Published online Jul 26, 2021. doi: 10.12998/wjcc.v9.i21.6102
Peer-review started: March 17, 2021
First decision: April 4, 2021
Revised: April 16, 2021
Accepted: May 19, 2021
Article in press: May 19, 2021
Published online: July 26, 2021
Immunohistochemical (IHC) staining for mismatch repair (MMR) proteins is useful for gastric cancer treatment and prognosis. Different IHC staining patterns reflect the complex biological phenomena underlying MMR deficiency. We herein report a rare IHC staining pattern of four MMR-related proteins in gastric cancer.
A “null” IHC staining pattern of four MMR-related proteins, including MLH1, PMS2, MSH2, and MSH6, in a 67-year-old male patient with gastric cancer pT3N3aM0 revealed promoter hypermethylation of MLH1. Next-generation sequencing showed that these four genes exhibited changes. One of these was the somatic mutation of the missing copy number in exon 14 of MSH2. Mutation analysis using peripheral blood showed no germline mutations in these four genes. The patient had no history of personal or family tumor history. We classified this case as sporadic. The patient returned to normal after operation, and there were no signs of tumor metastasis and recurrence. After six cycles of adjuvant chemotherapy, the patient was discharged in a stable condition. The patient had a mild reaction to chemotherapy and a good prognosis. At present, 16 mo after the operation, the patient's condition is stable.
Abnormal MMR protein expression, helpful for individualized follow-up care, helped identify a sporadic case lacking familial clinical management implications.
Core Tip: A “null” immunohistochemical staining pattern of mismatch repair (MMR) proteins was revealed, which helped identify this case as sporadic without familial clinical management implications. An in-depth understanding of the abnormal MMR expression is helpful for individualized follow-up treatment and assessment of pro
- Citation: Yue M, Liu JY, Liu YP. Unusual immunohistochemical “null” pattern of four mismatch repair proteins in gastric cancer: A case report. World J Clin Cases 2021; 9(21): 6102-6109
- URL: https://www.wjgnet.com/2307-8960/full/v9/i21/6102.htm
- DOI: https://dx.doi.org/10.12998/wjcc.v9.i21.6102
Gastric cancer is one of the most common cancers and a major cause of cancer-related deaths worldwide. Its molecular and clinical characteristics are complicated by histological and etiological heterogeneity. Gastric adenocarcinomas are divided into four subtypes according to their molecular features: Tumors exhibiting chromosomal instability, microsatellite instability (MSI)-high (MSI-H), Epstein–Barr virus (EBV) positivity, and genomic stability[1,2]. MSI and molecular typing are essential for the treatment and prognosis of gastric cancer[3,4].
Maintaining the mismatch repair (MMR) pathway is vital for accurate DNA replication and genome stability. Base pair insertion or loss occurs in the microsatellite region due to mismatch repair system defects, and replication errors cause MSI[5,6]. Immunohistochemistry (IHC) testing of four representative MMR-related proteins, MLH1, PMS2, MSH2, and MSH6, can detect a specific gene and help identify patients with Lynch syndrome (LS), an autosomal dominant disease caused by germline mutations in MMR-related genes[7]. IHC is a convenient and affordable method for MSI analysis and has a high sensitivity and specificity. Different IHC staining patterns reflect the complex biological phenomena underlying MMR deficiency. Therefore, it is essential to expand our understanding of abnormal IHC findings and their biological significance. In the present case, IHC testing of four MMR-related proteins revealed abnormal and rare protein expression with an unusual “null” pattern. These MMR-related genes exhibited changes tested by genetic analysis.
A 67-year-old male patient presented with upper abdominal discomfort and vomiting.
The patient experienced upper abdominal discomfort and vomiting for 4 mo. These events occurred without obvious inducement.
Helicobacter pylori infection was not observed. Left inguinal hernioplasty was performed 6 mo ago.
No personal or family history of tumors was noted.
No obvious abnormality.
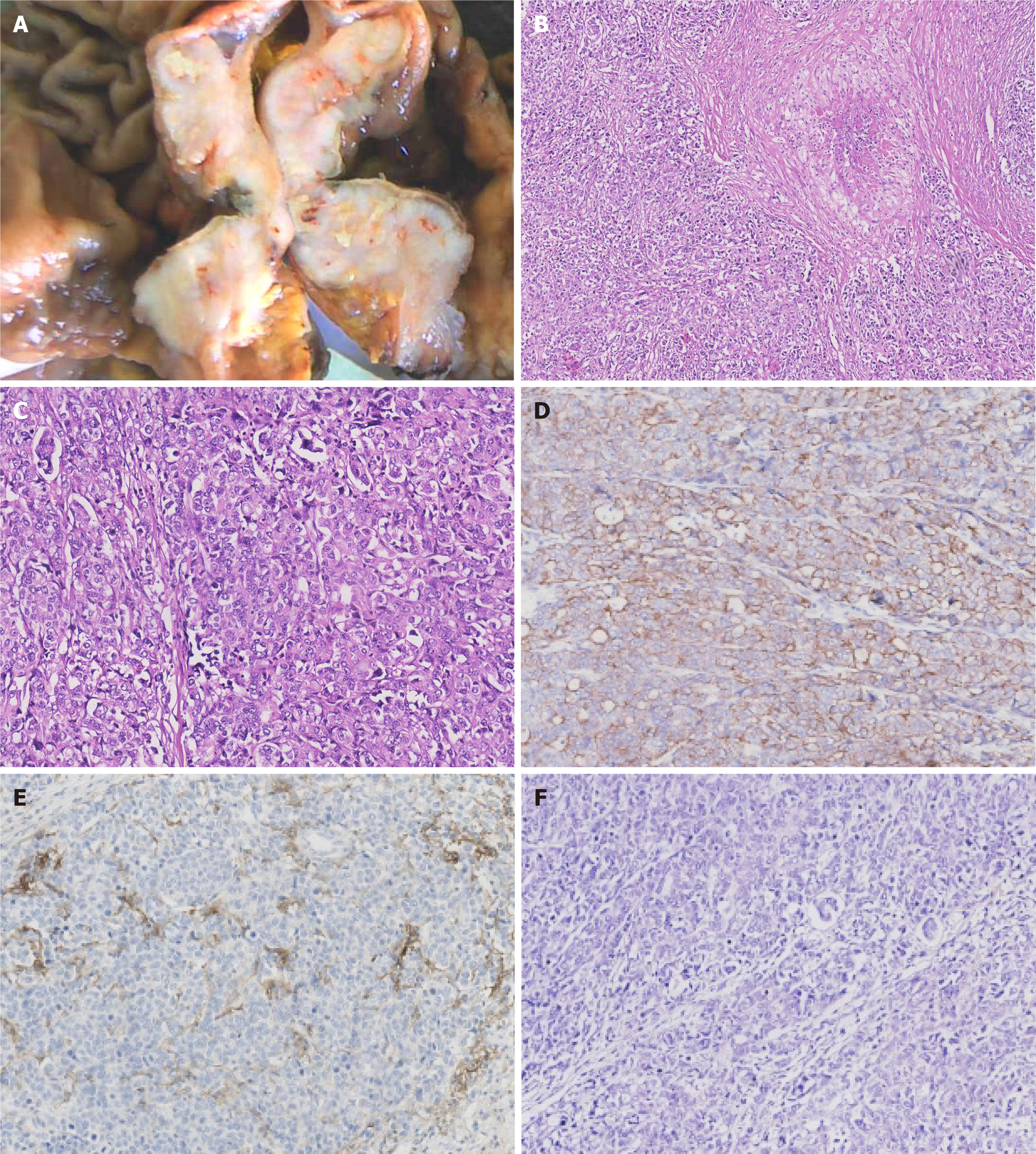
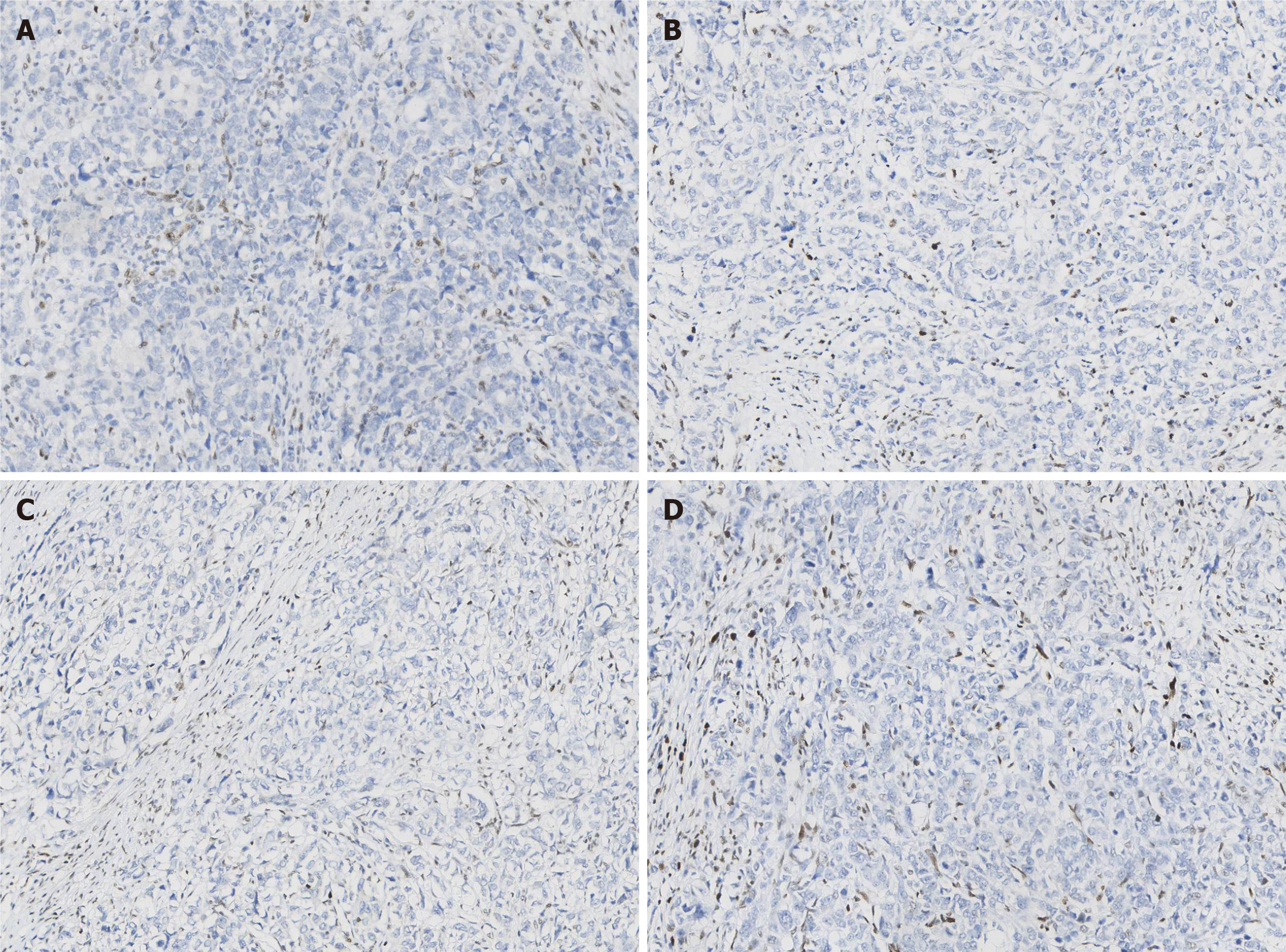
Paraffin-embedded blocks of gastric cancer tissues were used for pathologic diagnosis, IHC, and molecular analysis. The tumor showed an infiltrative growth pattern on gross examination (Figure 1A). Histologically, the tumor was poorly differentiated. It had a solid pattern with necrosis and was heterogeneous with glandular differentiation and prominent tumor-infiltrating lymphocytes (Figure 1B and C). The tumor cells were positive for low-molecular-weight cytokeratin (AE1/AE3, Ventana) (Figure 1D). The combined positive score (CPS) of programmed death-ligand 1 (PD-L1, 22C3, Dako) was high (Figure 1E). In situ hybridization of EBV-encoded small RNA was negative (Figure 1F). MMR proteins, including MLH1 (M1, Ventana), PMS2 (A16-4, Ventana), MSH2 (G219-1129, Ventana), and MSH6 (SP93, Ventana), were analyzed using IHC. All four proteins were negative in the tumor cells but positive in the positive control, stromal, and inflammatory cells. IHC results coincided with those of other paraffin blocks (Figure 2). HER2 (4B5, Ventana) was 1+, and Ki-67 (30-9, Ventana) was 60%.
Subsequently, a methylation-specific polymerase chain reaction assay of the MLH1 promoter region was performed, which revealed promoter hypermethylation of MLH1.
Next-generation sequencing (NGS) of genes including MLH1, MSH2, MSH6, and PMS2 was performed. In addition, 37 genes, including gastrointestinal tumor-related genes such as AKT1, ATM, BRCA1, BRCA2, CDH1, EGFR, ERBB2, HRAS, KIT, MET, PDGFRA, PIK3CA, PMS1, PTCH1, SDHB, SDHC, and SDHD; genes related to drug metabolism toxicity such as CYP2D6, DPYD, and UGT1A1; and genes related to gastrointestinal therapy, prognosis, and inheritance such as APC, BLM, BMPR1A, CHEK2, EPCAM, GALNT12, GREM1, MUTYH, POLD1, POLE, PTEN, SMAD4, STK11, TP53, KRAS, NRAS, and BRAF, were identified.
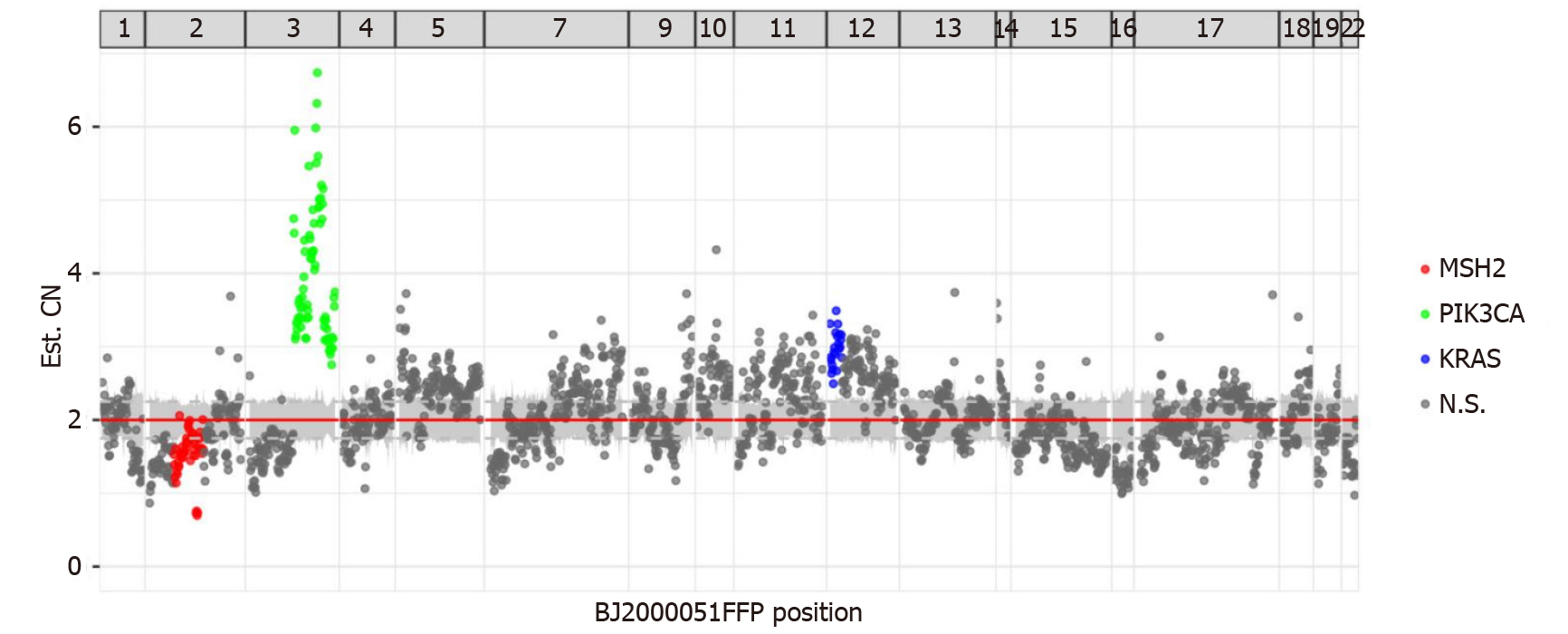
NGS demonstrated that MLH1, MSH2, MSH6, and PMS2 genes exhibited changes. There was a mutation in the splicing region of exon 12 of MLH1 (c.1039-13_1039-8del), a missense mutation in exon 11 of PMS2 (c.1799T>C, p.Met600Thr), a missing copy number in exon 14 of MSH2 (Figure 3), and a missense mutation in exon 4 of MSH6 (c.2693C>A, p.Pro898His). This mutation of MSH2 may lead to a shift in the subsequent coding frame of MSH2, leading to the loss of protein expression. Mutation analysis using peripheral blood showed no germline mutations in these four genes. The presence of MLH1 promoter methylation and mutation of MSH2 confirmed MSI-H. The patient had no personal or family tumor history, indicating that this MMR deficiency was highly likely sporadic in nature. Thus, there were no clinical manage
The tumor showed several other mutations, including copy number amplification of the PIK3CA gene (Figure 3), possibly leading to the upregulation of PIK3CA expression. The PTEN gene had a frameshift mutation in exon 7 (c.800del, p.Lys267fs) and a nonsense mutation in exon 5 (c.388C>T, p.Arg130). The exon began to shift from the amino acid residue 267 (lysine) in exon 7. Terminators are likely to be introduced into the new reading frame. The amino acid residue 130 encoded by exon 5 was mutated from arginine to the terminator. These two premature terminators may lead to meaningless mRNA degradation, resulting in protein loss. There was a missense mutation of p.g245s in exon 7 of the TP53 gene and mutations in the splicing region of exon 6 of the ATM gene (c.497-5_497-4del) and exon 12 of the MET gene (c. 2584-13_2584-9del).
Computed tomography showed space-occupying lesions in the gastric cardia and fundus and multiple slightly enlarged peripheral lymph nodes around the stomach.
Gastroscopy revealed a large mass in the gastric body. The endoscope could not pass through the gastric body.
Biopsy showed severe dysplasia and canceration of the glandular epithelium.
The results showed a 5-cm pT3N3aM0 adenocarcinoma. Furthermore, 7 of the 21 lymph nodes showed metastasis and a “null” IHC staining pattern of four MMR-related proteins, including MLH1, PMS2, MSH2, and MSH6. Thus, this case belongs to the MSI-H subtype of gastric adenocarcinoma.
Total gastrectomy was performed, and 21 lymph nodes were dissected. The patient returned to normal after operation. Nine months after the operation, the patient was admitted to the hospital for continued treatment. Ultrasound results showed several solid nodules in the patient's inguinal region, and biopsy results revealed normal lymph tissue. There were no signs of tumor metastasis or recurrence. After six cycles of adjuvant chemotherapy, the patient's response to chemotherapy was mild. The patient was satisfied with the treatment results.
The patient was asked to undergo follow-up every 3 mo. The patient was discharged in a stable condition for 16 mo after the operation. At present, the prognosis is good, but the long-term prognosis is yet to be determined.
Hagen et al[8] first reported this “null” IHC staining pattern of all four MMR proteins in colorectal cancer due to germline MSH2 mutation and somatic MLH1 hypermethylation in a 71-year-old female patient with LS. Using microscopy, the morphology observed in that case was similar to that observed in our case, but the gene expression was inconsistent. That patient had a history of colon cancer and ureteral cancer. She also had a strong family history of tumors. All these findings confirmed the diagnosis of LS. However, our case was sporadic.
Wang et al[9] found that the four MMR protein deficiencies in an 80-year-old female patient with colorectal cancer were highly likely sporadic, and no high-risk surveillance protocols were recommended to the patient or her family members. The morphology and gene expression observed in their case were similar to those observed in our case. They found promoter hypermethylation of MLH1 and double somatic truncating mutations in MSH2. They also found a BRAFV600E mutation. In our case, we found promoter hypermethylation of MLH1 and a missing copy number in exon 14 of MSH2. Westwood et al[10] found that the percentage of additional partial or complete loss of MSH2 and MSH6 in MLH1-deficient CRC patients was 0.48% (4/829). Additionally, IHC staining showed complete null expression of MMR proteins in only one colon cancer case, but it showed strong staining for all four proteins in rectal cancer. Unfortunately, biopsy was the only available procedure for testing in colon cancer. Further genetic analysis showed MLH1 promoter hypermethylation and BRAFV600E mutation. A summary of the “null” IHC staining pattern of all four MMR proteins reported in the literature is provided in Table 1.
| Case No. | Age (yr) | Sex | Location | MLH1 promoter methylation status | MSH2 status | BRAF status | Specimen tested | Ref. |
| 1 | 72 | Female | Colorectal cancer | Hypermethylation | Germline G587R mutation | Negative | Resection | Hagen et al[8], 2011 |
| 2 | 80 | Female | Colorectal cancer | Hypermethylation | Three somatic mutations (MSH2 c.1861 C>T (p.R621*) in exon 12, c.298G>A (p.V100I) and c.633dupG (p.K212Efs*20) | Mutation | Resection | Wang et al[9], 2017 |
| 3 | 76 | Male | Ascending colon | Hypermethylation | Unknown | Mutation | Biopsy | Westwood et al[10], 2019 |
MSI tumors with MLH1 methylation were associated with BRAFV600E mutation only in the colon but not in the stomach[11]. The BRAFV600E mutation was not detected in our case, which illustrates this point. Additionally, no abnormalities were found at the detection sites of KRAS, NRAS, and HRAS, which is also different from that in colon cancer.
All the above mentioned studies reported colorectal cancer cases but not gastric cancer cases. Cho et al[12] reported five cases in which all four MMR proteins were negative and none of 580 cases showed a single MMR protein loss by IHC; further genetic testing was not performed except for PCR analysis of MMR genes. In contrast, we retrospectively collected the data of 2808 cases of postoperative gastric cancer from the Fourth Hospital of Hebei Medical University from May 2017 to August 2020 and found that only this case was completely negative. Thus, the incidence rate was 0.0356%. Fifteen cases showed only negative results for PMS2, and three cases were negative for MSH6. The reasons for these differences remain unclear. The standard used by Cho et al[12] for judgement of negative results was complete loss or < 20% focal weak equivocal nuclear staining. We consider that our judgment standard was stricter; thus, the total number of negative cases was lower. Additionally, we tested more cases and identified cases with a single negative MMR protein. In another 464 cases of gastric cancer, the co-negative percentage of MLH1 and MSH2 was 4.4%[13]. These researchers used tissue microarrays for testing. However, there was potential heterogeneity in the use of tissue microarrays. In-depth genetic testing was not performed. Therefore, our case is the first to reveal this rare IHC staining pattern of four MMR proteins in gastric cancer and show differences in gene expression in colorectal cancer. We believe that this is not a common phenomenon in gastric cancer since it has not been previously reported in detail in the stomach.
Patients with MSI-H gastric cancer are usually older, with the majority being women. The cancer is mostly located in the distal part of the stomach; most of the cases are intestinal-type as per Lauren's classification and show less lymph node metastasis. They have a lower recurrence rate and better prognosis compared to those of other subgroups[1]. However, our patient was an aged man with cancer located near the cardia. Although the tumors were heterogeneous, the intestinal-type part only occupied a small portion of the tumor. Kawazoe et al[14] showed that PD-L1 expression (22C3) in immune cells (ICs) was associated with EBV positivity and lymph node metastasis. Our case showed the expression of PD-L1 only on ICs and high CPS with EBV negativity and lymph node metastasis. MSI status is considered a practical alternative to the immunotherapy response. Immunotherapy is recommended for patients with MSI-H[15]. However, our patient did not receive any immunotherapy, but the patient's condition remained stable after 9 mo, and there was no sign of recurrence or metastasis. Additionally, this patient had a large tumor size, many lymph node metastases, and specificity of gene expression. These findings are unusual compared with those in other cases; thus, the patient was asked to undergo follow-up regularly for safety precautions.
Here, we report a sporadic case with unusual MMR IHC and gene patterns in gastric adenocarcinoma. The patient had a mild reaction to chemotherapy and a good prognosis, and no clinical management implications for the family were identified. An in-depth understanding of the abnormal expression of MMR is helpful for individualized follow-up treatment and assessment of prognosis.
The authors thank all the clinicians and nurses involved in the treatment of this case.
Manuscript source: Unsolicited manuscript
Specialty type: Oncology
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Naem A, Villarejo-Campos P S-Editor: Gong ZM L-Editor: Wang TQ P-Editor: Wang LYT
| 1. | Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513:202-209. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 4230] [Cited by in F6Publishing: 4299] [Article Influence: 429.9] [Reference Citation Analysis (2)] |
| 2. | Katoh H, Ishikawa S. Genomic pathobiology of gastric carcinoma. Pathol Int. 2017;67:63-71. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 7] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 3. | Zappasodi R, Merghoub T, Wolchok JD. Emerging Concepts for Immune Checkpoint Blockade-Based Combination Therapies. Cancer Cell. 2018;33:581-598. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 311] [Cited by in F6Publishing: 328] [Article Influence: 54.7] [Reference Citation Analysis (0)] |
| 4. | Zhao X, Dai D, Li X, Shen B, Chen X, Shu Y, Wang D. A polymorphism within the mismatch repair gene predicts prognosis and adjuvant chemotherapy benefit in gastric cancer. Gastric Cancer. 2019;22:1121-1129. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 9] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 5. | Liu D, Keijzers G, Rasmussen LJ. DNA mismatch repair and its many roles in eukaryotic cells. Mutat Res. 2017;773:174-187. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 97] [Cited by in F6Publishing: 113] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 6. | Lee V, Murphy A, Le DT, Diaz LA Jr. Mismatch Repair Deficiency and Response to Immune Checkpoint Blockade. Oncologist. 2016;21:1200-1211. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 149] [Cited by in F6Publishing: 176] [Article Influence: 22.0] [Reference Citation Analysis (0)] |
| 7. | Kawakami H, Zaanan A, Sinicrope FA. Microsatellite instability testing and its role in the management of colorectal cancer. Curr Treat Options Oncol. 2015;16:30. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 226] [Cited by in F6Publishing: 257] [Article Influence: 32.1] [Reference Citation Analysis (0)] |
| 8. | Hagen CE, Lefferts J, Hornick JL, Srivastava A. "Null pattern" of immunoreactivity in a Lynch syndrome-associated colon cancer due to germline MSH2 mutation and somatic MLH1 hypermethylation. Am J Surg Pathol. 2011;35:1902-1905. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 25] [Cited by in F6Publishing: 22] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 9. | Wang T, Stadler ZK, Zhang L, Weiser MR, Basturk O, Hechtman JF, Vakiani E, Saltz LB, Klimstra DS, Shia J. Immunohistochemical null-phenotype for mismatch repair proteins in colonic carcinoma associated with concurrent MLH1 hypermethylation and MSH2 somatic mutations. Fam Cancer. 2018;17:225-228. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 17] [Cited by in F6Publishing: 16] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 10. | Westwood A, Glover A, Hutchins G, Young C, Brockmoeller S, Robinson R, Worrilow L, Wallace D, Rankeillor K, Adlard J, Quirke P, West N. Additional loss of MSH2 and MSH6 expression in sporadic deficient mismatch repair colorectal cancer due to MLH1 promoter hypermethylation. J Clin Pathol. 2019;72:443-447. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 13] [Cited by in F6Publishing: 10] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 11. | Liu Y, Sethi NS, Hinoue T, Schneider BG, Cherniack AD, Sanchez-Vega F, Seoane JA, Farshidfar F, Bowlby R, Islam M, Kim J, Chatila W, Akbani R, Kanchi RS, Rabkin CS, Willis JE, Wang KK, McCall SJ, Mishra L, Ojesina AI, Bullman S, Pedamallu CS, Lazar AJ, Sakai R; Cancer Genome Atlas Research Network; Thorsson V, Bass AJ, Laird PW. Comparative Molecular Analysis of Gastrointestinal Adenocarcinomas. Cancer Cell 2018; 33: 721-735. e8. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 337] [Cited by in F6Publishing: 318] [Article Influence: 53.0] [Reference Citation Analysis (0)] |
| 12. | Cho J, Kang SY, Kim KM. MMR protein immunohistochemistry and microsatellite instability in gastric cancers. Pathology. 2019;51:110-113. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 13] [Cited by in F6Publishing: 20] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 13. | Bae YS, Kim H, Noh SH. Usefulness of Immunohistochemistry for Microsatellite Instability Screening in Gastric Cancer. Gut Liver. 2015;9:629-635. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 25] [Cited by in F6Publishing: 28] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 14. | Kawazoe A, Shitara K, Kuboki Y, Bando H, Kojima T, Yoshino T, Ohtsu A, Ochiai A, Togashi Y, Nishikawa H, Doi T, Kuwata T. Clinicopathological features of 22C3 PD-L1 expression with mismatch repair, Epstein-Barr virus status, and cancer genome alterations in metastatic gastric cancer. Gastric Cancer. 2019;22:69-76. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 30] [Cited by in F6Publishing: 33] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 15. | Chivu-Economescu M, Matei L, Necula LG, Dragu DL, Bleotu C, Diaconu CC. New therapeutic options opened by the molecular classification of gastric cancer. World J Gastroenterol. 2018;24:1942-1961. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 29] [Cited by in F6Publishing: 32] [Article Influence: 5.3] [Reference Citation Analysis (0)] |