Published online Apr 21, 2010. doi: 10.3748/wjg.v16.i15.1859
Revised: February 2, 2010
Accepted: February 9, 2010
Published online: April 21, 2010
AIM: To investigate the sphingosine 1-phosphate (S1P) receptor expression profile in human esophageal cancer cells and the effects of S1P5 on proliferation and migration of human esophageal cancer cells.
METHODS: S1P receptor expression profile in human esophageal squamous cell carcinoma cell line Eca109 was detected by semi-quantitative reverse transcription polymerase chain reaction. Eca109 cells were stably transfected with S1P5-EGFP or control-EGFP constructs. The relation between the responses of cell proliferation and migration to S1P and S1P5 expression was evaluated by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide and migration assay, respectively.
RESULTS: Both normal human esophageal mucosal epithelium and Eca109 cells expressed S1P1, S1P2, S1P3 and S1P5, respectively. Esophageal mucosal epithelium expressed S1P5 at a higher level than Eca109 cell line. S1P5 over-expressing Eca109 cells displayed spindle cell morphology with elongated and extended filopodia-like projections. The proliferation response of S1P5-transfected Eca109 cells was lower than that of control vector-transfected cells with or without S1P stimulation (P < 0.05 or 0.01). S1P significantly inhibited the migration of S1P5-transfected Eca109 cells (P < 0.001). However, without S1P in transwell lower chamber, the number of migrated S1P5-transfected Eca109 cells was greater than that of control vector-transfected Eca109 cells (P < 0.001).
CONCLUSION: S1P binding to S1P5 inhibits the proliferation and migration of S1P5-transfected Eca109 cells. Esophageal cancer cells may down-regulate the expression of S1P5 to escape the inhibitory effect.
- Citation: Hu WM, Li L, Jing BQ, Zhao YS, Wang CL, Feng L, Xie YE. Effect of S1P5 on proliferation and migration of human esophageal cancer cells. World J Gastroenterol 2010; 16(15): 1859-1866
- URL: https://www.wjgnet.com/1007-9327/full/v16/i15/1859.htm
- DOI: https://dx.doi.org/10.3748/wjg.v16.i15.1859
Esophageal cancer is one of the most common malignancies worldwide, occurring mostly in developing countries with marked regional variations in incidence. Its mortality is very similar to its incidence due to relatively late diagnosis and inefficient treatment. About 300 000 people would die of esophageal cancer each year in the world[1]. The incidence and mortality of esophageal cancer are unusually high in China, especially in Henan, Shanxi, Hebei and Sichuan Provinces[2]. Even though a small number of esophageal cancer patients can survive more than 5 years after initial surgical treatment, over 60% of patients still die of metastasis and local recurrence[3,4]. The ability to reverse the outcome of esophageal cancer is limited due to a poor understanding of its biology.
Sphingosine 1-phosphate (S1P) is a pleiotropic phospholipid mediator that mediates diverse cellular responses, such as cell proliferation, differentiation, migration, adhesion, and morphogenesis. S1P with a low nanomolar affinity binds to its 5-related G-protein coupled receptors (GPCR), and is designated as S1P1-5 (originally termed EDG-1, 3, 5, 6, and 8, respectively). S1P1, S1P2, and S1P3 receptors are ubiquitously expressed, whereas S1P4 is only expressed in lymphoid tissues and platelets, and S1P5 is predominantly expressed in the central nervous system[5-11].
Previous reports have highlighted the importance of S1P and its receptors for the growth, metastasis and infiltration of tumors, such as gastric cancer[5], thyroid cancer[12], ovarian cancer[13-16], and glioma[17]. In general, S1P stimulates the migration of S1P3 dominantly expressing cell lines and inhibits the migration of S1P2 expressing cell lines. S1P1, S1P2 and S1P3 contribute positively to S1P-stimulated glioma cell proliferation, and S1P5 blocks glioma cell proliferation. S1P1 and S1P3 enhance glioma cell migration and invasion, while S1P2 inhibits cell migration but unexpectedly enhances glioma cell invasiveness by stimulating cell adhesion[17].
The possible role and importance of S1P and its receptors in the growth and metastasis of human esophageal cancer have not been investigated. In this study, we focused on esophageal squamous cell carcinoma (ESCC), characterized S1P receptor expression patterns and investigated the role of S1P receptors in proliferation and migration of ESCC cells. ESCC cell line Eca109 expressed 4 S1P receptors (S1P1, S1P2, S1P3 and S1P5). Compared with normal esophageal mucosa epithelium, Eca109 cells expressed a very low level of S1P5. Furthermore, by over-expressing human S1P5 in Eca109 cells, the effects of S1P5 on esophageal cancer cells were partly clarified.
S1P was obtained from Cayman Chemical (Ann Arbor, MI, USA). Dispase II, fibronectin and fatty acid-free bovine serum albumin (BSA) were purchased from Sigma (St Louis, MO, USA). SV total RNA isolation system and pGEM-T easy vector system were bought from Promega (Madison, WI, USA). PrimeScript™ reverse transcription polymerase chain reaction (RT-PCR) kit and PrimerSTAR HS DNA polymerase with GC buffer were purchased from TakaRa Biotech (Dalian, China). Lipofectamine 2000 and G418 were obtained from Invitrogen (Carlsbad, CA, USA).
Normal human esophageal mucosa samples were obtained from surgically resected esophagus of patients with esophageal carcinoma, and then confirmed by an experienced pathologist to contain neither macroscopic tumor nor histologically detectable metaplastic cells or cancer cells. Samples were collected with the signed informed consent from patients. Tunica mucosa was separated from the tissues with forceps, and cut into 1 cm × 1 cm sections which were then transferred to a 100 mm culture dish containing 10 mL PBS with 0.25% dispase II and incubated for 18-24 h at 4°C. Epithelial layers were separated from the full-thickness mucosa with forceps.
Human ESCC cell line Eca109 was obtained from Shanghai Institute of Cell Biology, Chinese Academy of Sciences. Eca109 cells were cultured in RPMI 1640 medium supplemented with 10% (v/v) fetal bovine serum (FBS) and 100 U/mL of penicillin and streptomycin at 37°C in a humidified incubator containing 5% CO2.
Total RNA was isolated with the SV total RNA isolation system following its manufacturer’s instructions. RNA quality and integrity were checked by absorbance spectrometry and agarose gel electrophoresis. Semi-quantitative RT-PCR was performed to confirm differential expression of S1P receptors in Eca109 cell line. The sequences of primers used are as follows: 5'-TATCAGCGCGGACAAGGAGAACAG-3' (sense) and 5'-ATAGGCAGGCCACCCAGGATGAG-3' (antisense) for S1P1 receptor (429-bp product), 5'-TCGGCCTTCATCGTCATCCTCT-3' (sense) and 5'-CCTCCCGGGCAAACCACTG-3' (antisense) for S1P2 receptor (220-bp product), 5'-CTGCCTGCACAATCTCCCTGACTG-3’ (sense) and 5'-GGCCCGCCGCATCTCCT-3' (antisense) for S1P3 receptor (394-bp product), 5'-GAGAGCGGGGCCACCAAGAC-3' (sense) and 5'-GGTTGACCGCCGAGTTGAGGAC-3' (antisense) for S1P4 receptor (454-bp product), 5'-GGAGTAGTTCCCGAAGGACC-3' (sense) and 5'-TCTAGAATCCACGGGGTCTG-3' (antisense) for S1P5 receptor (236-bp product), 5'-GATGACATCAAGAAGGTGGTGAA-3' (sense) and 5'-GTCTTACTCCTTGGAGGCCATGT-3' (antisense) for glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (246-bp product). Thirty-two cycles of amplification were performed using a thermal cycler, Mastercycle® gradient (Eppendorf, Hamburg, Germany) at 94°C for 30 s, at 60°C for 30 s, and at 72°C for 1 min. The PCR products were electrophoresed on a 1.6% agarose gel and visualized with ethidium bromide staining.
Human S1P5 open reading frame was cloned from normal esophageal mucosal epithelium cDNA via PCR using the PrimerSTAR HS DNA polymerase with GC buffer. The sequences of primers used are 5'-CATTGAAGCTTCCACCATGGAGTCGGGGCTGCTG-3' (sense) and 5'-CATTGTCTAGAGTCTGCAGCCGGTTCTGATA-3' (antisense). The sense primer was designed to add a HindIII site upstream of a consensus Kozak sequence and the S1P5 initiating Met (italics). The antisense primer was designed to remove the S1P5 stop codon and to add a XbaI site. PCR conditions, established to amplify the S1P5 receptor sequence, were 94°C for 1 min, followed by 35 cycles at 94°C for 30 s, at 60°C for 30 s, at 72°C for 2 min, and a final extension at 72°C for 8 min. The PCR products were electrophoresed on 0.8% agarose gel and the corresponding band was cut and purified. The purified PCR products were ligated into the pGEM-T easy vector and the sequence was confirmed by sequence analysis. The S1P5 gene was spliced out with HindIII/XbaI and ligated into the Myc-His-EGFP-N1 frame, which was purified from similarly digested S1P1-Myc-His-EGFP-N1 as previously described[18]. The initiating methionine of the GFP open reading frame in pEGFP-N1 was mutated to alanine. The resulting plasmid was designated as S1P5-EGFP. A HindIII/XbaI-digested DNA oligonucleotide containing both sense (5'-CATTGAAGCTTCCACCATGGAATTCTGCAGTCGACGGTACTCTAGACCG-3') and antisense (5'-CGGTCTAGAGTACCGTCGACTGCAGAATTCCATGGTGGAAGCTTCAATG-3') sequences was cloned into similarly digested Myc-His-EGFP-N1 frame as a control vector. The DNA oligonucleotide contained initiating Met and multiple cloning sites. The plasmid was designated as control-EGFP.
Cells were transfected using Lipofectamine 2000, according to its manufacturer’s instructions. Briefly, 5 × 105 Eca109 cells in 2 mL of growth medium without antibiotics were seeded into each well of 6-well plates and grown overnight, then transfected using Lipofectamine 2000. The cells were split at a 1:20 dilution into a fresh growth medium 24 h after transfection. G418 was added at a concentration of 0.8 mg/mL the following day. Resistant colonies were picked to a 96-well plate 14 d after selection and several clones were analyzed. The cells were subsequently maintained in a medium containing G418 at 0.2 mg/mL.
Cells (5 × 103/well in 100 μL/well) were seeded into a 96-well plate and cultured in RPMI 1640 medium containing 10% FBS for 8 h at 37°C in an atmosphere containing 5% CO2 to allow attachment. The medium was then aspirated and a fresh medium containing 0.1% fatty acid-free BSA and varying concentrations of S1P was added into three replicate wells every 48 h. After incubation at 37°C in an atmosphere containing 5% CO2 for 72 h, the number of living cells was calculated by 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT) assay. Briefly, a MTT solution was added into each well and the cells were incubated for an additional 4 h. The number of living cells was then counted by measuring the absorbance at 490 nm.
Migration assay was performed on a 6.5 mm-diameter transwell chamber with an 8 μm pore size (Corning, NY, USA). The underside of transwell membranes was precoated with fibronectin (10 μg/mL) and allowed to dry. Cells at 70%-80% confluence were serum starved overnight, detached from culture plates by 0.02% EDTA, washed and resuspended in RPMI 1640 medium with 0.1% fatty acid-free BSA at 1 × 106 cells/mL. For migration assay, cells (1 × 105 cells/well) were added into the upper chamber of transwell chambers separated by inserts with 8 μm pores in the presence or absence of S1P. Following an 8-h incubation, nonmigrating cells on the upper surface of membranes were removed with cotton swabs, fixed and stained with 0.1% crystal violet in phosphate-buffered saline (PBS) at the ambient temperature, and then destained with PBS. Migrating cells were counted in 5 randomly selected high-power fields per membrane under an Olympus IX71 inverted microscope.
Data are expressed as mean ± SD and shown as error bars in the figures. Statistical comparisons were made by Student’s t-test. One-way ANOVA with a post hoc Bonferroni testing was employed in case of multiple comparisons. P < 0.05 was considered statistically significant.
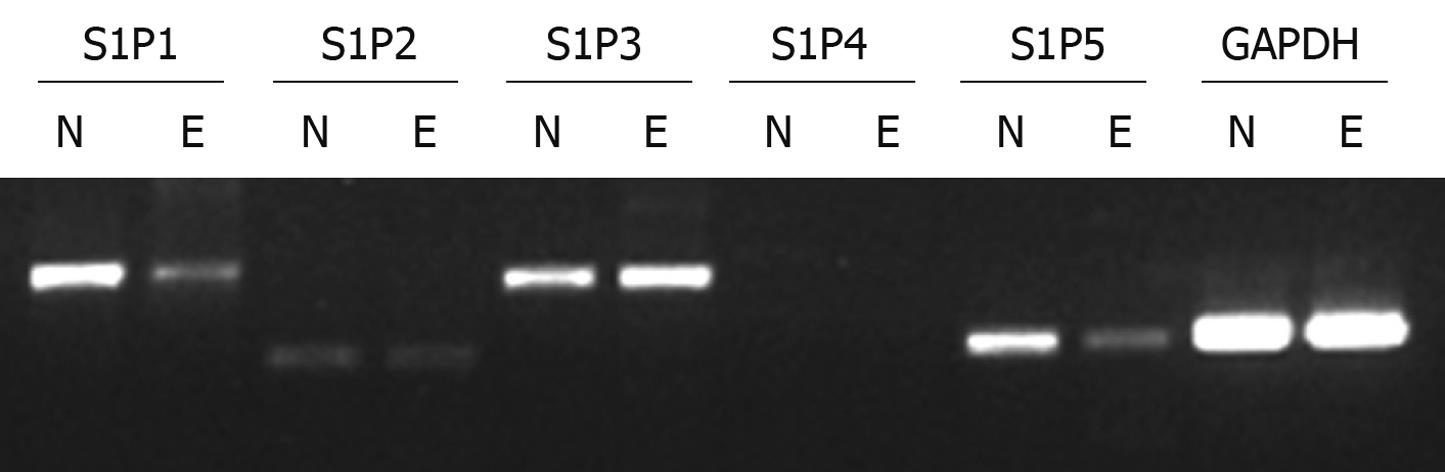
Semi-quantitative RT-PCR of mRNA was performed to observe which S1P receptors are expressed in Eca109 cell line and normal human esophageal mucosal epithelium. For comparison, data were also normalized to the expression of the reference gene GAPDH, which gave similar results. Four S1P receptors were found in normal human esophageal mucosal epithelium with the following rank order of mRNA abundance: S1P1>S1P5>S1P3>S1P2. In contrast to Eca109 cell line, the mRNA abundance in S1P receptors was S1P3>S1P1>S1P5>S1P2. S1P4 expression was absent or minimal in Eca109 cell line and normal human esophageal mucosal epithelium (Figure 1). Eca109 cell line expressed S1P3 at a higher level than normal esophageal mucosal epithelium. In contrast, normal esophageal mucosal epithelium expressed S1P1 and S1P5 at a higher level than Eca109 cell line. On the basis of these results, S1P5 was chosen for further study.
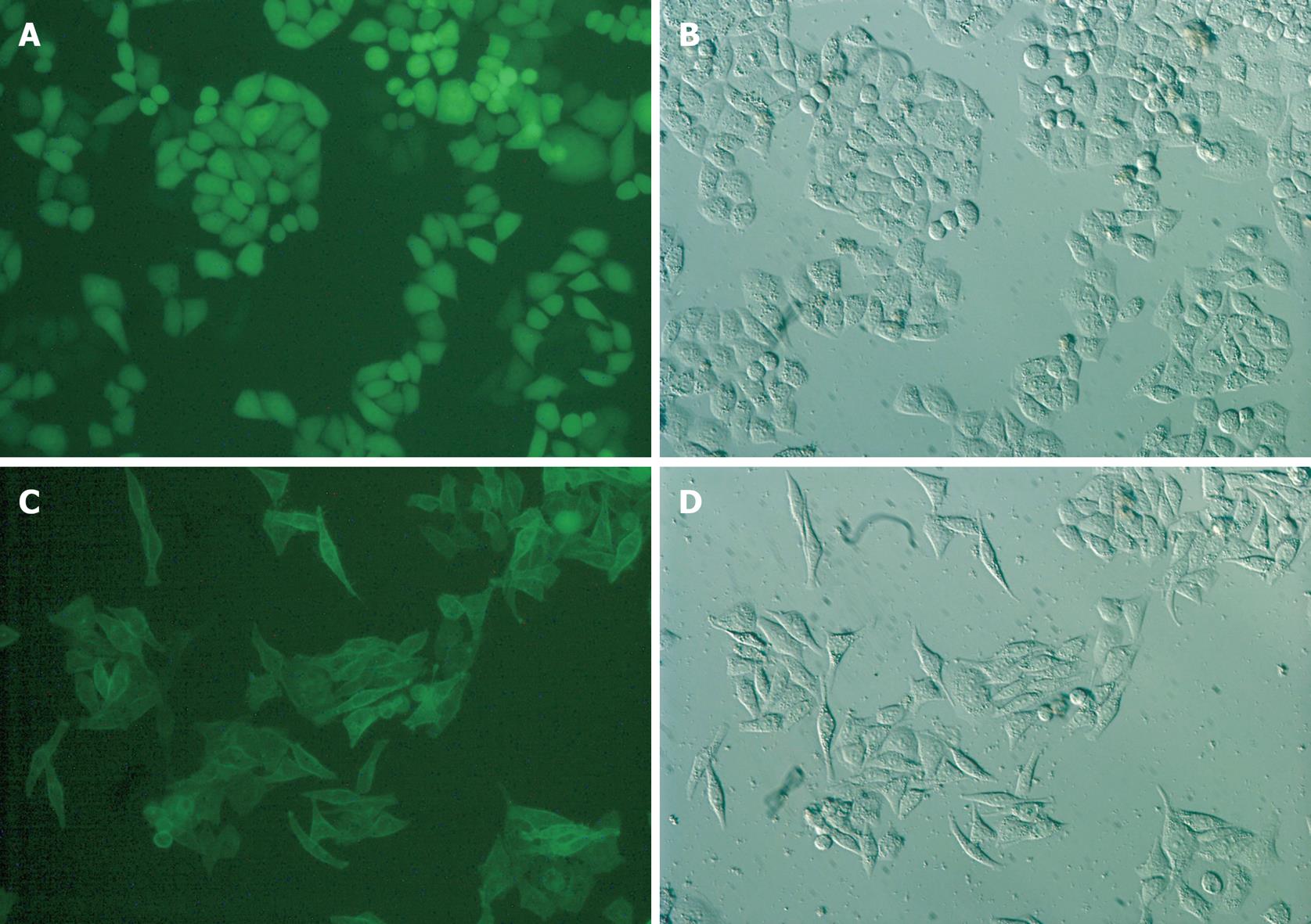
Eca109 cells transfected with the S1P5-EGFP or Control-EGFP constructs were cultured in a medium containing 10% FBS. Their localization was visualized by fluorescence microscopy. As shown in Figure 2, control-EGFP was localized at the cytosol of Eca109 cells. In contrast, the S1P5-EGFP was localized at the plasma membrane. Interestingly, control-EGFP-transfected cells displayed the characteristic Eca109 cell morphology. In contrast, S1P5-EGFP-transfected cells displayed spindle cell morphology with elongated and extended filopodia-like projections in a medium containing 10% FBS or 0.1% fatty acid-free BSA.
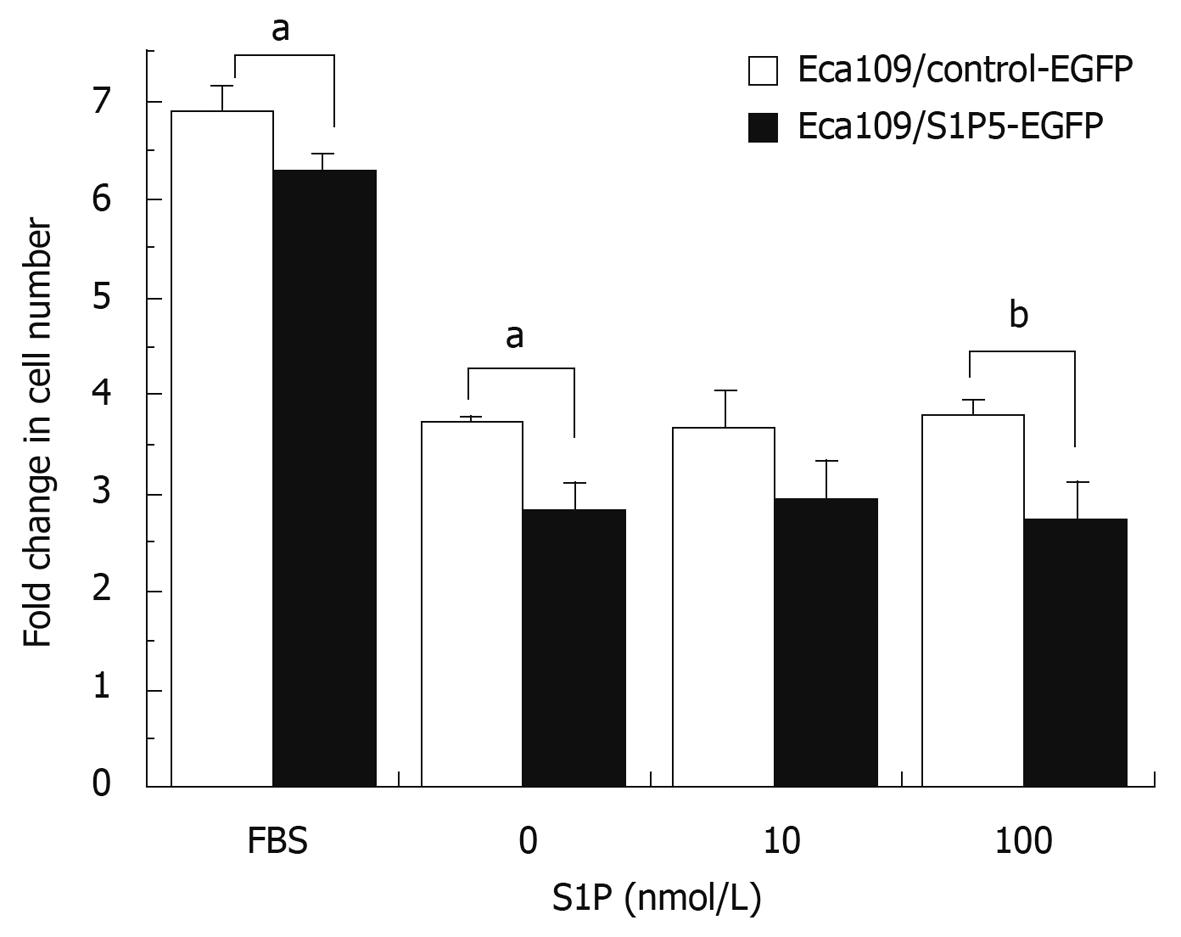
Since S1P regulates the proliferation of many cell types, including cancer cells, it is of great interest to investigate whether S1P5 or S1P binding to S1P5 has an effect on the proliferation of control vector and S1P5 transfected Eca109 cells. To assess induction of proliferation, stably transfected cells in 96-well plates containing 10% FBS or 0.1% fatty acid-free BSA were counted after treatment with a vehicle (DMSO) and 10 or 100 nmol/L S1P for 72 h. The number of cells from both control vector and S1P5 transfectants did not increase in response to S1P (P > 0.05), suggesting that S1P does not behave as a mitogen at S1P5 (Figure 3).
Interestingly, the proliferation response of S1P5-transfected Eca109 cells was lower than that of control vector-transfected cells in a medium containing 10% FBS (8% lower, P < 0.05) or 0.1% fatty acid-free BSA with or without S1P (20%-29% lower, P < 0.05 or 0.01), suggesting that S1P5 has an intrinsic activity and inhibits cell proliferation.
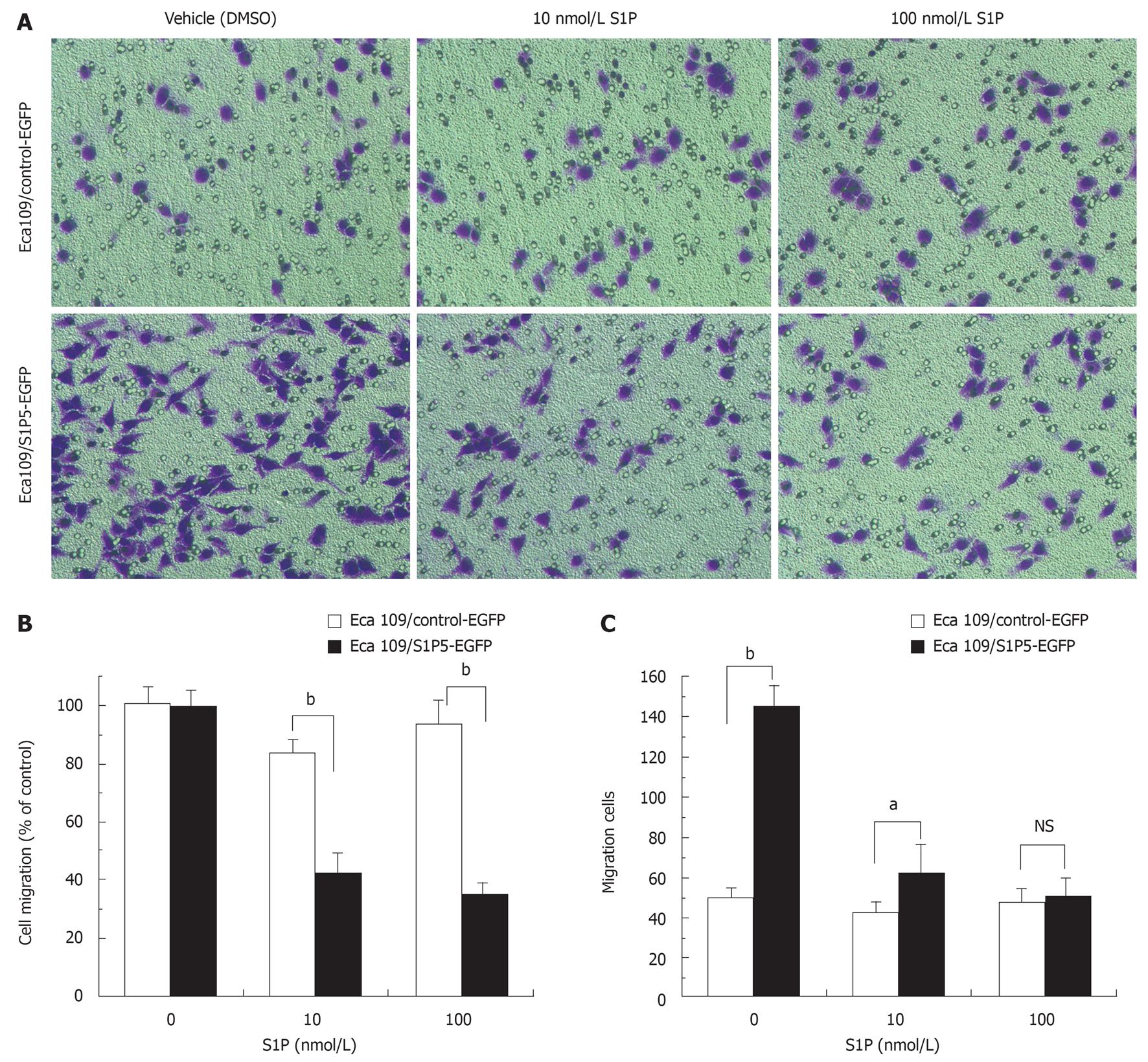
Migration of Eca109 cells was quantified by transwell-based assay. Transwell membranes were precoated with fibronectin. At a concentration of 10 or 100 nmol/L S1P, the migration of S1P5-EGFP-transfected Eca109 cells was significantly lower than that of control-EGFP-transfected Eca109 cells (48% vs 84% at 10 nmol/L S1P, 39% vs 94% at 100 nmol/L S1P, P < 0.001) (Figure 4A and B). Interestingly, without S1P in the lower chamber, the migrated number of S1P5-EGFP transfected Eca109 cells (145 ± 10) was more than that of control-EGFP- transfected Eca109 cells (51 ± 4, P <0.001) (Figure 4A and C). Both cells failed to migrate on gelatin-coated or non-coated transwell membranes (data not shown). Control-EGFP and S1P5-EGFP transfected Eca109 cells exhibited spontaneous migration without S1P, and the addition of S1P to the lower chamber potently inhibited cell migration.
In the present study, normal esophageal mucosal epithelium expressed S1P5 at a high level, and Eca109 cells expressed S1P5 at a low level. S1P5 was predominantly expressed in the white matter tracts and oligodendrocytes, and was particularly abundant in the anterior commissure, corpus callosum, and optic tract. S1P5 mRNA was detected in spleen, peripheral blood leukocytes, placenta, lung, aorta, and fetal spleen. A low level of signals has been detected in many tissues[19,20]. In this study, normal human esophageal mucosal epithelium expressed S1P5 at a high level, suggesting that S1P5 is also a ubiquitous S1P receptor and plays an important role in different tissues and cells.
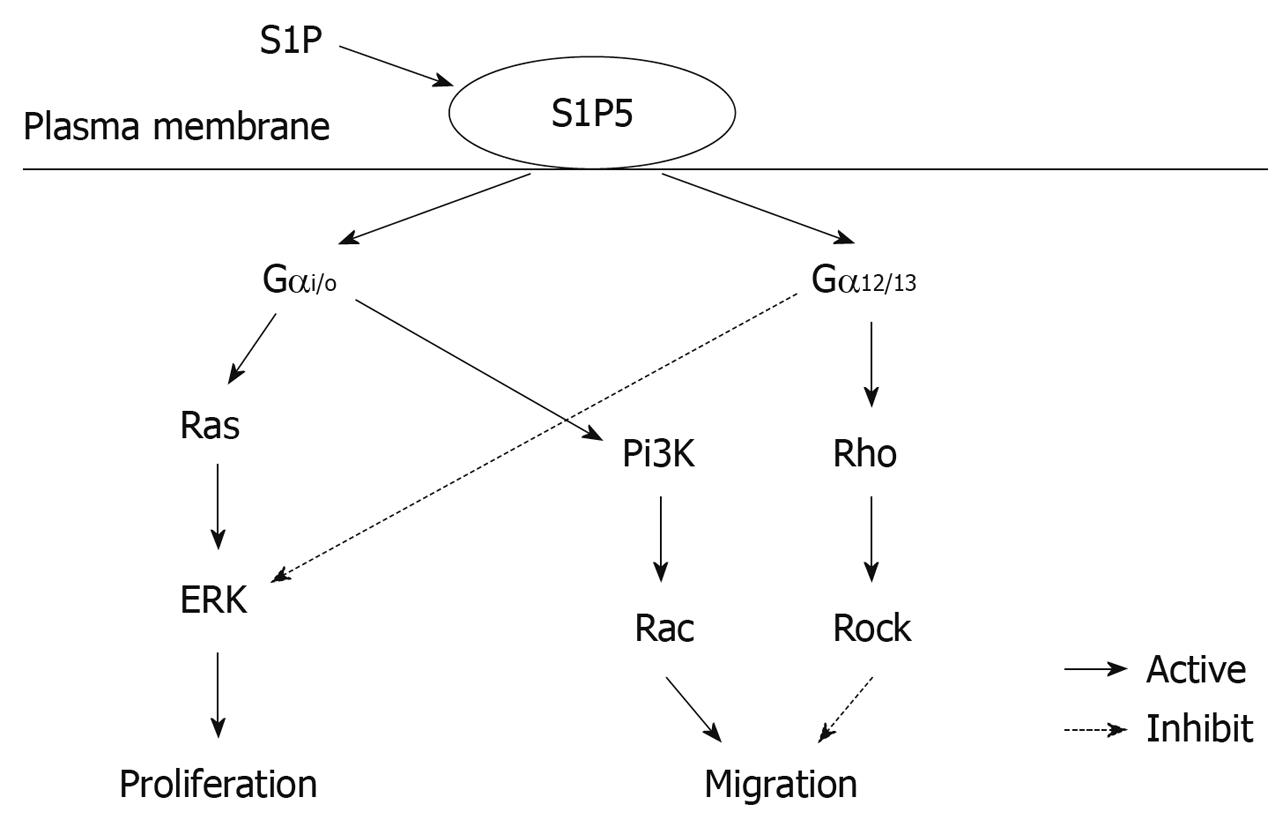
S1P is believed to be generated intracellularly by all cell types during sphingolipid degradation. Plasma S1P is mainly hematopoietic in origin with erythrocytes as a major contributor to the sources of extracellular S1P, whereas lymph S1P is from a distinct radiation-resistant source[21,22]. The concentration of S1P in plasma and serum is 0.1-0.6 μmol/L and 0.4-1.1 μmol/L, respectively, while the S1P level in tissues is 0.5-75 pmol/mg[23]. Although S1P was originally thought to act as an intracellular second messenger, it has been widely accepted as an intercellular agonist. S1P binding to its receptors activates different intracellular signaling pathways depending on which intracellular Gα protein they couple to. The S1P1 receptor primarily couples through Gαi/o, whereas S1P2 and S1P3 can couple through Gαi/o, Gαq and Gα12/13, S1P4 and S1P5 can couple through Gαi/o and Gα12/13[6,7,24-26]. Signaling through Gαi/o has been associated with the activation of small guanosine triphosphatase (GTPase) Ras and extracellular signal-regulated kinase (ERK) to promote proliferation, and induction of Pi3K and small GTPase Rac to promote migration. Signaling through Gα12/13 can promote the activation of small GTPase Rho and Rho-associated kinase (ROCK) to inhibit migration[6]. Furthermore, S1P5 is also a constitutively active receptor with the ability to intrinsically stimulate various effector pathways albeit at a different extent in the absence of stimulating ligand[27].
In the present study, normal esophageal mucosal epithelium expressed S1P5 at a higher level than Eca109 cell line. Therefore, Eca109 cells were transfected with the expression constructs of S1P5 or control vector in order to delineate the effects of S1P5 on the growth, migration and morphogenesis of esophageal cancer cells. Eca109 cells that overexpress S1P5 were analyzed by phase contrast microscopy. Increased expression of S1P5 induced cell spindle change with elongated and extended filopodia-like projections in a medium containing 10% FBS or 0.1% fatty acid-free BSA. This morphological change is characterized by motile cells. However, transfection of several mammalian cell lines (CHO-K1, HEK293, NIH3T3, RH7777) with the S1P5 receptor induced cell rounding change, which is more pronounced in the presence of S1P-containing serum[27]. Differences in morphological alteration may correlate well with cell type or malignant degree, because CHO-K1, HEK293, NIH3T3 and RH7777 are the normally-derived mammalian cell lines.
In this study, the expression of S1P5 receptor in Eca109 cells reduced the proliferation of Eca109 cells whether or not the medium contained S1P, indicating that the S1P5 receptor is constitutively active in the absence of stimulating ligand. Antiproliferative effect is an intrinsic ability of the S1P5 receptor that is insensitive to ligand stimulation. Analogously, S1P5-transfected CHO cells exhibit an intrinsic activity and inhibit serum-stimulated cell proliferation[28]. S1P5 can negatively and constitutively regulate cellular proliferation by reducing ERK1/2 activity via protein-tyrosine phosphatase. Gα12/13 is a likely candidate G protein for the inhibitory effects of S1P5 on ERK activity. It has been shown that mutation- activated Gα12/13, but not Gαq, Gαs, or Gαi2, can inhibit EGF stimulation of ERK activity in COS-7 cells in a Ras- and Raf-independent manner[29]. Down-regulation of S1P5 in esophageal cancer cells may weaken its antiproliferative effect.
In this study, overexpression of S1P5 receptor constitutively increased the migration of esophageal cancer cells to fibronectin in the absence of S1P, which was consistent with morphological alteration. However, addition of 10 and 100 nmol/L S1P significantly inhibited the constitutive migration of S1P5-transfected Eca109 cells. It has been shown that small GTPases of the Rho family, including Rac, Cdc42, and RhoA, are well-known regulators of cell migration[5,30]. Cell migration induced by S1P receptors (S1P1 and S1P3) involves Gαi/o-mediated activation of phosphatidylinositol 3-kinase (Pi3K) and Rac. S1P receptors (S1P2 and S1P5) mediate inhibition of cell migration by Gα12/13-evoked stimulation of Rho[12]. It was reported that fibronectin binding to αvβ3 integrin can decrease Rho activity and stimulate cell motility and spreading, suggesting that enhanced Rho activity may reduce integrin-driven cell migration[7,31]. Our data support that S1P5 may constitutively induce cell migration due to Gαi/o-activated Rac and inhibit cell migration due to Gα12/13-stimulated Rho. In a microenvironment containing S1P interstitial fluid, esophageal cancer cells may escape the inhibitory effect of S1P-S1P5 on their migration by down-regulating the expression of S1P5. The putative signaling pathway of S1P5 is shown in Figure 5.
In summary, S1P5 is a constitutively active receptor with the ability to intrinsically inhibit cell proliferation and induce cell migration. Cell migration can be inhibited by stimulating extracellular S1P. In normal esophageal mucosa cells, a high expression level of S1P5 may inhibit cell proliferation and migration while esophageal cancer cells may down-regulate the expression of S1P5 to promote cell proliferation and escape the inhibitory effect of S1P-S1P5 on cell migration. Our study is the first to show that the deficiency in inhibitory effect of S1P-S1P5 is of importance in the growth and metastasis of esophageal cancer cells. Further study is needed on the role of S1P and S1P5 receptor in esophageal cancer.
Esophageal cancer is one of the most common malignancies worldwide. Its mortality is very high due to relatively late diagnosis and inefficient treatment. The ability to reverse the outcome of esophageal cancer is limited due to a poor understanding of its biology. Progression of esophageal cancer may be associated with sphingosine 1-phosphate (S1P) and its receptors S1P1-5, which play an important role in other cancers.
S1P is a pleiotropic phospholipid mediator. S1P binding to its 5 receptors S1P1-5 mediates diverse cellular responses such as cell proliferation, differentiation, migration, adhesion, and morphogenesis. Recent studies have shown that S1P and its receptors are related with cancer growth and metastasis. In this study, esophageal squamous cell carcinoma (ESCC), S1P receptor expression pattern, and the role of S1P receptors in proliferation and migration of ESCC cells were studied.
Recent reports have highlighted the importance of S1P and its receptors for the growth, metastasis and infiltration of tumors, such as gastric cancer, thyroid cancer, ovarian cancer, and glioma. This study showed that deficiency in inhibitory effect of S1P-S1P5 may be of importance in the growth and metastasis of esophageal cancer.
The results of this study indicate that deficiency in inhibitory effect of S1P-S1P5 may be of importance in the growth and metastasis of esophageal cancer. S1P5 or its associated signaling molecules may serve as a future strategy in biotherapy for esophageal cancer.
S1P is a bioactive lipid that regulates central cellular processes such as cell proliferation, differentiation, migration, adhesion, and morphogenesis. S1P is present in plasma and serum at high nanomolar concentrations. S1P binding to related G-protein coupled receptors is designated as S1P1-5. S1P1, S1P2, and S1P3 receptors which are ubiquitously expressed, whereas S1P4 is restricted to lymphoid tissues and platelets and S1P5 is predominantly expressed in the central nervous system. Binding of S1P to its receptors activates different signaling pathways via the heterotrimeric G proteins.
The paper is well written and presents an interesting aspect of the pathophysiology of esophageal mucosa which is related to the evolution of esophageal cancer.
Peer reviewers: Vincenzo Villanacci, MD, 2nd Department of Pathology, Spedali Civili Piazzale Spedali Civili 1, Brescia, 25100, Italy; Toru Hiyama, MD, PhD, Health Service Center, Hiroshima University, 1-7-1 Kagamiyama, Higashihiroshima 739-8521, Japan
S- Editor Wang YR L- Editor Wang XL E- Editor Ma WH
| 1. | Miyazaki T, Kato H, Fukuchi M, Nakajima M, Kuwano H. EphA2 overexpression correlates with poor prognosis in esophageal squamous cell carcinoma. Int J Cancer. 2003;103:657-663. [Cited in This Article: ] |
| 2. | Stoner GD, Gupta A. Etiology and chemoprevention of esophageal squamous cell carcinoma. Carcinogenesis. 2001;22:1737-1746. [Cited in This Article: ] |
| 3. | Krasna MJ, Mao YS, Sonett JR, Tamura G, Jones R, Suntharalingam M, Meltzer SJ. P53 gene protein overexpression predicts results of trimodality therapy in esophageal cancer patients. Ann Thorac Surg. 1999;68:2021-2024; discussion 2024-2025. [Cited in This Article: ] |
| 4. | Xie YE, Tang EJ, Zhang DR, Ren BX. Down-regulation of Bcl-XL by RNA interference suppresses cell growth and induces apoptosis in human esophageal cancer cells. World J Gastroenterol. 2006;12:7472-7477. [Cited in This Article: ] |
| 5. | Yamashita H, Kitayama J, Shida D, Yamaguchi H, Mori K, Osada M, Aoki S, Yatomi Y, Takuwa Y, Nagawa H. Sphingosine 1-phosphate receptor expression profile in human gastric cancer cells: differential regulation on the migration and proliferation. J Surg Res. 2006;130:80-87. [Cited in This Article: ] |
| 6. | Brinkmann V. Sphingosine 1-phosphate receptors in health and disease: mechanistic insights from gene deletion studies and reverse pharmacology. Pharmacol Ther. 2007;115:84-105. [Cited in This Article: ] |
| 7. | Novgorodov AS, El-Alwani M, Bielawski J, Obeid LM, Gudz TI. Activation of sphingosine-1-phosphate receptor S1P5 inhibits oligodendrocyte progenitor migration. FASEB J. 2007;21:1503-1514. [Cited in This Article: ] |
| 8. | Sanchez T, Hla T. Structural and functional characteristics of S1P receptors. J Cell Biochem. 2004;92:913-922. [Cited in This Article: ] |
| 9. | Rosen H, Goetzl EJ. Sphingosine 1-phosphate and its receptors: an autocrine and paracrine network. Nat Rev Immunol. 2005;5:560-570. [Cited in This Article: ] |
| 10. | Taha TA, Argraves KM, Obeid LM. Sphingosine-1-phosphate receptors: receptor specificity versus functional redundancy. Biochim Biophys Acta. 2004;1682:48-55. [Cited in This Article: ] |
| 11. | Anliker B, Chun J. Cell surface receptors in lysophospholipid signaling. Semin Cell Dev Biol. 2004;15:457-465. [Cited in This Article: ] |
| 12. | Balthasar S, Samulin J, Ahlgren H, Bergelin N, Lundqvist M, Toescu EC, Eggo MC, Törnquist K. Sphingosine 1-phosphate receptor expression profile and regulation of migration in human thyroid cancer cells. Biochem J. 2006;398:547-556. [Cited in This Article: ] |
| 13. | Park KS, Kim MK, Lee HY, Kim SD, Lee SY, Kim JM, Ryu SH, Bae YS. S1P stimulates chemotactic migration and invasion in OVCAR3 ovarian cancer cells. Biochem Biophys Res Commun. 2007;356:239-244. [Cited in This Article: ] |
| 14. | Smicun Y, Gil O, Devine K, Fishman DA. S1P and LPA have an attachment-dependent regulatory effect on invasion of epithelial ovarian cancer cells. Gynecol Oncol. 2007;107:298-309. [Cited in This Article: ] |
| 15. | Smicun Y, Reierstad S, Wang FQ, Lee C, Fishman DA. S1P regulation of ovarian carcinoma invasiveness. Gynecol Oncol. 2006;103:952-959. [Cited in This Article: ] |
| 16. | Hong G, Baudhuin LM, Xu Y. Sphingosine-1-phosphate modulates growth and adhesion of ovarian cancer cells. FEBS Lett. 1999;460:513-518. [Cited in This Article: ] |
| 17. | Young N, Van Brocklyn JR. Roles of sphingosine-1-phosphate (S1P) receptors in malignant behavior of glioma cells. Differential effects of S1P2 on cell migration and invasiveness. Exp Cell Res. 2007;313:1615-1627. [Cited in This Article: ] |
| 18. | Watterson KR, Johnston E, Chalmers C, Pronin A, Cook SJ, Benovic JL, Palmer TM. Dual regulation of EDG1/S1P(1) receptor phosphorylation and internalization by protein kinase C and G-protein-coupled receptor kinase 2. J Biol Chem. 2002;277:5767-5777. [Cited in This Article: ] |
| 19. | Im DS, Clemens J, Macdonald TL, Lynch KR. Characterization of the human and mouse sphingosine 1-phosphate receptor, S1P5 (Edg-8): structure-activity relationship of sphingosine1-phosphate receptors. Biochemistry. 2001;40:14053-14060. [Cited in This Article: ] |
| 20. | Terai K, Soga T, Takahashi M, Kamohara M, Ohno K, Yatsugi S, Okada M, Yamaguchi T. Edg-8 receptors are preferentially expressed in oligodendrocyte lineage cells of the rat CNS. Neuroscience. 2003;116:1053-1062. [Cited in This Article: ] |
| 21. | Spiegel S, Milstien S. Sphingosine-1-phosphate: an enigmatic signalling lipid. Nat Rev Mol Cell Biol. 2003;4:397-407. [Cited in This Article: ] |
| 22. | Pappu R, Schwab SR, Cornelissen I, Pereira JP, Regard JB, Xu Y, Camerer E, Zheng YW, Huang Y, Cyster JG. Promotion of lymphocyte egress into blood and lymph by distinct sources of sphingosine-1-phosphate. Science. 2007;316:295-298. [Cited in This Article: ] |
| 23. | Oskeritzian CA, Milstien S, Spiegel S. Sphingosine-1-phosphate in allergic responses, asthma and anaphylaxis. Pharmacol Ther. 2007;115:390-399. [Cited in This Article: ] |
| 24. | Rosen H, Sanna MG, Cahalan SM, Gonzalez-Cabrera PJ. Tipping the gatekeeper: S1P regulation of endothelial barrier function. Trends Immunol. 2007;28:102-107. [Cited in This Article: ] |
| 25. | Kluk MJ, Hla T. Signaling of sphingosine-1-phosphate via the S1P/EDG-family of G-protein-coupled receptors. Biochim Biophys Acta. 2002;1582:72-80. [Cited in This Article: ] |
| 26. | Takuwa Y. Subtype-specific differential regulation of Rho family G proteins and cell migration by the Edg family sphingosine-1-phosphate receptors. Biochim Biophys Acta. 2002;1582:112-120. [Cited in This Article: ] |
| 27. | Niedernberg A, Blaukat A, Schöneberg T, Kostenis E. Regulated and constitutive activation of specific signalling pathways by the human S1P5 receptor. Br J Pharmacol. 2003;138:481-493. [Cited in This Article: ] |
| 28. | Malek RL, Toman RE, Edsall LC, Wong S, Chiu J, Letterle CA, Van Brocklyn JR, Milstien S, Spiegel S, Lee NH. Nrg-1 belongs to the endothelial differentiation gene family of G protein-coupled sphingosine-1-phosphate receptors. J Biol Chem. 2001;276:5692-5699. [Cited in This Article: ] |
| 29. | Voyno-Yasenetskaya TA, Faure MP, Ahn NG, Bourne HR. Galpha12 and Galpha13 regulate extracellular signal-regulated kinase and c-Jun kinase pathways by different mechanisms in COS-7 cells. J Biol Chem. 1996;271:21081-21087. [Cited in This Article: ] |
| 30. | Evers EE, Zondag GC, Malliri A, Price LS, ten Klooster JP, van der Kammen RA, Collard JG. Rho family proteins in cell adhesion and cell migration. Eur J Cancer. 2000;36:1269-1274. [Cited in This Article: ] |