-tumor effects on cultured and primary leukemia cells[106,107], as well as primary cells isolated from luminal-type breast cancer[106]. More importantly, they can reduce the number of CD34+ stem cells in leukemia, and mammosphere-forming breast cancer cells. Importantly, CD34+ cells are the origin and cause of tumor recurrence and chemoresistance[108].
Studies using cell lines[109] and pre-clinical mouse models[110] reveal promising results that may open up new avenues for the intervention and management of HCC. Drugs targeting epigenetic changes in HCC are summarized in Table 2. Andersen et al[110] showed that treatment with the DNMT inhibitor zebularine can prevent and effectively inhibit tumor growth in xenograft mouse models that are sensitive to this drug. Zebularine-resistant cell lines, however, showed up-regulation of oncogenic activities that instead promote liver cancer growth. These findings suggest that this drug may only benefit a specific sub-population of HCC patients. Treatment with zebularine was also shown to inhibit cell proliferation and to induce apoptosis of the HepG2 cell line. However, the DNA methylation levels of tumor suppressor genes p53 and p21 were not affected by zebularine treatment, while the anti-apoptotic protein BCL-2 is down-regulated, indicating that a DNA-methylation-independent pathway exists for increased p53 and p21 protein levels[109]. Additional cell-based studies show that 5-aza-2’-deoxycytidine exerts anti-tumor effects by inhibiting telomerase activity, accompanied by reactivation of p16 and c-Myc expression[111].
| Epigenetic modification | Targets | Drug(s) | Cell line/Animal model/Clinical trial phase | Results | Ref. |
| DNA methylation | DNA methyltransferase | Zebularine | Huh7 and KMCH cell linesHuman xenograft models | Zebularine-sensitive cell lines (Huh7 and KMCH) showed preferential demethylation of genes for tumor suppression, apoptosis, and cell cycle regulationIn vivo inhibition of tumor growth in xenograft model | [110] |
| DNA methyltransferases | Zebularine | HepG2 cell line | Zebularine treatment inhibited cell proliferation and induced apoptosis in HepG2 cell line | [109] | |
| DNA methyltransferases | 5-aza-2’-deoxycytidine | MMC-7721 and HepG2 cell lines | Inhibited telomerase activity, accompanied by reactivation of p16 and c-Myc. DAC synergized with cisplatin on growth inhibition | [111] | |
| Histone acetylation | Histone deacetylase | Belinostat | PLC/PRF/5, Hep3B and HepG2 cell lines | Inhibited cell growth and induced apoptosis | [112] |
| Histone deacetylase | Belinostat | Multi-center phase I/II clinical trial | Stabilized tumor in non-resectable advanced HCC | [115] | |
| Histone deacetylase | Suberoylanilide hydroxamic acid | HepG2, Hep3B and SK- Hep1 cell lines | Tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis | [114] | |
| Combination | Histone deacetylase + tyrosine kinase-inhibitors | Panobinostat + sorafinib | Huh7, Hep3B and HepG2 cell lines HCC xenograft model | Induction of apoptosisCombined panobinostat and sorafenib decreased vessel density, tumor volume and increased survival in HCC xenografts | [124] |
The use of HDAC inhibitors in HCC has been investigated in preclinical and clinical studies. In preclinical studies, belinostat inhibited cell growth in a HCC cell line[112], while suberoylanilide hydroxamic acid sensitized HCC cells to acetylation of p53[113] and TNF-related apoptosis-inducing ligand (TRAIL)-induced apoptosis[114]. In a multi-center phase I/II clinical trial, belinostat was found to stabilize unresectable advanced HCC[115]. Another important aspect of this clinical trial is the identification of HR23B as a potential biomarker for predicting the response to belinostat[115]. The combination of suberoylanilide hydroxamic and dihydroartemisinin significantly halted the growth of liver cancer tumor xenografts[116]. Of note, treatment with HDAC inhibitors was observed to induce cell death while simultaneously activating tumor-progression genes[117]. These observations indicate that a more in-depth understanding of epigenetic mechanisms is needed to obtain further insights into the in vivo determinants of responses to epigenetic drugs.
A recent addition to the family of epigenetic drugs is GSK-1, the first chemical inhibitor synthesized for targeting histone demethylases JMJD3/UTX (KDM6A/KDM6B) in a selective and potent manner[118]. These histone demethylases (JMJD3/UTX) catalyze removal of the trimethylation marker of H3K27. JMJD3 is induced via nuclear factor-κB exclusively in macrophages upon lipopolysaccharide (LPS) stimulation, providing an important link between inflammation and epigenetic reprogramming[119]. This small molecule specifically inhibits the LPS-induced pro-inflammatory response by sustaining the repressive effect of H3K27 on the TNF-α gene, as well as by inhibiting recruitment of RNA polymerase II during transcription, thereby reducing TNF-α expression[118]. Chronic inflammation due to chronic HBV and HCV infection, or obesity, gives rise to liver injury that slowly progresses to HCC, with elevated levels of pro-inflammatory cytokines TNF-α and interleukin (IL)-6 in HCC[120,121]. By suppressing TNF-α expression, GSK-1 may serve as a potential candidate drug for HCC.
MiRNAs with tumor suppressor roles in HCC, such as miR-26a, are feasible anti-cancer agents. Specifically, reduced expression levels of miR-26a have been reported in human HCCs[64], whereas systemic restoration of miR-26a expression by an adeno-associated virus suppresses cancer cell proliferation in a liver-specific Myc transgenic mouse model[122]. MiR-26a exerts these effects by directly targeting and down-regulating cyclins D2 and E2, consequently inducing cell cycle arrest. Importantly, as reported in that study, the cytotoxic effect of this miRNA is minimal in major organs. In another mouse model, in which liver cancer was induced by administration of the chemical hepatocarcinogen diethylnitrosamine, restoration of miR-124 by systemic injection significantly reduced liver tumor size[123]. Likewise, no cytotoxic effects on vital organs were detected.
Aberrant expression of several HDACs and copy number gains of HDAC3 and HDAC5 were detected in HCC patients[124], providing the rationale for treating HCC with HDAC inhibitors. In preclinical models of HCC, the combination of the pan-HDAC inhibitor panobinostat and sorafenib significantly decreased tumor growth and improved survival in murine xenograft models, compared to either drug used alone. Detailed molecular mechanisms include induction of apoptosis, acetylation of histone 3, down-regulation of BIRC5, or up-regulation of CDH1. This observation implies that such combination treatment may also achieve favorable clinical outcome for HCC patients.
Besides reversing aberrant epigenetic modifications in diseased conditions, epigenetic drugs can also induce host immunogenicity by increasing tumor antigen presentation. Systemic administration of the DNMT inhibitor 5-aza-2’-deoxycytidine induces the cancer/testis antigen (CTA) and augments adoptive immunotherapy by making cancer cells more visible to immunotherapy[125]. Under the inflammatory microenvironment of melanoma, melanocytes exhibit cell plasticity by converting between differentiated and undifferentiated status caused by TNF-α. Gradually, cells acquire resistance to adoptive cell transfer therapy[126]. The immunomodulatory activity of 5-aza-2’-deoxycytidine in vivo suggests its clinical use to design novel strategies of CTA-based chemo-immunotherapy for melanoma patients[127]. Such a concept may also be extended to the treatment of HCC, especially since HCC routinely occurs on a background of inflammation resulting from chronic HBV or HCV infection, where increased levels of IL-6[128] and TNF-α[121] are readily detected.
Like other types of cancers, HCC is associated with multiple genetic mutations and epigenetic aberrations. Whereas gene mutations are not easily amenable for therapy, epigenetic aberrations that appear frequently in HCC may serve as new targets[129]. To date, global DNA hypomethylation, promoter methylation, aberrant expression of miRNAs, and dysregulated expression of other epigenetic regulatory genes such as EZH2 are the best-known epigenetic abnormalities in HCC. Epigenetic drugs targeting these abnormalities may reverse the expression of dysregulated genes that are important in apoptosis and cell cycle arrest, thereby controlling the development and/or progression of HCC. Concerns of using epigenetic drugs are their off-target effects, which may activate genes having the opposite effects. Detailed mechanisms of action should be further investigated in multiple cancer types. Future research should aim at understanding how best to identify patient groups that would benefit most from the prescribed therapy, depending on the clinical setting, for example, disease stage, disease background, which drugs will give the best outcome, and when they should be administered. Useful biomarker(s) should provide guidance in identifying patients who may have optimal responses and reduced likelihood of relapse. Last but not least, most epigenetic drugs are used mainly in advanced tumors. Whether these drugs can be used as a preventative measure or in high risk patients who may benefit from future therapy remains to be explored.
P- Reviewers: Alpini G, Limpaiboon T, Naghibalhossaini F, Shang H S- Editor: Song XX L- Editor: Wang TQ E- Editor: Liu XM
| 1. | Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61:69-90. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 23762] [Cited by in F6Publishing: 25175] [Article Influence: 1936.5] [Reference Citation Analysis (3)] |
| 2. | El-Serag HB. Hepatocellular carcinoma. N Engl J Med. 2011;365:1118-1127. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2881] [Cited by in F6Publishing: 2977] [Article Influence: 229.0] [Reference Citation Analysis (0)] |
| 3. | Altekruse SF, McGlynn KA, Reichman ME. Hepatocellular carcinoma incidence, mortality, and survival trends in the United States from 1975 to 2005. J Clin Oncol. 2009;27:1485-1491. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1196] [Cited by in F6Publishing: 1285] [Article Influence: 85.7] [Reference Citation Analysis (0)] |
| 4. | El-Serag HB, Rudolph KL. Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology. 2007;132:2557-2576. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3846] [Cited by in F6Publishing: 4102] [Article Influence: 241.3] [Reference Citation Analysis (2)] |
| 5. | Egger G, Liang G, Aparicio A, Jones PA. Epigenetics in human disease and prospects for epigenetic therapy. Nature. 2004;429:457-463. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2210] [Cited by in F6Publishing: 2104] [Article Influence: 105.2] [Reference Citation Analysis (0)] |
| 6. | Jones PA, Takai D. The role of DNA methylation in mammalian epigenetics. Science. 2001;293:1068-1070. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1432] [Cited by in F6Publishing: 1352] [Article Influence: 58.8] [Reference Citation Analysis (0)] |
| 7. | Karpf AR, Jones DA. Reactivating the expression of methylation silenced genes in human cancer. Oncogene. 2002;21:5496-5503. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 198] [Cited by in F6Publishing: 209] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 8. | Das PM, Singal R. DNA methylation and cancer. J Clin Oncol. 2004;22:4632-4642. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 797] [Cited by in F6Publishing: 822] [Article Influence: 41.1] [Reference Citation Analysis (0)] |
| 9. | Herceg Z, Paliwal A. Epigenetic mechanisms in hepatocellular carcinoma: how environmental factors influence the epigenome. Mutat Res. 2011;727:55-61. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 131] [Cited by in F6Publishing: 118] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 10. | Hernandez-Vargas H, Lambert MP, Le Calvez-Kelm F, Gouysse G, McKay-Chopin S, Tavtigian SV, Scoazec JY, Herceg Z. Hepatocellular carcinoma displays distinct DNA methylation signatures with potential as clinical predictors. PLoS One. 2010;5:e9749. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 151] [Cited by in F6Publishing: 149] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 11. | Song MA, Tiirikainen M, Kwee S, Okimoto G, Yu H, Wong LL. Elucidating the landscape of aberrant DNA methylation in hepatocellular carcinoma. PLoS One. 2013;8:e55761. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 83] [Cited by in F6Publishing: 89] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 12. | Shen J, Wang S, Zhang YJ, Kappil M, Wu HC, Kibriya MG, Wang Q, Jasmine F, Ahsan H, Lee PH. Genome-wide DNA methylation profiles in hepatocellular carcinoma. Hepatology. 2012;55:1799-1808. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 150] [Cited by in F6Publishing: 151] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 13. | Nishida N, Kudo M, Nagasaka T, Ikai I, Goel A. Characteristic patterns of altered DNA methylation predict emergence of human hepatocellular carcinoma. Hepatology. 2012;56:994-1003. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 111] [Cited by in F6Publishing: 119] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 14. | Revill K, Wang T, Lachenmayer A, Kojima K, Harrington A, Li J, Hoshida Y, Llovet JM, Powers S. Genome-wide methylation analysis and epigenetic unmasking identify tumor suppressor genes in hepatocellular carcinoma. Gastroenterology. 2013;145:1424-1435.e25. [PubMed] [Cited in This Article: ] |
| 15. | Bhasin M, Reinherz EL, Reche PA. Recognition and classification of histones using support vector machine. J Comput Biol. 2006;13:102-112. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 58] [Cited by in F6Publishing: 64] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 16. | Peterson CL, Laniel MA. Histones and histone modifications. Curr Biol. 2004;14:R546-R551. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 887] [Cited by in F6Publishing: 841] [Article Influence: 42.1] [Reference Citation Analysis (0)] |
| 17. | Cedar H, Bergman Y. Linking DNA methylation and histone modification: patterns and paradigms. Nat Rev Genet. 2009;10:295-304. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1601] [Cited by in F6Publishing: 1563] [Article Influence: 104.2] [Reference Citation Analysis (0)] |
| 18. | Shahbazian MD, Grunstein M. Functions of site-specific histone acetylation and deacetylation. Annu Rev Biochem. 2007;76:75-100. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1112] [Cited by in F6Publishing: 1139] [Article Influence: 67.0] [Reference Citation Analysis (0)] |
| 19. | Lachner M, Jenuwein T. The many faces of histone lysine methylation. Curr Opin Cell Biol. 2002;14:286-298. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 610] [Cited by in F6Publishing: 642] [Article Influence: 29.2] [Reference Citation Analysis (0)] |
| 20. | Zhang Y, Reinberg D. Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev. 2001;15:2343-2360. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1142] [Cited by in F6Publishing: 1086] [Article Influence: 47.2] [Reference Citation Analysis (0)] |
| 21. | He C, Xu J, Zhang J, Xie D, Ye H, Xiao Z, Cai M, Xu K, Zeng Y, Li H. High expression of trimethylated histone H3 lysine 4 is associated with poor prognosis in hepatocellular carcinoma. Hum Pathol. 2012;43:1425-1435. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 83] [Cited by in F6Publishing: 83] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 22. | Cai MY, Hou JH, Rao HL, Luo RZ, Li M, Pei XQ, Lin MC, Guan XY, Kung HF, Zeng YX. High expression of H3K27me3 in human hepatocellular carcinomas correlates closely with vascular invasion and predicts worse prognosis in patients. Mol Med. 2011;17:12-20. [PubMed] [Cited in This Article: ] |
| 23. | Spivakov M, Fisher AG. Epigenetic signatures of stem-cell identity. Nat Rev Genet. 2007;8:263-271. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 287] [Cited by in F6Publishing: 302] [Article Influence: 17.8] [Reference Citation Analysis (0)] |
| 24. | Margueron R, Reinberg D. The Polycomb complex PRC2 and its mark in life. Nature. 2011;469:343-349. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2234] [Cited by in F6Publishing: 2321] [Article Influence: 178.5] [Reference Citation Analysis (0)] |
| 25. | Pasini D, Bracken AP, Hansen JB, Capillo M, Helin K. The polycomb group protein Suz12 is required for embryonic stem cell differentiation. Mol Cell Biol. 2007;27:3769-3779. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 535] [Cited by in F6Publishing: 532] [Article Influence: 31.3] [Reference Citation Analysis (0)] |
| 26. | Kleer CG, Cao Q, Varambally S, Shen R, Ota I, Tomlins SA, Ghosh D, Sewalt RG, Otte AP, Hayes DF. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc Natl Acad Sci USA. 2003;100:11606-11611. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1203] [Cited by in F6Publishing: 1257] [Article Influence: 59.9] [Reference Citation Analysis (0)] |
| 27. | Collett K, Eide GE, Arnes J, Stefansson IM, Eide J, Braaten A, Aas T, Otte AP, Akslen LA. Expression of enhancer of zeste homologue 2 is significantly associated with increased tumor cell proliferation and is a marker of aggressive breast cancer. Clin Cancer Res. 2006;12:1168-1174. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 203] [Cited by in F6Publishing: 208] [Article Influence: 11.6] [Reference Citation Analysis (0)] |
| 28. | Varambally S, Dhanasekaran SM, Zhou M, Barrette TR, Kumar-Sinha C, Sanda MG, Ghosh D, Pienta KJ, Sewalt RG, Otte AP. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature. 2002;419:624-629. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1984] [Cited by in F6Publishing: 1982] [Article Influence: 90.1] [Reference Citation Analysis (0)] |
| 29. | Sudo T, Utsunomiya T, Mimori K, Nagahara H, Ogawa K, Inoue H, Wakiyama S, Fujita H, Shirouzu K, Mori M. Clinicopathological significance of EZH2 mRNA expression in patients with hepatocellular carcinoma. Br J Cancer. 2005;92:1754-1758. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 170] [Cited by in F6Publishing: 183] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 30. | Cai MY, Tong ZT, Zheng F, Liao YJ, Wang Y, Rao HL, Chen YC, Wu QL, Liu YH, Guan XY. EZH2 protein: a promising immunomarker for the detection of hepatocellular carcinomas in liver needle biopsies. Gut. 2011;60:967-976. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 130] [Cited by in F6Publishing: 129] [Article Influence: 9.9] [Reference Citation Analysis (0)] |
| 31. | Yonemitsu Y, Imazeki F, Chiba T, Fukai K, Nagai Y, Miyagi S, Arai M, Aoki R, Miyazaki M, Nakatani Y. Distinct expression of polycomb group proteins EZH2 and BMI1 in hepatocellular carcinoma. Hum Pathol. 2009;40:1304-1311. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 42] [Cited by in F6Publishing: 39] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 32. | Cheng AS, Lau SS, Chen Y, Kondo Y, Li MS, Feng H, Ching AK, Cheung KF, Wong HK, Tong JH. EZH2-mediated concordant repression of Wnt antagonists promotes β-catenin-dependent hepatocarcinogenesis. Cancer Res. 2011;71:4028-4039. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 155] [Cited by in F6Publishing: 172] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 33. | Au SL, Wong CC, Lee JM, Fan DN, Tsang FH, Ng IO, Wong CM. Enhancer of zeste homolog 2 epigenetically silences multiple tumor suppressor microRNAs to promote liver cancer metastasis. Hepatology. 2012;56:622-631. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 209] [Cited by in F6Publishing: 228] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 34. | Aoki R, Chiba T, Miyagi S, Negishi M, Konuma T, Taniguchi H, Ogawa M, Yokosuka O, Iwama A. The polycomb group gene product Ezh2 regulates proliferation and differentiation of murine hepatic stem/progenitor cells. J Hepatol. 2010;52:854-863. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 57] [Cited by in F6Publishing: 61] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 35. | Wang L, Zhang X, Jia LT, Hu SJ, Zhao J, Yang JD, Wen WH, Wang Z, Wang T, Zhao J. c-Myc-mediated epigenetic silencing of microRNA-101 contributes to dysregulation of multiple pathways in hepatocellular carcinoma. Hepatology. 2013;Epub ahead of print. [PubMed] [Cited in This Article: ] |
| 36. | Varambally S, Cao Q, Mani RS, Shankar S, Wang X, Ateeq B, Laxman B, Cao X, Jing X, Ramnarayanan K. Genomic loss of microRNA-101 leads to overexpression of histone methyltransferase EZH2 in cancer. Science. 2008;322:1695-1699. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 808] [Cited by in F6Publishing: 812] [Article Influence: 50.8] [Reference Citation Analysis (0)] |
| 37. | Terradillos O, Billet O, Renard CA, Levy R, Molina T, Briand P, Buendia MA. The hepatitis B virus X gene potentiates c-myc-induced liver oncogenesis in transgenic mice. Oncogene. 1997;14:395-404. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 224] [Cited by in F6Publishing: 225] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 38. | Madden CR, Finegold MJ, Slagle BL. Hepatitis B virus X protein acts as a tumor promoter in development of diethylnitrosamine-induced preneoplastic lesions. J Virol. 2001;75:3851-3858. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 95] [Cited by in F6Publishing: 108] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 39. | Rakotomalala L, Studach L, Wang WH, Gregori G, Hullinger RL, Andrisani O. Hepatitis B virus X protein increases the Cdt1-to-geminin ratio inducing DNA re-replication and polyploidy. J Biol Chem. 2008;283:28729-28740. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 43] [Cited by in F6Publishing: 47] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 40. | Studach L, Wang WH, Weber G, Tang J, Hullinger RL, Malbrue R, Liu X, Andrisani O. Polo-like kinase 1 activated by the hepatitis B virus X protein attenuates both the DNA damage checkpoint and DNA repair resulting in partial polyploidy. J Biol Chem. 2010;285:30282-30293. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 38] [Cited by in F6Publishing: 41] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 41. | Studach LL, Menne S, Cairo S, Buendia MA, Hullinger RL, Lefrançois L, Merle P, Andrisani OM. Subset of Suz12/PRC2 target genes is activated during hepatitis B virus replication and liver carcinogenesis associated with HBV X protein. Hepatology. 2012;56:1240-1251. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 35] [Cited by in F6Publishing: 39] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 42. | Wang WH, Studach LL, Andrisani OM. Proteins ZNF198 and SUZ12 are down-regulated in hepatitis B virus (HBV) X protein-mediated hepatocyte transformation and in HBV replication. Hepatology. 2011;53:1137-1147. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 46] [Cited by in F6Publishing: 48] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 43. | Carthew RW, Sontheimer EJ. Origins and Mechanisms of miRNAs and siRNAs. Cell. 2009;136:642-655. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3447] [Cited by in F6Publishing: 3500] [Article Influence: 233.3] [Reference Citation Analysis (0)] |
| 44. | Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281-297. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 25833] [Cited by in F6Publishing: 26770] [Article Influence: 1338.5] [Reference Citation Analysis (0)] |
| 45. | Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857-866. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 5705] [Cited by in F6Publishing: 5849] [Article Influence: 324.9] [Reference Citation Analysis (0)] |
| 46. | Pedersen IM, Cheng G, Wieland S, Volinia S, Croce CM, Chisari FV, David M. Interferon modulation of cellular microRNAs as an antiviral mechanism. Nature. 2007;449:919-922. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 686] [Cited by in F6Publishing: 727] [Article Influence: 42.8] [Reference Citation Analysis (0)] |
| 47. | Lecellier CH, Dunoyer P, Arar K, Lehmann-Che J, Eyquem S, Himber C, Saïb A, Voinnet O. A cellular microRNA mediates antiviral defense in human cells. Science. 2005;308:557-560. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 686] [Cited by in F6Publishing: 685] [Article Influence: 36.1] [Reference Citation Analysis (0)] |
| 48. | Hu W, Wang X, Ding X, Li Y, Zhang X, Xie P, Yang J, Wang S. MicroRNA-141 represses HBV replication by targeting PPARA. PLoS One. 2012;7:e34165. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 71] [Cited by in F6Publishing: 82] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 49. | Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, Tuschl T. Identification of tissue-specific microRNAs from mouse. Curr Biol. 2002;12:735-739. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2502] [Cited by in F6Publishing: 2504] [Article Influence: 113.8] [Reference Citation Analysis (0)] |
| 50. | Chen Y, Shen A, Rider PJ, Yu Y, Wu K, Mu Y, Hao Q, Liu Y, Gong H, Zhu Y. A liver-specific microRNA binds to a highly conserved RNA sequence of hepatitis B virus and negatively regulates viral gene expression and replication. FASEB J. 2011;25:4511-4521. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 129] [Cited by in F6Publishing: 141] [Article Influence: 10.8] [Reference Citation Analysis (0)] |
| 51. | Shimakami T, Yamane D, Jangra RK, Kempf BJ, Spaniel C, Barton DJ, Lemon SM. Stabilization of hepatitis C virus RNA by an Ago2-miR-122 complex. Proc Natl Acad Sci USA. 2012;109:941-946. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 291] [Cited by in F6Publishing: 287] [Article Influence: 23.9] [Reference Citation Analysis (0)] |
| 52. | Janssen HL, Reesink HW, Lawitz EJ, Zeuzem S, Rodriguez-Torres M, Patel K, van der Meer AJ, Patick AK, Chen A, Zhou Y. Treatment of HCV infection by targeting microRNA. N Engl J Med. 2013;368:1685-1694. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1665] [Cited by in F6Publishing: 1619] [Article Influence: 147.2] [Reference Citation Analysis (0)] |
| 53. | Kutay H, Bai S, Datta J, Motiwala T, Pogribny I, Frankel W, Jacob ST, Ghoshal K. Downregulation of miR-122 in the rodent and human hepatocellular carcinomas. J Cell Biochem. 2006;99:671-678. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 450] [Cited by in F6Publishing: 492] [Article Influence: 28.9] [Reference Citation Analysis (0)] |
| 54. | Murakami Y, Yasuda T, Saigo K, Urashima T, Toyoda H, Okanoue T, Shimotohno K. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene. 2006;25:2537-2545. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 876] [Cited by in F6Publishing: 880] [Article Influence: 48.9] [Reference Citation Analysis (0)] |
| 55. | Tsai WC, Hsu PW, Lai TC, Chau GY, Lin CW, Chen CM, Lin CD, Liao YL, Wang JL, Chau YP. MicroRNA-122, a tumor suppressor microRNA that regulates intrahepatic metastasis of hepatocellular carcinoma. Hepatology. 2009;49:1571-1582. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 450] [Cited by in F6Publishing: 465] [Article Influence: 31.0] [Reference Citation Analysis (0)] |
| 56. | Zhang X, Zhang E, Ma Z, Pei R, Jiang M, Schlaak JF, Roggendorf M, Lu M. Modulation of hepatitis B virus replication and hepatocyte differentiation by MicroRNA-1. Hepatology. 2011;53:1476-1485. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 149] [Cited by in F6Publishing: 154] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 57. | Xu X, Fan Z, Kang L, Han J, Jiang C, Zheng X, Zhu Z, Jiao H, Lin J, Jiang K. Hepatitis B virus X protein represses miRNA-148a to enhance tumorigenesis. J Clin Invest. 2013;123:630-645. [PubMed] [Cited in This Article: ] |
| 58. | Wang XW, Heegaard NH, Orum H. MicroRNAs in liver disease. Gastroenterology. 2012;142:1431-1443. [PubMed] [Cited in This Article: ] |
| 59. | Callegari E, Elamin BK, Sabbioni S, Gramantieri L, Negrini M. Role of microRNAs in hepatocellular carcinoma: a clinical perspective. Onco Targets Ther. 2013;6:1167-1178. [PubMed] [Cited in This Article: ] |
| 60. | Wang W, Zhao LJ, Tan YX, Ren H, Qi ZT. Identification of deregulated miRNAs and their targets in hepatitis B virus-associated hepatocellular carcinoma. World J Gastroenterol. 2012;18:5442-5453. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 75] [Cited by in F6Publishing: 83] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 61. | Wang W, Zhao LJ, Tan YX, Ren H, Qi ZT. MiR-138 induces cell cycle arrest by targeting cyclin D3 in hepatocellular carcinoma. Carcinogenesis. 2012;33:1113-1120. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 163] [Cited by in F6Publishing: 176] [Article Influence: 14.7] [Reference Citation Analysis (0)] |
| 62. | Shen Q, Cicinnati VR, Zhang X, Iacob S, Weber F, Sotiropoulos GC, Radtke A, Lu M, Paul A, Gerken G. Role of microRNA-199a-5p and discoidin domain receptor 1 in human hepatocellular carcinoma invasion. Mol Cancer. 2010;9:227. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 132] [Cited by in F6Publishing: 154] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 63. | Jiang J, Gusev Y, Aderca I, Mettler TA, Nagorney DM, Brackett DJ, Roberts LR, Schmittgen TD. Association of MicroRNA expression in hepatocellular carcinomas with hepatitis infection, cirrhosis, and patient survival. Clin Cancer Res. 2008;14:419-427. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 387] [Cited by in F6Publishing: 414] [Article Influence: 25.9] [Reference Citation Analysis (0)] |
| 64. | Ji J, Shi J, Budhu A, Yu Z, Forgues M, Roessler S, Ambs S, Chen Y, Meltzer PS, Croce CM. MicroRNA expression, survival, and response to interferon in liver cancer. N Engl J Med. 2009;361:1437-1447. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 624] [Cited by in F6Publishing: 638] [Article Influence: 42.5] [Reference Citation Analysis (0)] |
| 65. | Liang L, Wong CM, Ying Q, Fan DN, Huang S, Ding J, Yao J, Yan M, Li J, Yao M. MicroRNA-125b suppressesed human liver cancer cell proliferation and metastasis by directly targeting oncogene LIN28B2. Hepatology. 2010;52:1731-1740. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 190] [Cited by in F6Publishing: 208] [Article Influence: 14.9] [Reference Citation Analysis (0)] |
| 66. | Yang H, Fang F, Chang R, Yang L. MicroRNA-140-5p suppresses tumor growth and metastasis by targeting transforming growth factor β receptor 1 and fibroblast growth factor 9 in hepatocellular carcinoma. Hepatology. 2013;58:205-217. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 199] [Cited by in F6Publishing: 217] [Article Influence: 19.7] [Reference Citation Analysis (0)] |
| 67. | Wei W, Hou J, Alder O, Ye X, Lee S, Cullum R, Chu A, Zhao Y, Warner SM, Knight DA. Genome-wide microRNA and messenger RNA profiling in rodent liver development implicates mir302b and mir20a in repressing transforming growth factor-beta signaling. Hepatology. 2013;57:2491-2501. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 17] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 68. | Tsai WC, Hsu SD, Hsu CS, Lai TC, Chen SJ, Shen R, Huang Y, Chen HC, Lee CH, Tsai TF. MicroRNA-122 plays a critical role in liver homeostasis and hepatocarcinogenesis. J Clin Invest. 2012;122:2884-2897. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 612] [Cited by in F6Publishing: 631] [Article Influence: 52.6] [Reference Citation Analysis (0)] |
| 69. | Coulouarn C, Factor VM, Andersen JB, Durkin ME, Thorgeirsson SS. Loss of miR-122 expression in liver cancer correlates with suppression of the hepatic phenotype and gain of metastatic properties. Oncogene. 2009;28:3526-3536. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 531] [Cited by in F6Publishing: 565] [Article Influence: 37.7] [Reference Citation Analysis (0)] |
| 70. | Sell S, Leffert HL. Liver cancer stem cells. J Clin Oncol. 2008;26:2800-2805. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 170] [Cited by in F6Publishing: 162] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 71. | Rountree CB, Mishra L, Willenbring H. Stem cells in liver diseases and cancer: recent advances on the path to new therapies. Hepatology. 2012;55:298-306. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 77] [Cited by in F6Publishing: 82] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 72. | Yamashita T, Ji J, Budhu A, Forgues M, Yang W, Wang HY, Jia H, Ye Q, Qin LX, Wauthier E. EpCAM-positive hepatocellular carcinoma cells are tumor-initiating cells with stem/progenitor cell features. Gastroenterology. 2009;136:1012-1024. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 891] [Cited by in F6Publishing: 905] [Article Influence: 60.3] [Reference Citation Analysis (0)] |
| 73. | Munz M, Baeuerle PA, Gires O. The emerging role of EpCAM in cancer and stem cell signaling. Cancer Res. 2009;69:5627-5629. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 375] [Cited by in F6Publishing: 404] [Article Influence: 26.9] [Reference Citation Analysis (0)] |
| 74. | Ma S, Chan KW, Hu L, Lee TK, Wo JY, Ng IO, Zheng BJ, Guan XY. Identification and characterization of tumorigenic liver cancer stem/progenitor cells. Gastroenterology. 2007;132:2542-2556. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 899] [Cited by in F6Publishing: 896] [Article Influence: 52.7] [Reference Citation Analysis (0)] |
| 75. | Suetsugu A, Nagaki M, Aoki H, Motohashi T, Kunisada T, Moriwaki H. Characterization of CD133+ hepatocellular carcinoma cells as cancer stem/progenitor cells. Biochem Biophys Res Commun. 2006;351:820-824. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 423] [Cited by in F6Publishing: 458] [Article Influence: 25.4] [Reference Citation Analysis (0)] |
| 76. | Yang ZF, Ho DW, Ng MN, Lau CK, Yu WC, Ngai P, Chu PW, Lam CT, Poon RT, Fan ST. Significance of CD90+ cancer stem cells in human liver cancer. Cancer Cell. 2008;13:153-166. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 883] [Cited by in F6Publishing: 897] [Article Influence: 56.1] [Reference Citation Analysis (0)] |
| 77. | Yang ZF, Ngai P, Ho DW, Yu WC, Ng MN, Lau CK, Li ML, Tam KH, Lam CT, Poon RT. Identification of local and circulating cancer stem cells in human liver cancer. Hepatology. 2008;47:919-928. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 263] [Cited by in F6Publishing: 292] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 78. | Ji J, Yamashita T, Budhu A, Forgues M, Jia HL, Li C, Deng C, Wauthier E, Reid LM, Ye QH. Identification of microRNA-181 by genome-wide screening as a critical player in EpCAM-positive hepatic cancer stem cells. Hepatology. 2009;50:472-480. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 397] [Cited by in F6Publishing: 436] [Article Influence: 29.1] [Reference Citation Analysis (0)] |
| 79. | Ji J, Yamashita T, Wang XW. Wnt/beta-catenin signaling activates microRNA-181 expression in hepatocellular carcinoma. Cell Biosci. 2011;1:4. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 77] [Cited by in F6Publishing: 87] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 80. | Zhang J, Luo N, Luo Y, Peng Z, Zhang T, Li S. microRNA-150 inhibits human CD133-positive liver cancer stem cells through negative regulation of the transcription factor c-Myb. Int J Oncol. 2012;40:747-756. [PubMed] [Cited in This Article: ] |
| 81. | Fang L, Zhang HB, Li H, Fu Y, Yang GS. miR-548c-5p inhibits proliferation and migration and promotes apoptosis in CD90(+) HepG2 cells. Radiol Oncol. 2012;46:233-241. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 18] [Cited by in F6Publishing: 20] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 82. | Cheetham SW, Gruhl F, Mattick JS, Dinger ME. Long noncoding RNAs and the genetics of cancer. Br J Cancer. 2013;108:2419-2425. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 582] [Cited by in F6Publishing: 596] [Article Influence: 54.2] [Reference Citation Analysis (0)] |
| 83. | Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011;1:391-407. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1266] [Cited by in F6Publishing: 1429] [Article Influence: 119.1] [Reference Citation Analysis (0)] |
| 84. | Khalil AM, Guttman M, Huarte M, Garber M, Raj A, Rivea Morales D, Thomas K, Presser A, Bernstein BE, van Oudenaarden A. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci USA. 2009;106:11667-11672. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2182] [Cited by in F6Publishing: 2276] [Article Influence: 151.7] [Reference Citation Analysis (0)] |
| 85. | Redon S, Reichenbach P, Lingner J. The non-coding RNA TERRA is a natural ligand and direct inhibitor of human telomerase. Nucleic Acids Res. 2010;38:5797-5806. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 257] [Cited by in F6Publishing: 266] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 86. | Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature. 2010;465:1033-1038. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1717] [Cited by in F6Publishing: 1801] [Article Influence: 128.6] [Reference Citation Analysis (0)] |
| 87. | Panzitt K, Tschernatsch MM, Guelly C, Moustafa T, Stradner M, Strohmaier HM, Buck CR, Denk H, Schroeder R, Trauner M. Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology. 2007;132:330-342. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 587] [Cited by in F6Publishing: 613] [Article Influence: 36.1] [Reference Citation Analysis (0)] |
| 88. | Du Y, Kong G, You X, Zhang S, Zhang T, Gao Y, Ye L, Zhang X. Elevation of highly up-regulated in liver cancer (HULC) by hepatitis B virus X protein promotes hepatoma cell proliferation via down-regulating p18. J Biol Chem. 2012;287:26302-26311. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 274] [Cited by in F6Publishing: 306] [Article Influence: 25.5] [Reference Citation Analysis (0)] |
| 89. | Huang JF, Guo YJ, Zhao CX, Yuan SX, Wang Y, Tang GN, Zhou WP, Sun SH. Hepatitis B virus X protein (HBx)-related long noncoding RNA (lncRNA) down-regulated expression by HBx (Dreh) inhibits hepatocellular carcinoma metastasis by targeting the intermediate filament protein vimentin. Hepatology. 2013;57:1882-1892. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 251] [Cited by in F6Publishing: 268] [Article Influence: 24.4] [Reference Citation Analysis (0)] |
| 90. | Ji P, Diederichs S, Wang W, Böing S, Metzger R, Schneider PM, Tidow N, Brandt B, Buerger H, Bulk E. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene. 2003;22:8031-8041. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1584] [Cited by in F6Publishing: 1716] [Article Influence: 81.7] [Reference Citation Analysis (0)] |
| 91. | Gutschner T, Hämmerle M, Eissmann M, Hsu J, Kim Y, Hung G, Revenko A, Arun G, Stentrup M, Gross M. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013;73:1180-1189. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1087] [Cited by in F6Publishing: 1230] [Article Influence: 102.5] [Reference Citation Analysis (0)] |
| 92. | Lai MC, Yang Z, Zhou L, Zhu QQ, Xie HY, Zhang F, Wu LM, Chen LM, Zheng SS. Long non-coding RNA MALAT-1 overexpression predicts tumor recurrence of hepatocellular carcinoma after liver transplantation. Med Oncol. 2012;29:1810-1816. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 394] [Cited by in F6Publishing: 464] [Article Influence: 35.7] [Reference Citation Analysis (0)] |
| 93. | Yang Z, Zhou L, Wu LM, Lai MC, Xie HY, Zhang F, Zheng SS. Overexpression of long non-coding RNA HOTAIR predicts tumor recurrence in hepatocellular carcinoma patients following liver transplantation. Ann Surg Oncol. 2011;18:1243-1250. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 530] [Cited by in F6Publishing: 595] [Article Influence: 45.8] [Reference Citation Analysis (0)] |
| 94. | Yuan SX, Yang F, Yang Y, Tao QF, Zhang J, Huang G, Yang Y, Wang RY, Yang S, Huo XS. Long noncoding RNA associated with microvascular invasion in hepatocellular carcinoma promotes angiogenesis and serves as a predictor for hepatocellular carcinoma patients’ poor recurrence-free survival after hepatectomy. Hepatology. 2012;56:2231-2241. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 285] [Cited by in F6Publishing: 322] [Article Influence: 26.8] [Reference Citation Analysis (0)] |
| 95. | Yang F, Zhang L, Huo XS, Yuan JH, Xu D, Yuan SX, Zhu N, Zhou WP, Yang GS, Wang YZ. Long noncoding RNA high expression in hepatocellular carcinoma facilitates tumor growth through enhancer of zeste homolog 2 in humans. Hepatology. 2011;54:1679-1689. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 482] [Cited by in F6Publishing: 535] [Article Influence: 41.2] [Reference Citation Analysis (0)] |
| 96. | Sánchez Y, Huarte M. Long non-coding RNAs: challenges for diagnosis and therapies. Nucleic Acid Ther. 2013;23:15-20. [PubMed] [Cited in This Article: ] |
| 97. | Gish RG. Hepatocellular carcinoma: overcoming challenges in disease management. Clin Gastroenterol Hepatol. 2006;4:252-261. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 43] [Cited by in F6Publishing: 46] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 98. | Maluccio M, Covey A. Recent progress in understanding, diagnosing, and treating hepatocellular carcinoma. CA Cancer J Clin. 2012;62:394-399. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 577] [Cited by in F6Publishing: 666] [Article Influence: 55.5] [Reference Citation Analysis (0)] |
| 99. | Tsim NC, Frampton AE, Habib NA, Jiao LR. Surgical treatment for liver cancer. World J Gastroenterol. 2010;16:927-933. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 40] [Cited by in F6Publishing: 42] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 100. | Llovet JM, Ricci S, Mazzaferro V, Hilgard P, Gane E, Blanc JF, de Oliveira AC, Santoro A, Raoul JL, Forner A. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359:378-390. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 9016] [Cited by in F6Publishing: 9500] [Article Influence: 593.8] [Reference Citation Analysis (1)] |
| 101. | Cheng AL, Kang YK, Chen Z, Tsao CJ, Qin S, Kim JS, Luo R, Feng J, Ye S, Yang TS. Efficacy and safety of sorafenib in patients in the Asia-Pacific region with advanced hepatocellular carcinoma: a phase III randomised, double-blind, placebo-controlled trial. Lancet Oncol. 2009;10:25-34. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3854] [Cited by in F6Publishing: 4336] [Article Influence: 271.0] [Reference Citation Analysis (0)] |
| 102. | Kelley RK, Nimeiri HS, Munster PN, Vergo MT, Huang Y, Li CM, Hwang J, Mulcahy MF, Yeh BM, Kuhn P. Temsirolimus combined with sorafenib in hepatocellular carcinoma: a phase I dose-finding trial with pharmacokinetic and biomarker correlates. Ann Oncol. 2013;24:1900-1907. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 43] [Cited by in F6Publishing: 45] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 103. | Chan SL, Yeo W. Targeted therapy of hepatocellular carcinoma: present and future. J Gastroenterol Hepatol. 2012;27:862-872. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 59] [Cited by in F6Publishing: 66] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 104. | Yoo CB, Jones PA. Epigenetic therapy of cancer: past, present and future. Nat Rev Drug Discov. 2006;5:37-50. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 952] [Cited by in F6Publishing: 918] [Article Influence: 51.0] [Reference Citation Analysis (0)] |
| 105. | Fenaux P, Mufti GJ, Hellstrom-Lindberg E, Santini V, Finelli C, Giagounidis A, Schoch R, Gattermann N, Sanz G, List A. Efficacy of azacitidine compared with that of conventional care regimens in the treatment of higher-risk myelodysplastic syndromes: a randomised, open-label, phase III study. Lancet Oncol. 2009;10:223-232. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1862] [Cited by in F6Publishing: 1945] [Article Influence: 129.7] [Reference Citation Analysis (0)] |
| 106. | Issa JP, Garcia-Manero G, Giles FJ, Mannari R, Thomas D, Faderl S, Bayar E, Lyons J, Rosenfeld CS, Cortes J. Phase 1 study of low-dose prolonged exposure schedules of the hypomethylating agent 5-aza-2’-deoxycytidine (decitabine) in hematopoietic malignancies. Blood. 2004;103:1635-1640. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 624] [Cited by in F6Publishing: 607] [Article Influence: 28.9] [Reference Citation Analysis (0)] |
| 107. | Issa JP, Gharibyan V, Cortes J, Jelinek J, Morris G, Verstovsek S, Talpaz M, Garcia-Manero G, Kantarjian HM. Phase II study of low-dose decitabine in patients with chronic myelogenous leukemia resistant to imatinib mesylate. J Clin Oncol. 2005;23:3948-3956. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 231] [Cited by in F6Publishing: 246] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 108. | Tsai HC, Li H, Van Neste L, Cai Y, Robert C, Rassool FV, Shin JJ, Harbom KM, Beaty R, Pappou E. Transient low doses of DNA-demethylating agents exert durable antitumor effects on hematological and epithelial tumor cells. Cancer Cell. 2012;21:430-446. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 477] [Cited by in F6Publishing: 465] [Article Influence: 38.8] [Reference Citation Analysis (0)] |
| 109. | Nakamura K, Aizawa K, Nakabayashi K, Kato N, Yamauchi J, Hata K, Tanoue A. DNA methyltransferase inhibitor zebularine inhibits human hepatic carcinoma cells proliferation and induces apoptosis. PLoS One. 2013;8:e54036. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 26] [Cited by in F6Publishing: 33] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 110. | Andersen JB, Factor VM, Marquardt JU, Raggi C, Lee YH, Seo D, Conner EA, Thorgeirsson SS. An integrated genomic and epigenomic approach predicts therapeutic response to zebularine in human liver cancer. Sci Transl Med. 2010;2:54ra77. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 66] [Cited by in F6Publishing: 80] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 111. | Tao SF, Zhang CS, Guo XL, Xu Y, Zhang SS, Song JR, Li R, Wu MC, Wei LX. Anti-tumor effect of 5-aza-2’-deoxycytidine by inhibiting telomerase activity in hepatocellular carcinoma cells. World J Gastroenterol. 2012;18:2334-2343. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 20] [Cited by in F6Publishing: 28] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 112. | Ma BB, Sung F, Tao Q, Poon FF, Lui VW, Yeo W, Chan SL, Chan AT. The preclinical activity of the histone deacetylase inhibitor PXD101 (belinostat) in hepatocellular carcinoma cell lines. Invest New Drugs. 2010;28:107-114. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 44] [Cited by in F6Publishing: 52] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 113. | Carlisi D, Vassallo B, Lauricella M, Emanuele S, D’Anneo A, Di Leonardo E, Di Fazio P, Vento R, Tesoriere G. Histone deacetylase inhibitors induce in human hepatoma HepG2 cells acetylation of p53 and histones in correlation with apoptotic effects. Int J Oncol. 2008;32:177-184. [PubMed] [Cited in This Article: ] |
| 114. | Carlisi D, Lauricella M, D’Anneo A, Emanuele S, Angileri L, Di Fazio P, Santulli A, Vento R, Tesoriere G. The histone deacetylase inhibitor suberoylanilide hydroxamic acid sensitises human hepatocellular carcinoma cells to TRAIL-induced apoptosis by TRAIL-DISC activation. Eur J Cancer. 2009;45:2425-2438. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 64] [Cited by in F6Publishing: 63] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 115. | Yeo W, Chung HC, Chan SL, Wang LZ, Lim R, Picus J, Boyer M, Mo FK, Koh J, Rha SY. Epigenetic therapy using belinostat for patients with unresectable hepatocellular carcinoma: a multicenter phase I/II study with biomarker and pharmacokinetic analysis of tumors from patients in the Mayo Phase II Consortium and the Cancer Therapeutics Research Group. J Clin Oncol. 2012;30:3361-3367. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 140] [Cited by in F6Publishing: 151] [Article Influence: 12.6] [Reference Citation Analysis (0)] |
| 116. | Zhang CZ, Pan Y, Cao Y, Lai PB, Liu L, Chen GG, Yun J. Histone deacetylase inhibitors facilitate dihydroartemisinin-induced apoptosis in liver cancer in vitro and in vivo. PLoS One. 2012;7:e39870. [PubMed] [Cited in This Article: ] |
| 117. | Lin KT, Wang YW, Chen CT, Ho CM, Su WH, Jou YS. HDAC inhibitors augmented cell migration and metastasis through induction of PKCs leading to identification of low toxicity modalities for combination cancer therapy. Clin Cancer Res. 2012;18:4691-4701. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 64] [Cited by in F6Publishing: 69] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 118. | Kruidenier L, Chung CW, Cheng Z, Liddle J, Che K, Joberty G, Bantscheff M, Bountra C, Bridges A, Diallo H. A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature. 2012;488:404-408. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 658] [Cited by in F6Publishing: 694] [Article Influence: 57.8] [Reference Citation Analysis (0)] |
| 119. | De Santa F, Totaro MG, Prosperini E, Notarbartolo S, Testa G, Natoli G. The histone H3 lysine-27 demethylase Jmjd3 links inflammation to inhibition of polycomb-mediated gene silencing. Cell. 2007;130:1083-1094. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 696] [Cited by in F6Publishing: 726] [Article Influence: 42.7] [Reference Citation Analysis (0)] |
| 120. | Naugler WE, Sakurai T, Kim S, Maeda S, Kim K, Elsharkawy AM, Karin M. Gender disparity in liver cancer due to sex differences in MyD88-dependent IL-6 production. Science. 2007;317:121-124. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1362] [Cited by in F6Publishing: 1400] [Article Influence: 82.4] [Reference Citation Analysis (0)] |
| 121. | Park EJ, Lee JH, Yu GY, He G, Ali SR, Holzer RG, Osterreicher CH, Takahashi H, Karin M. Dietary and genetic obesity promote liver inflammation and tumorigenesis by enhancing IL-6 and TNF expression. Cell. 2010;140:197-208. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1169] [Cited by in F6Publishing: 1283] [Article Influence: 91.6] [Reference Citation Analysis (1)] |
| 122. | Kota J, Chivukula RR, O’Donnell KA, Wentzel EA, Montgomery CL, Hwang HW, Chang TC, Vivekanandan P, Torbenson M, Clark KR. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell. 2009;137:1005-1017. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1287] [Cited by in F6Publishing: 1327] [Article Influence: 88.5] [Reference Citation Analysis (0)] |
| 123. | Hatziapostolou M, Polytarchou C, Aggelidou E, Drakaki A, Poultsides GA, Jaeger SA, Ogata H, Karin M, Struhl K, Hadzopoulou-Cladaras M. An HNF4α-miRNA inflammatory feedback circuit regulates hepatocellular oncogenesis. Cell. 2011;147:1233-1247. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 354] [Cited by in F6Publishing: 378] [Article Influence: 31.5] [Reference Citation Analysis (0)] |
| 124. | Lachenmayer A, Toffanin S, Cabellos L, Alsinet C, Hoshida Y, Villanueva A, Minguez B, Tsai HW, Ward SC, Thung S. Combination therapy for hepatocellular carcinoma: additive preclinical efficacy of the HDAC inhibitor panobinostat with sorafenib. J Hepatol. 2012;56:1343-1350. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 152] [Cited by in F6Publishing: 164] [Article Influence: 13.7] [Reference Citation Analysis (0)] |
| 125. | Guo ZS, Hong JA, Irvine KR, Chen GA, Spiess PJ, Liu Y, Zeng G, Wunderlich JR, Nguyen DM, Restifo NP. De novo induction of a cancer/testis antigen by 5-aza-2’-deoxycytidine augments adoptive immunotherapy in a murine tumor model. Cancer Res. 2006;66:1105-1113. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 115] [Cited by in F6Publishing: 120] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 126. | Landsberg J, Kohlmeyer J, Renn M, Bald T, Rogava M, Cron M, Fatho M, Lennerz V, Wölfel T, Hölzel M. Melanomas resist T-cell therapy through inflammation-induced reversible dedifferentiation. Nature. 2012;490:412-416. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 395] [Cited by in F6Publishing: 429] [Article Influence: 35.8] [Reference Citation Analysis (0)] |
| 127. | Coral S, Sigalotti L, Colizzi F, Spessotto A, Nardi G, Cortini E, Pezzani L, Fratta E, Fonsatti E, Di Giacomo AM. Phenotypic and functional changes of human melanoma xenografts induced by DNA hypomethylation: immunotherapeutic implications. J Cell Physiol. 2006;207:58-66. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 42] [Cited by in F6Publishing: 43] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 128. | Porta C, De Amici M, Quaglini S, Paglino C, Tagliani F, Boncimino A, Moratti R, Corazza GR. Circulating interleukin-6 as a tumor marker for hepatocellular carcinoma. Ann Oncol. 2008;19:353-358. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 108] [Cited by in F6Publishing: 121] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 129. | Ozen C, Yildiz G, Dagcan AT, Cevik D, Ors A, Keles U, Topel H, Ozturk M. Genetics and epigenetics of liver cancer. N Biotechnol. 2013;30:381-384. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 71] [Cited by in F6Publishing: 79] [Article Influence: 7.2] [Reference Citation Analysis (0)] |