Copyright
©The Author(s) 2025.
World J Clin Oncol. Aug 24, 2025; 16(8): 107208
Published online Aug 24, 2025. doi: 10.5306/wjco.v16.i8.107208
Published online Aug 24, 2025. doi: 10.5306/wjco.v16.i8.107208
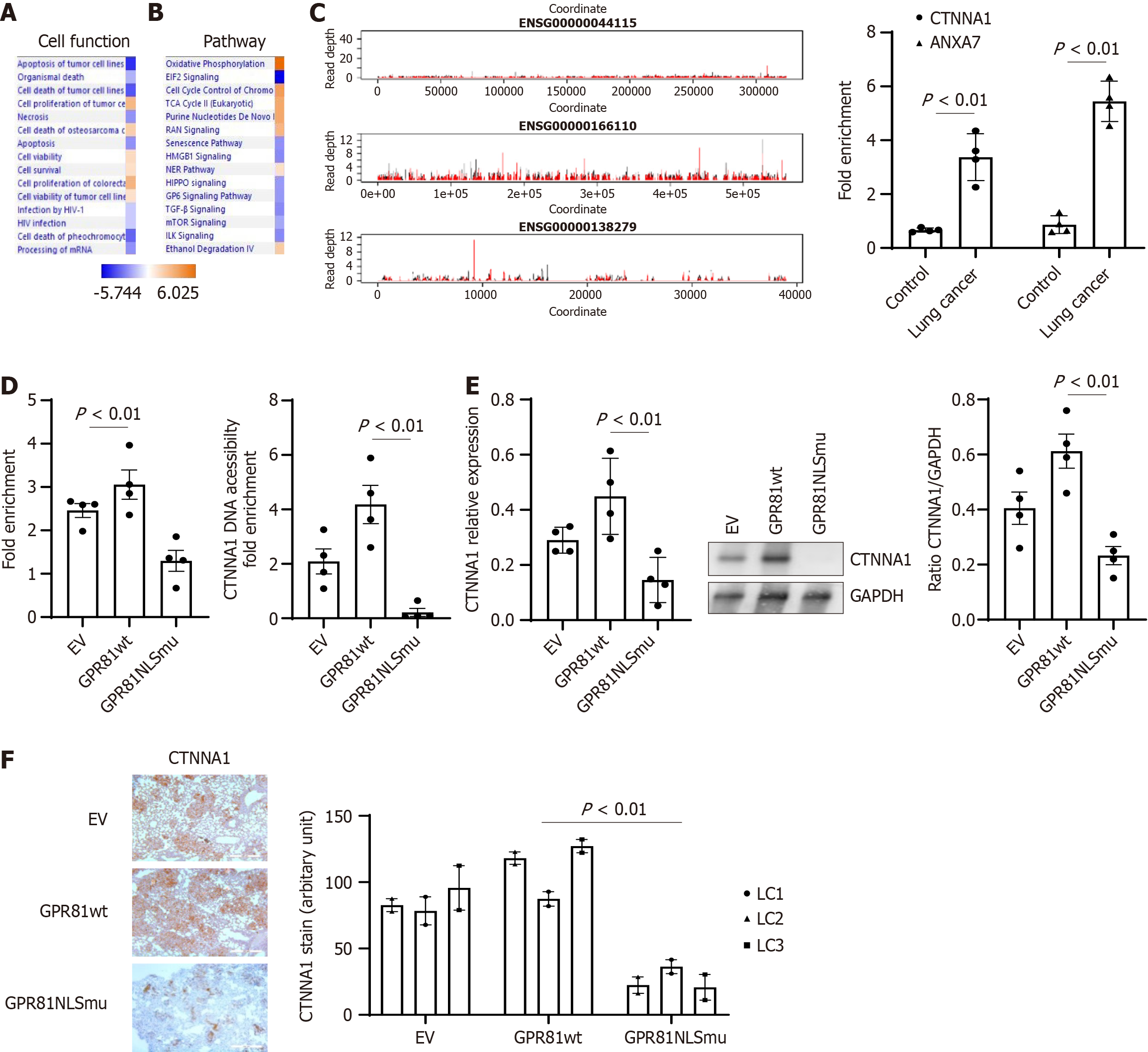
Figure 6 GPR81 interacts with genes and change gene expression.
Genes identified from GPR81 immunoprecipitated chromatin in lung cancer cells in Chromatin immunoprecipitation (ChIP) sequencing analysis were applied to Ingenuity Pathway Analysis (IPA). A: Top cell functions associated with the lung cancer cell dataset as shown by IPA pathway analysis; B: Top canonical pathways associated with proteins from the lung cancer cell dataset as shown by IPA pathway analysis. Cell functions, or pathways identified are represented on the Y-axis. The X-axis corresponds to the –log of the P-value (Fisher’s exact test) and the orange points on each pathway bar represent the ratio of the number of proteins in a given pathway that meet the cutoff criteria, divided by the total number of proteins that map to that pathway; C: The read depth plots for the GPR81 ChIP peaks of gene interest. Each plot shows the gene body. The red line shows the mean read depth across the +lactate samples and the black line shows the mean across the -lactate samples. The vertical blue lines show the boundaries of the gene body. Right panel: ChIP assays were performed on cross linked chromatin from nuclear fractions from control and lung cancer cells with CTNNA1 and ANXA7 primers; D: ChIP assays were performed on cross linked chromatin from nuclear fractions from GPR81 wt and GPR81 NLSmutant cells (n = 4 cell lines each) using immune GPR81 antibody (Isotype IgG as blank control) (left panel). Right panel: DNA Accessibility Assays were performed on cross linked chromatin from nuclear fractions from GPR81 wt and GPR81 NLSmutant cells (n = 4 cell lines A549, H661, H858, H645); E: CTNNA1 expression level was quantified with reverse transcriptase-PCR and western blot analysis in above set of cells; F: Anti-CTNNA1 was used to assess the distribution of human cells expressing CTNNA1 protein expressing cells from mice lung regions from mice receiving lung cancer cells transduced with empty vector, GPR81 wild-type (GPR81wt) (left panel) or GPR81 NLS mutant (GPR81NLSmu) (right panel). Scale bar = 200 μmol/L. Quantification was summed up in the bottom figure. Image J was used to quantify CTNNA1 protein positive cells in lung cancer cell explanted mice lung tissue sections.
- Citation: Yang L, Kono T, Gilbertsen A, Li Y, Sun B, Jacobson BA, Karam S, Dehm SM, Henke CA, Kratzke RA. GPR81 nuclear transportation is critical for cancer growth and progression in lung and other solid cancers. World J Clin Oncol 2025; 16(8): 107208
- URL: https://www.wjgnet.com/2218-4333/full/v16/i8/107208.htm
- DOI: https://dx.doi.org/10.5306/wjco.v16.i8.107208