INTRODUCTION
Globally, about 350 million people are chronically infected with hepatitis B virus (HBV), of which 15 million are positive for hepatitis D virus (HDV) antibodies[1]. HDV causes chronic or fulminant “delta hepatitis”, in form of co- or super-infection in HBV infected patients[2]. Delta virus is considered as a very deleterious pathogen since its infection commonly leads to progression of hepatic fibrosis, cirrhosis and increased risk of hepatocellular carcinoma[2,3]. There are eight known genotypes of HDV (from 1 to 8), of which genotype 1 has a worldwide distribution and genotype 3 has been associated with the most severe outcome of liver disease[4,5]. With a virion size of 36 nm and a 1.7 Kb genomic circular RNA, HDV is the smallest known human virus. Its genome encodes only two structural proteins termed small- and large-HD-antigens (S- and L-HDAg). The proteins are transcribed from the same open reading frame (ORF) and are identical except for a 19 amino acid extension in C-terminal domain of L-HDAg[2].
HDV requires the function of a helper virus as an envelope source for virion envelopment and propagation. This function can be provided through HBV (all genotypes from A to H) or other Orthohepadnaviridae members, such as Woodchuck hepatitis virus (WHV), by sharing the surface proteins[2]. The 19 amino acid extension of L-HDAg, which is called the “packaging signal”, is responsible for this interaction[6]. While HBV thereby provides an essential basis for HDV viremia and infectivity, most clinical studies reported that HBV replication is diminished in HBV-HDV-infected patients and that HDV co-infection is associated with lower HBV viremia than HBV mono-infection[7]. However, HBV-DNA, HDV-RNA and HBsAg apparently fluctuate in longitudinally studied patients indicating ongoing and dynamic interactions between HBV and HDV in infected cells[8].
Although the direct contact between HBsAg and HDAg for HDV virion envelopment can be considered the main interaction, other less well understood mechanisms may also interfere with the replication of both viruses in infected cells[9]. Here we describe possible mechanisms for HBV/HDV interactions and their probable molecular cross-talks in infected cells. These mechanisms include HBsAg-HDAg interactions and HDV-trans-controlling of HBV genome replication/transcription, cellular transcriptional pathways and RNA polymerase activity in dually infected hepatocytes.
HBsAg-HDAg INTERACTIONS
HBV encodes three surface proteins with different initiation-of-replication sites from one ORF. These proteins are large, medium and small HBsAgs (L-, M- and S-HBsAg)[10]. As an integral protein, S-HBsAg (226 amino acids) is anchored in the lipid bilayer of the endoplasmic reticulum (ER) through its N-terminal (residues 4-28 and 80-100) and C-terminal (residues 165-226) transmembrane domains (TMDs). It also includes an antigenic loop (Ag loop, residues 101-164) with immunodominant epitopes, facing the ER lumen. The rest of residues located between TMDs face the cytoplasm and are called cytosolic loops (CYLs). These are expected to be residues 29 to 79 (CYL-I) and 194 to 201 (CYL-II)[11]. The M-HBsAg (281 amino acids), contains the whole S-HBsAg plus an N-terminal preS2 region facing the ER lumen. The L-HBsAg (389-400 amino acids), contains preS1, preS2 (preS) and S domains[12]. This protein has two conformations based on the positioning of preS in ER membrane towards the cytoplasm (for virion formation) or ER lumen (for receptor binding)[11]. All three types of HBsAgs are found on the surface of mature HDV particles[2]. The schematic features of HBsAg proteins and their localization in ER membrane are shown in Figure 1.
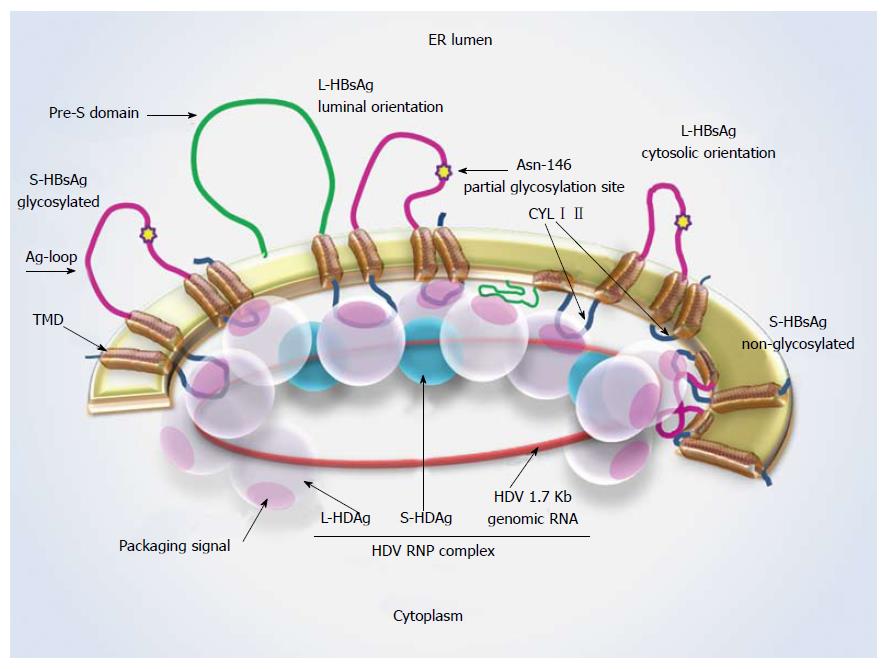
Figure 1 Hepatitis D virus-ribonucleoprotein complex interaction with S-hepatitis B virus surface antigen.
Schematic representation of L- and S-HBsAg locations in the ER membrane and the interaction sites with the HDV-RNP complex. The Asn-146 glycosylation site is highlighted by a star. About half of the residues remain non-glycosylated at this site. A cytosolic orientation has been suggested for the non-glycosylated Ag loop, which may interfere with HBsAg-HDAg interaction. L-HDAg mediates HDV assembly at the late stage of viral replication through forming connections with HDV-RNA, S-HDAg and HBsAg. Ag loop: Antigenic loop; CYL I, II: Cytosolic loop I, II; ER: Endoplasmic reticulum; L-/ S-HBsAg: Large and small hepatitis B virus surface antigens; L-/ S-HDAg: Large and small hepatitis D virus proteins; RNP: Ribonucleoprotein; TMD: Transmembrane domain.
Both HDV small and large proteins form connections to one another as well as to HDV RNA through RNA binding domains to assemble the HDV ribonucleoprotein (RNP) complex[6]. The L-HDAg is responsible for RNP localization in the ER membrane through a CXXX farnesylation signal (C stands for cytosine, and X for any amino acid) and also interactions with HBV surface proteins through its packaging signal (Figure 1)[2]. The packaging signal is very genotype specific in HDV (74% divergence between genotypes 1 and 2) and plays an important role in the envelopment. Although an association between HDV-1/HBV-A and -D and HDV-3/HBV-F and -A has been observed, independent investigations suggest that the co-infections are mainly representative of common genotypes of each of the viruses in certain geographical areas and not specific for distinct HBV or HDV genotypes[13-15]. Studies indicate that HDV genotype 2 is associated with a less aggressive disease compared to genotype 1 which has been attributed to the higher packaging efficiency of genotype 1 than that of genotype 2[16,17]. Moreover, a variation in the packaging efficiency has been observed among different isolates of the same genotype, which reflects the critical role of this length in HDAg interactions with HBsAg. It has been reported that the hydrophobic nature of the L-HDAg C-terminal domain provided by C211-farnesylation as well as the number of hydrophobic residues of the packaging signal (which differs among HDV genotypes) enhance HDV interactions with surface proteins of HBV and therefore the packaging efficiency[17].
While HBV requires both S- and L-HBAgs for viral assembly, HDV needs S-HBsAg for its in vitro virion packaging and L-HBsAg for infectivity[18,19]. Therefore, it is very likely that most of the HDAg binding sites are located on the S domain of HBsAg[20]. While an intact HBsAg is not able to interact with L-HDAg, a denatured form of HBsAg is competent for such interactions suggesting that L-HDAg has no connection to the external domains of HBsAg but rather to the domains inside the particles[21]. The cytoplasmic orientation of the CYLs of the S protein, provides a reasonable condition for these sites to interact with pre-mature HDV virions in the cytosol (Figure 1)[22].
The importance of S-HBsAg residues 24 to 28 and 56 to 80 for HDV secretion has been shown in previous studies[23,24]. Also, a C-terminal truncation of HBsAg by 50 amino acids inhibits HDV envelopment and secretion[25]. Based on these data and also from our recent observation indicating a high rate of amino acid selection at CYLs of S-HBsAg in HBV isolates from HBV/HDV infected patients (own unpublished observations), these domains are expected to make a significant contribution in HDV packaging. Of special importance are tryptophan residues at positions 196, 199 and 201 at the C-terminal domain of S-HBsAg, which are suggested to have a central localization in binding interface with HDV-RNP complex[26].
Mutational studies revealed that in addition to the receptor binding site on the pre-S1 domain of L-HBsAg, the Ag loop is also responsible for HDV virion infectivity[27]. On the other hand, in vitro experiments using mutant HBsAg with deletions in Ag loop resulted in the lack of subviral particles as well as HDV virion secretion[27]. In more detailed studies it was shown that N-glycosylation of S-HBsAg, which is mediated by the C-terminal domain of S protein and occurs partially on Asn-146, affects HDV envelopment and secretion[12]. Based on these studies, HDV secretion is delayed or reduced (about ten folds) in the presence of non-glycosylated HBsAg, while HBV and HBsAg formation is not affected[21]. Although a weakened interaction between HBsAg and other components of HDV-RNP complex (rather than L-HDAg) has been suggested for this reduction, based on the luminal positioning of Ag loop in the ER membrane, a direct interaction of this domain and HDV components is unlikely[21,27]. Different mechanisms have been suggested to explain the effects of the antigenic domain, especially in its non-glycosylated form, on HDV packaging and secretion. One is a modified maturation and trafficking process for non-glycosylated HBsAg, which in turn will affect the rate of interactions with HDV[21]. The association of HBsAg with calnexin (a molecular chaperon in ER membrane) is also affected by the glycosylation process[28]. Therefore, a non-glycosylated HBsAg is more prone to misfolding and late maturation[29]. Furthermore, it has been suggested that a non-glycosylated Ag loop faces the cytoplasm, which possibly masks the cytosolic interaction sites with HDV or hinders appropriate connections between HBsAg and HDV (Figure 1)[11]. Due to deferent propagation responses of HBV and HDV to non-glycosylated Ag loop, it is possible that these viruses apply different mechanisms to interact with HBsAg[21]. Likewise, the lateral S-S interactions between S protein carbohydrates, which play a critical role in virion stability, are suggested to occur differently for HBV and HDV due to their particle sizes[12].
INDIRECT INTERACTIONS BETWEEN HBV AND HDV AFFECTING VIRAL REPLICATION
There are several indications of low HBV replication levels in patients co-infected with HDV[7,9,30]. On the other hand, longitudinal analyses of HBV/HDV co-infected individuals demonstrated a fluctuating pattern of HBV and HDV replication over time[8]. In case that HDV is temporarily or permanently the dominant virus during dual infection with HBV, there should be a molecular scenario for these viruses to control each other’s replication. Most of the studies so far, indicate a controlling role of HDV over HBV replication or its protein expression in infected cells[7,9,31].
Previous investigations showed that HBV DNA in the host cell genome can produce enough surface antigen molecules for HDV virion assembly even in the absence of precore and pregenomic RNAs and regardless of an active HBV replication[32,33]. These cells, which still produce some of the viral products, may be selected through immune responses, appear as a result of a resolved infection or just due to the support of the infected cells for parts of the viral proteins such as envelope antigens but not the complete replication of the virus[33-35]. Nonetheless, regarding the role of HBV as an envelope provider, HDV nucleoproteins can be considered as competitors with HBV for HBsAg. Therefore, they may induce a selective suppression on HBV replication associated with an increase in PreS/S RNAs and HBsAg levels in co-infected patients[9]. Investigations on the effects of S- and L-HDAgs on HBV replication have shown that these proteins inhibit HBV replication through a strong suppression of HBV enhancers (EnhI and II) and also trans-activation of the IFN-alpha-inducible MxA gene[31]. The inhibitory effects of L-HDAg on RNA polymerase II, which is involved in replication of both HBV and HDV viruses, might be another reason for reduced HBV replication in the presence of co-infection with HDV[36].
Another indication of HBV controlled replication/gene expression in HDV infected cells is the presence of basal core promoter (BCP) and precore (PC) mutations in the HBV genome of patients co-infected with HDV. Occurrence of HBV BCP and PC mutations is associated with lower levels of HBV DNA in both serum and liver without affecting HDV replication and clinical manifestations in patients[9,37]. In contrast, PC/BCP point mutations with HBeAg negative phenotype can significantly increase HBV viremia and replication of polymerase mutated strains in HDV-negative patients[38].
Other instances of indirect effects of HBV and HDV on each other’s replication include the synergistic activation of serum response element (SRE)-dependent pathways by HBxAg and L-HDAg, thus affecting factors which are involved in transcription regulation mediated by SRE[39]. From the cross-talks between HBV and HDV we can also refer to the NF-κB activation, which results from ER stress (induced by HBsAgs) or TNFα secretion from immune cells (in response to HBV infection) and correlates with L-HDAg nuclear export and HDV secretion[40].
CONCLUSION
The clinical observation of aggravated liver disease in patients with HBV/HDV co- or super-infection has prompted intense research on molecular interactions between both viruses. A major interaction between HBV and HDV is that they share a surface protein supply; this fact is currently being translated into novel therapeutic approaches using entry inhibitors in clinical trials for delta hepatitis[41]. However, in spite of the direct interaction sites between HBsAg and HDV-RNP, it seems that the key interference between HBV and HDV cannot be devoted to a certain domain or residue of the S protein but connection spots are rather distributed along the HBsAg. Moreover, besides the main reason for HBV/HDV interactions to share a surface protein supply, this is not the only interface between the two viruses in infected cells. Further investigations are required to unravel yet unknown molecular interactions that are employed by HBV or HDV to dominate in dual infections.
P- Reviewer: Rodriguez-Frias F, Rizzetto M S- Editor: Ji FF L- Editor: A E- Editor: Jiao XK