Published online Nov 24, 2020. doi: 10.5306/wjco.v11.i11.868
Peer-review started: March 29, 2020
First decision: June 20, 2020
Revised: July 29, 2020
Accepted: November 4, 2020
Article in press: November 4, 2020
Published online: November 24, 2020
Epithelial ovarian cancer (EOC) is the most lethal gynaecological malignancy in the western world. The majority of women presenting with the disease are asymptomatic and it has been dubbed the “silent killer”. To date there is no effective minimally invasive method of stratifying those with the disease or screening for the disease in the general population. Recent molecular and pathological discoveries, along with the advancement of scientific technology, means there is a real possibility of having disease-specific liquid biopsies available within the clinical environment in the near future. In this review we discuss these discoveries, particularly in relation to the most common and aggressive form of EOC, and their role in making this possibility a reality.
Core Tip: Epithelial ovarian cancer (EOC), particularly high-grade serous carcinoma, is a gynaecological malignancy with a poor survival rate. Currently there is no effective disease-specific biomarker, which could improve detection rates and treatment algorithms, for any of the EOC types - this is an area of unmet clinical need. Circulating tumour DNA (ctDNA) analysis has emerged as a potential blood-based “liquid biopsy” for early detection, diagnosis, staging and prognosis, monitoring response to treatment, monitoring minimal residual disease and relapse and identifying acquired drug resistance mechanisms. However, there are several obstacles to the development of cfDNA-based biomarkers which are discussed further in this in-depth review.
- Citation: Feeney L, Harley IJ, McCluggage WG, Mullan PB, Beirne JP. Liquid biopsy in ovarian cancer: Catching the silent killer before it strikes. World J Clin Oncol 2020; 11(11): 868-889
- URL: https://www.wjgnet.com/2218-4333/full/v11/i11/868.htm
- DOI: https://dx.doi.org/10.5306/wjco.v11.i11.868
Epithelial ovarian cancer (EOC) is the most lethal gynaecological malignancy in the western world. In 2012 there were 152000 and 4300 deaths from ovarian cancer worldwide and in the United Kingdom, respectively[1]. This equates to twelve women dying from ovarian cancer within the United Kingdom every day.
The majority of women presenting with EOC are asymptomatic. In those women that do experience symptoms they are often vague and non-specific. The nature of the symptoms means that women generally present to their doctor with advanced stage disease. This has led to ovarian cancer being termed the “silent killer”. There have been major publicity campaigns, both regionally and nationally, involving the Department of Health and cancer charities to increase the level of awareness of ovarian cancer symptomatology; in an effort to improve earlier diagnosis.
There are five main types of EOC and these are included in the current World health Organization (WHO) classification: High-grade serous carcinoma (HGSC, 70%), endometrioid carcinoma (EC, 10%), clear cell carcinoma (CCC, 10%), mucinous carcinomas (MC, 3%), and low-grade serous carcinomas (LGSC, 3%)[2]. HGSC accounts for approximately 70% of all EOCs and approximately 90% of advanced stage EOCs (FIGO stage III-IV), making it the most common and most deadly.
The EOC types differ significantly morphologically, clinically, and at a molecular level. LGSC exhibit KRAS, BRAF, and ERBB2 mutations in around two thirds of cases whereas TP53 mutations are very rare in these tumours[3,4]. CTNNB1 (encoding β-catenin) and PTEN mutations, along with microsatellite instability, are associated with low-grade ECs via specific signalling pathways[5,6]. MC display KRAS mutations in more than 50% of specimens and identical mutations have been elucidated in benign, borderline and malignant areas from the same neoplastic lesion suggesting that a KRAS mutation is an early event in mucinous tumour pathogenesis[7]. EC and CCC have been shown to be associated with endometriosis in the ovary or pelvis in 15% to 70% of cases; this is almost certainly an underestimate since endometriosis may be overgrown by the tumour and it is likely that a large majority of EC and CCC arise in endometriosis[8,9]. In fact, there is recent evidence to show that long interspersed element-1 hypomethylation is an early molecular event involved in the transformation of EC and CCC from endometriosis[10]. The tumour suppressor gene, AT-Rich Interaction Domain 1A (ARID1A) also seems to play a role in early malignant transformation of endometriosis to EC or CCC[11-13]. Mutations in this gene have been identified in up to 50% of CCCs and approximately one third of ECs[12].
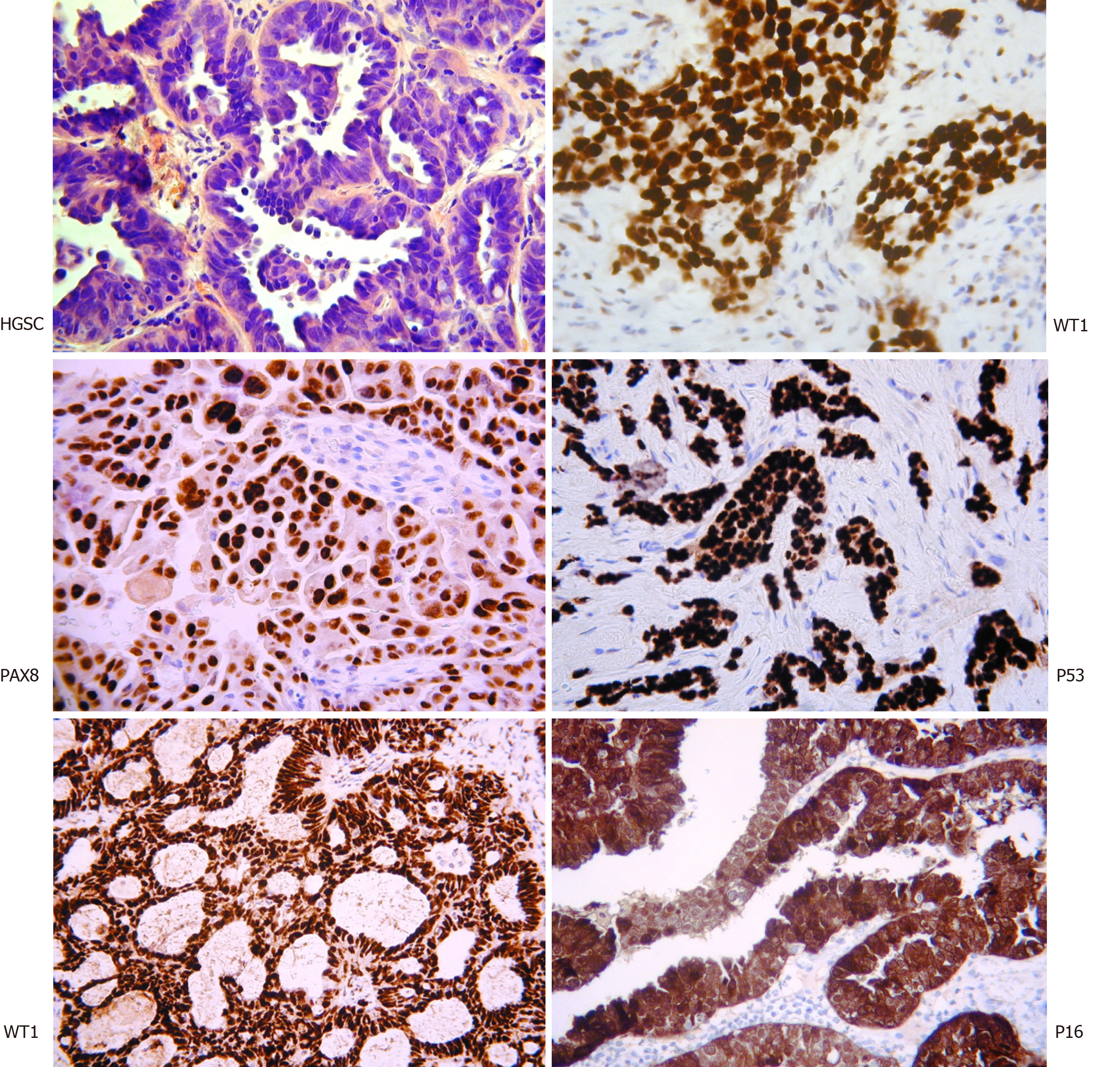
HGSC differs significantly from the other subtypes. The classical histological appearance is of intermediate sized tumour cells, marked nuclear atypia, and necrotic areas (Figure 1). Immunostaining with WT1, PAX8, P16, and P53 assist with the diagnosis. Interestingly, WT1 staining helps discriminate between HGSC and pseudo-endometrioid. Molecularly, it is characterised by the ubiquitous presence of TP53 mutations and CCNE1 gene (encoding cyclin E1) amplification in 20% of cases[14-19]. It is, however, rarely associated with mutations such as KRAS, BRAF, ERBB2, HER2, PTEN, CTNNB1, ARID1A and PIK3CA[6,7,14,15,16,19]. Germline mutations in the BRCA1 or BRCA2 genes are present in 6.5%-19% of HGSCs and a smaller proportion have somatic mutations[20]. Whilst BRCA1 somatic mutations are rare in sporadic disease (< 10%), BRCA1 is reported to be downregulated in 15%-72% via mechanisms of epigenetic inactivation[20]. To complicate things further, the HGSC subtype is also highly molecularly heterogeneous, with the possible inclusion of further subpopulations that display distinct gene expression profiles and variable responses to current chemotherapy regimens[21].
The diagnosis of women with symptoms suspicious of EOC utilises the biomarker serum cancer antigen 125 (CA125), which was first described by Bast et al[22] in 1981. It was identified by the murine monoclonal antibody OC-125 as an antigenic determinant on a high molecular-weight glycoprotein. In adults, CA125 is expressed in tissues derived from both coelomic and Mullerian epithelia. It is also expressed by epithelia of the pancreas, colon, gall bladder, lung, kidney, and stomach[23].
In clinical practice, serum CA125 and preliminary radiological imaging, in the form of an abdomino-pelvic ultrasound, results are used to calculate the risk of malignancy index as a method of triaging patients for tertiary/quaternary referral[24]. Although it is used to aid diagnosis, only 50% of early stage EOCs show expression[23,25].
CA125 is most effective as a marker of disease status in patients undergoing chemotherapy treatment for EOC[26,27]. Unfortunately, it is not specific to malignancy, expressed by several other benign conditions including diverticulitis, endometriosis, liver cirrhosis, uterine fibroids, menstruation, pregnancy, pelvic infection, and uterine leiomyomata[28-33]. It is also elevated by other malignancies such as pancreatic, bladder, breast, liver, and lung cancers[33].
With earlier diagnosis the key to improved survival from EOC, there has been considerable effort to identify alternative biomarkers or develop combination markers with CA125, without overwhelming success[34]. To improve the sensitivity of CA125 the Risk of Ovarian Cancer Algorithm (ROCA) was developed[35]. This algorithm compares the CA125 level of cases with a profile of healthy women. It calculates a risk estimate based on this comparison and the age-specific incidence of EOC. The algorithm was employed within the UKCTOCS trial and various others with varying degrees of success[36-38].
The only novel biomarker that has showed promise, since CA125, is human epididymis protein 4 (HE4)[39]. This biomarker has been shown to improve diagnostic accuracy, both alone and in combination with CA125[29,40-42]. A recent meta-analysis looked at diagnostic performance with two control groups (healthy women and women with benign gynaecological disease)[40]. In the analysis vs “healthy women” the sensitivity and specificity for HE4 in diagnosing ovarian cancer were 83% (95%CI: 77%-88%) and 90% (95%CI: 87%-92%), respectively. Receiver operator characteristic (ROC) analysis revealed an area under the curve (AUC) of 0.9271. In the women with benign disease analysis, the sensitivity and specificity were 74% (95%CI: 69%-78%) and 90% (95%CI: 87%-92%) respectively. The ROC analysis AUC was 0.8853. This suggests HE4 carries potential as an early-warning biomarker. Contrastingly, another meta-analysis looking at CA125 and HE4 combined diagnostic performance showed HE4 to be no better than CA125 for predicting EOC[43].
A panel of biomarkers that included CA125, HE4, transthyretin, CA15.3, and CA72.4 was evaluated using specimens assembled from multiple cohort and randomised trial[44]. Phase II and III biomarker studies concluded that CA125 remained the “single-best biomarker” for EOC. Another retrospective study evaluated seven proteomic biomarkers (apolipoprotein A1, truncated transthyretin, transferrin, hepcidin, beta-2 microglobulin, connective tissue activating protein III, and inter-alpha-trypsin inhibitor heavy-chain) in pre-diagnostic blood samples[45]. The addition of the seven protein biomarkers to CA125 did not improve sensitivity compared to CA125 alone.
The combination of CA125 and HE4 was developed into an algorithm in an effort to improve EOC detection[46]. The Risk of Ovarian Malignancy Algorithm (ROMA) successfully classified 93.8% of EOC patients as high risk. The model has been assessed in a range of populations with varying degrees of accuracy[42,47-53]. It is for this reason that HE4 or ROMA has not translated into routine clinical care, with some groups requesting further in-depth validation.
The evidence that population screening impacts cancer-specific mortality is ever-accumulating. There are examples of successful cancer screening programmes across many countries, particularly for breast, bowel, and cervical cancers. A significant reduction in mortality has been seen in the latter[54]. The real success of these screening programmes is the understanding of the pathogenesis of the disease. Currently there is no effective screening method for ovarian cancer and the United States Preventative Services Task Force (USPSTF) maintains its position that it should not be undertaken, in any form[55].
Ovarian cancer is relatively rare with a low prevalence in the general population. An effective and acceptable screening strategy must have not only a high sensitivity for early-stage disease (> 75%) but must also have a very high specificity (99.6%) to achieve a positive predictive value (PPV) of 10%. In real terms, that equates to a threshold of no more than 10 staging laparotomies to identify one ovarian cancer[56].
The main methods of ovarian cancer screening assessed to date are pelvic ultrasound scanning and serially measuring the serum biomarker, CA125. Pelvic ultrasound in expert hands is a highly sensitive diagnostic method[57]. Unfortunately, because it relies heavily on individual expertise, discrimination between benign and malignant pelvic masses in routine clinical practice is challenging. Serum CA125 is most effective as a marker of disease status in patients undergoing chemotherapy treatment for EOC[26]. It is not particularly specific to malignancy and can be expressed by a number of other benign conditions including endometriosis, pelvic infection, and uterine leiomyomata. It is elevated in only 50% of early stage EOCs, and thus can cause unnecessary medical intervention and significant patient distress[25].
There have been a few large prospective trials examining ovarian cancer screening. Each utilised different screening strategies but all aimed to assess the same primary outcome: Mortality.
The United States Prostate Lung Colorectal Ovarian Cancer Screening Trial (USPLCO) comprised over 78000 women aged 55-74 years in two arms; annual transvaginal ultrasound and serum CA125 vs routine clinical care[58]. The study showed no difference in mortality between the two cohorts. The United Kingdom Collaborative Trial in Ovarian Cancer Screening (UKCTOCS) is the largest screening trial to date, enrolling over 200000 postmenopausal women randomised into no intervention or quarterly (initially annual) screening[38]. The screening cohort was broken down into transvaginal ultrasound alone or serum CA125 screening, interpreted by the ROCA algorithm, with second-line transvaginal ultrasound (termed multimodal screening). UKCTOCS finally reported mortality data in 2016. At a median follow-up of 11.1 years, ovarian cancer was diagnosed in 1282 (0.6%) women. There was no difference across subgroups with 0.7% in the multimodal screening group, 0.6% in the ultrasound only group, and 0.6% in the control group. Of these women, 0.29% in the MMS group, 0.30% in the USS group, and 0.34% in the control group died of ovarian cancer. The primary analysis revealed a non-statistically significant mortality reduction of 15% (P = 0.10) with MMS and 11% (P = 0.21) with USS alone.
Although the primary end point did not reach statistical significance, further analysis suggested improved mortality reductions for the MMS group compared to the control group after 7 years; 8% reduction for the first 7 years compared to 23% for years 7 to 14; suggesting a “late effect” mortality benefit[38]. In response to the “late effect” mortality trend proposed by the UKCTOCS study, extended mortality results were reported for the USPLCO trial with a median 15 year follow up (range 13 to 19 years)[59]. However, following this extended analysis, the study reiterated its initial findings and reported no change in the mortality benefit of screening using CA125 and TVUS.
To date there have been no disease-specific biomarkers validated for any of the five main EOC types. Moving forward, this approach would be more appropriate given that these represent different tumour types with a different pathogenesis and behaviour and require different management. Currently, the spotlight is on HGSC as it is the most frequent and most aggressive type of EOC.
For some time, scientists and clinicians have been unable to identify a pre-invasive stage to HGSC, and the disease did not appear to fit this model. This is likely explained by the fact the disease spreads so aggressively quite early in its course, making the pathological detection of early stage disease elusive[60]. There has been considerable effort employed to define the molecular mechanisms of HGSC and, until recently, its pathogenesis remained undefined. Historically, it was thought HGSC developed from the ovarian surface epithelium (OSE) due to errors in cell replication associated with the repair of trauma incurred by ovulation[61]. Several epidemiological studies supported this theory with evidence that women with an increased number of lifetime ovulations are at a much greater risk of developing HGSC[62-67].
However, in recent years, overwhelming pathological evidence has emerged that supports the theory that the distal fallopian tube (fimbria) is the origin of HGSC[68]. The fact that fallopian tube epithelium is embryologically mullerian and HGSCs are mullerian in nature suggests a potential site of origin here for HGSC. Furthermore, malignant transformation of fallopian tubal epithelium yields almost exclusively HGSC[69]. HGSC occurring concurrently with fallopian tubal mucosal disease was first documented by Bannatyne et al[70] but this was interpreted to represent a second primary tumour focus rather than being directly related to the ovarian disease. Subsequently a number of research groups acknowledged that there was an increased risk of primary fallopian tubal HGSC in BRCA1/2 mutation carriers[71-73]. Zweemer et al[74] presented the first evidence of primary tubal HGSC in BRCA1 mutants and, subsequently, a large prospective study of BRCA1 mutation carriers revealed a 120-fold increased risk, compared to the general population, of primary fallopian tube carcinoma[75]. In 2006, Finch et al[76] published clinical and pathological findings of prophylactic salpingo–oophorectomy specimens from 159 BRCA1/2 mutation carriers. Seven (4.4%) occult fallopian tube cancers were identified in these women, in the absence of symptoms. Multiple other investigators also recorded the presence of occult tubal tumours at prophylactic salpingoophorectomy[76-83]. The incidence of these occult lesions ranged from 6%-40% with up to 100% of them occurring in the fallopian tube, usually in the absence of ovarian involvement. Further investigation has revealed most HGSCs arise from the distal fallopian tube via an in-situ carcinomatous lesion referred to as serous tubal intraepithelial carcinoma (STIC)[84,85].
There is now compelling evidence from both our group and others worldwide that most, but not all, HSGCs arise from the distal fallopian tube, and not the ovary, from STIC[86-88]. Detailed assessment of the FTs in cases of extrauterine HGSC shows involvement of the fimbriae in 70% and STIC in approximately 50% of the cases[89]. At a molecular level; Cyclin E1 (CCNE1), remodelling and spacing factor 1 (Rsf-1), and fatty acid synthase (FASN) are all upregulated in HGSC. Rsf-1 encodes an important ATP-dependent chromatin remodelling protein which is an integral component of DNA replication and cell cycle progression[90,91]. FASN is a cytoplasmic enzyme involved in tumour initiation and progression[92,93]. These oncogenes are all overexpressed in STIC lesions[94,95]. Another oncogene integral to this disease process is TP53[96]. Mutation of this tumour suppressor is known to be associated with HGSC and recent evidence shows it be characteristic of HGSC and essentially ubiquitous[15,97,98]. The fact that STIC lesions and concurrent HGSC have been shown to contain identical TP53 mutations further indicates a clonal relationship between the two entities[98,99]. Gene expression profiling of a unique sample set containing normal OSE, normal FT, STIC, ovarian HGSC, and omental metastases was performed by our group. Bioinformatic analysis revealed that the tumour samples clustered in one cohort and normal FT samples clustered together and separately from the normal OSE samples. Notably, the normal OSE samples clustered separately from all other profiled samples and STIC samples clustered within the tumour cohort. Multi-dimensional scaling analysis confirmed the strong common biology present between STIC and the two tumour groups. It also affirmed that OSE has no significant common genetic biology to the other samples. This study further confirms the fallopian tubal origin of HGSC[86].
The proportion of HGSCs derived from the FT is currently unknown, mainly due to the fact HGSC usually presents at an advanced stage making the detection of precursor lesions difficult and depends on the criteria used to determine a tubal primary. However, in the most detailed prospective study, using criteria for site assignment of extrauterine HGSC adopted by the International Collaboration on Cancer Reporting (ICCR), 83% of cases were determined to be of tubal origin and almost all the remainder of ovarian origin with a primary peritoneal origin being extremely rare when strict criteria are used[100-103].
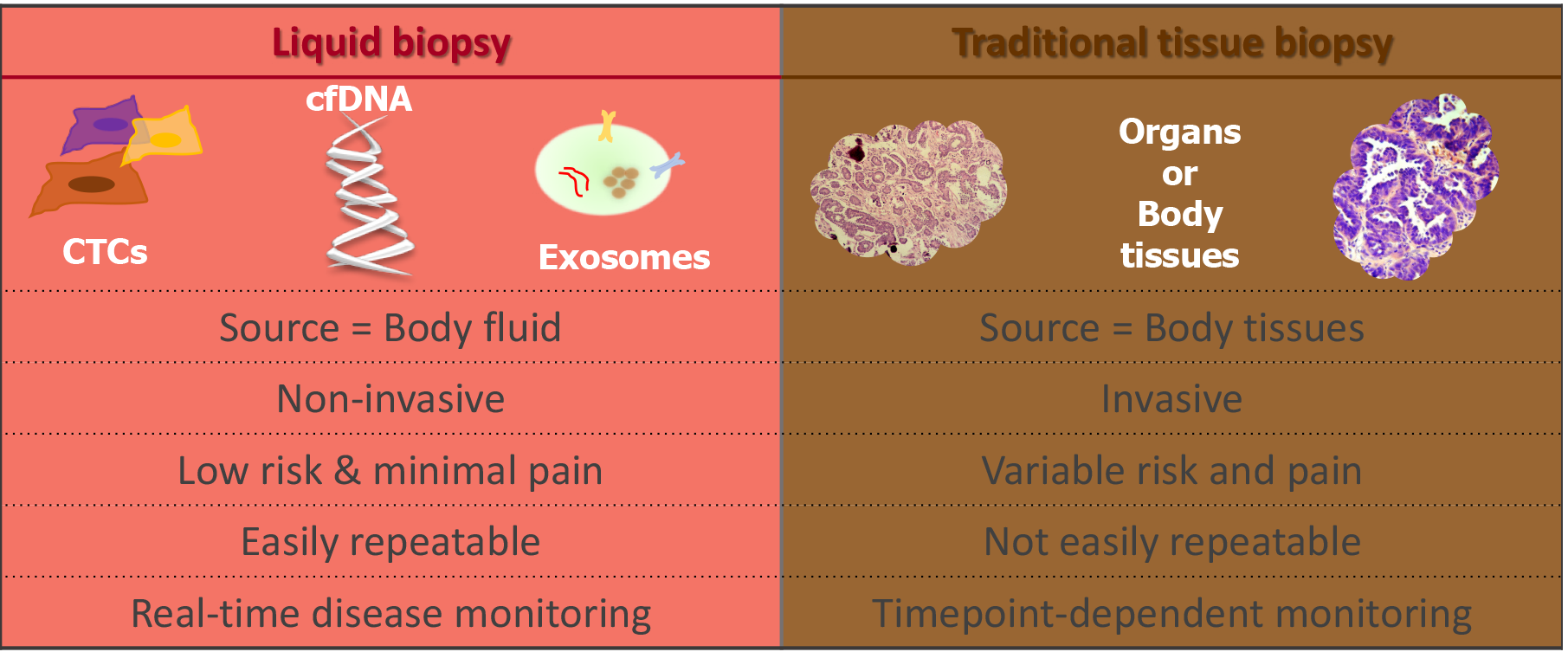
Precision oncology seeks to obtain molecular information about cancer to improve patient outcomes. Tissue biopsy samples are widely used to characterise tumours; however, this method of tumour analysis has limitations. The term “liquid biopsy” was first used to describe methods that can derive the same diagnostic information from a blood sample, or other body fluid, that is typically derived from a tissue biopsy sample[104]. In recent years, the focus of precision medicine is increasingly turning towards liquid biopsies as they are minimally invasive and can be repeated at multiple time points facilitating “real-time” disease monitoring[105].
Liquid biopsy (Figure 2) can include measurement of soluble factors, such as circulating tumour nucleic acids (DNA/RNA), circulating tumour cells (CTCs), proteins, and extracellular vesicles such as exosomes. All of which have been investigated for potential as diagnostic, predictive and prognostic biomarkers[106].
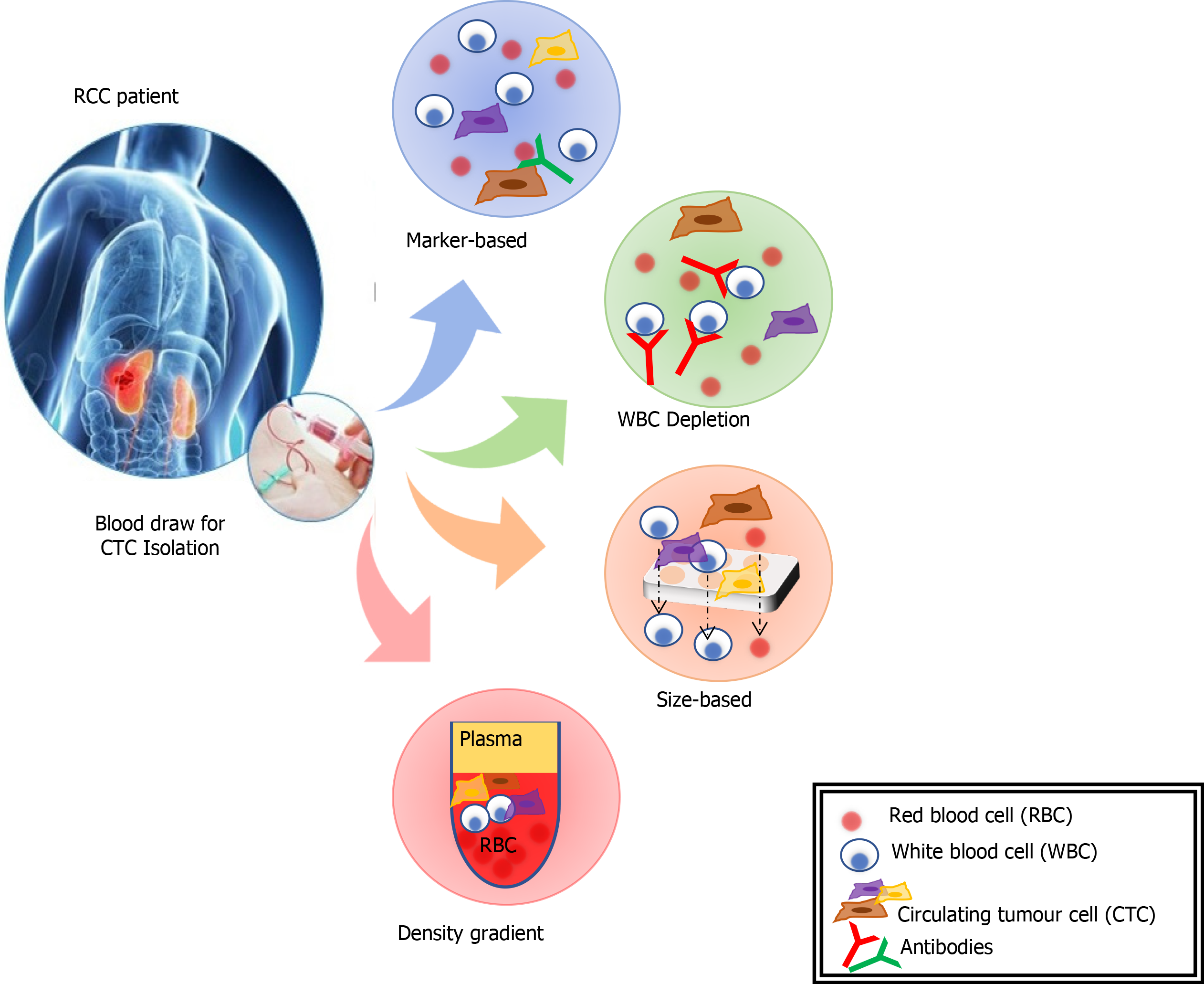
CTCs are cells originating from a solid tumour that are detectable in the peripheral blood. They are considered a prerequisite step in establishing distant metastases[107]. The detection of CTCs in peripheral blood (Figure 3) is a novel type of cancer biomarker[108]. CTCs can be isolated from blood samples and used to follow patients over time. They can provide significant information that will better characterise underlying disease biology and metastases. However, CTCs are rare, and their isolation, quantification and molecular characterization carry many challenges. An average metastatic carcinoma patient has between 5 and 50 CTCs for every 7.5 mL of blood[109]. This places technical limitations on the ability to identify and characterise the sub-population of cells that carry the relevant genetic information. CTCs tend to aggregate with leucocytes so adequate cell-surface markers and separation techniques need to be utilised to improve purity[109].
CTCs were first detected on the background of malignant melanoma and have subsequently been identified associated with a range of solid tumours[110]. There are a range of approaches for isolation of CTCs including; mRNA-based, protein-based, and cell size-based. The mRNA-based strategy involves RqPCR techniques. This carries some limitations including: (1) Amplification of non-specific products, (2) Lack of validated protocols for sample processing, RNA-preparation, cDNA synthesis and PCR conditions, and (3) A lack of sample quality control measures[107]. These issues all raise the possibility of variations in sensitivity and specificity of a biomarker employing this method. Separation and enrichment of CTCs, using magnetic bead technologies, followed by flow cytometry or immunohistochemistry is another method of quantification and characterisation of CTCs[106]. This method eliminates some of the concerns with RqPCR techniques, but further research needs to be done to clarify the reproducibility of different techniques. The isolation by size of epithelial tumour cells is a direct method for CTC identification and has been applied in various epithelial cancers[107]. The CTCs are collected by filtration and, following staining for specific markers, the cells are identified and quantified by immunohistochemistry or molecular pathological techniques.
The only clinically validated, FDA-approved blood test for CTCs is the CELLSEARCH® CTC Test (Janssen Diagnostics, New Jersey, United States). It has shown promise as a prognostic indicator in prostate, colorectal, and breast cancers[111-113]. This technology was assessed within the EOC population and while it did provide evidence that ovarian CTCs were present in blood it did not correlate with clinical outcomes[113]. A further study assessed a unique cell adhesion matrix based, functional cell enrichment and identification platform[114]. There was clear evidence that elevated CTCs correlated with worse OS and PFS. In fact, this method proved more specific than CA125 in detecting EOC malignancy in high-risk patients. A recent meta-analysis assessed eleven studies comprising a total of 1129 patients[115]. It reported CTC status to be a significant prognostic indicator (OS:HR 1.61, 95%CI: 1.22-2.13; PFS:HR 1.44, 95%CI: 1.18-1.75). A subgroup analysis showed the RqPCR methodology to be superior to both CELLSEARCH® and immunohistochemical methods.
Despite the potential role of CTCs in cancer diagnostics, CTC methods are generally used for research purposes, and only a few methods have been accepted for clinical application. This is because of the difficulties caused by CTC heterogeneity, CTC separation from blood, and a lack of thorough clinical validation[116].
In recent years there has been a significant amount of research into the use of circulating cell free DNA (cfDNA) as a biomarker. The presence of circulatory cfDNA was identified over seventy years ago[117]. Along with technological evolution, subsequent research has demonstrated that cancer cells release cfDNA fragments into the circulation and other bodily fluids, termed circulating tumour DNA (ctDNA), and these fragments carry all the genetic and epigenetic characteristics of the primary tumour[118]. Fragments of cfDNA in blood samples are between 150 and 200 bp long, of which up to 90% originates from the tumour[118]. It is reported that tumours containing approximately 50 million malignant cells release enough DNA for the detection of tumour cfDNA in blood[119]. Of note, this is well below the limit of resolution of radiological imaging (approximately 1 billion cells)[120].
As a result, ctDNA analysis has emerged as a potential blood-based “liquid biopsy” for early detection, diagnosis, staging and prognosis, monitoring response to treatment, monitoring minimal residual disease and relapse and identifying acquired drug resistance mechanisms.
There are several hypotheses as to the origin and mechanism of release of cfDNA/ctDNA into the circulation; however, the precise mechanism(s) have yet to be determined. Early studies suggested that ctDNA enters the circulation following lysis of cells on the interface between the tumour and circulation[121]. Another theory proposed that ctDNA may originate from the destruction of tumour micro metastases and circulating cancer cells[122]. Current consensus suggests that most cfDNA in healthy individuals is released from the bone marrow and white blood cells, whereas ctDNA in cancer patients is derived from necrotic and apoptotic cancer cells. Three possible mechanisms resulting in the shedding of DNA from both healthy and tumour cells have been described: Apoptosis, necrosis, and active cellular release[123]. Apoptosis causes the systematic cleavage of chromosomal DNA into multiples of 160-180 bp stretches, resulting in the extracellular presence of mono-(approximately 166 bp) and poly-nucleosomes (332 bp, 498 bp)[124]. Necrosis results in nuclear chromatin clumping and non-specific digestion, producing DNA fragments that are typically larger than 10000 bp. DNA fragments derived from active cellular secretions have been shown to range between 1000 and 3000 bp. Most of the DNA present in plasma occurs as fragments around 180 bases and 360 bases in size and reflects the likely apoptotic origin of the DNA[125].
In cancer patients, cfDNA may originate from multiple sources, including cancer cells, cells from the tumour microenvironment and normal cells such as haematopoietic stem cells, muscle cells and epithelial cells. Once present in circulation, cfDNA levels are influenced by multiple factors including its: (1) Dynamic association and disassociation with extracellular vesicles and several serum proteins, (2) Rate of binding, dissociation and internalisation by cells; and (3) Rate of digestion or clearance, including the activity of deoxyribonuclease I (DNAse), renal excretion into urine, and uptake by the liver and spleen[126]. Furthermore, cfDNA levels can be elevated due to the lysis of white blood cells (WBCs) and release of germline DNA, thus diluting ctDNA concentrations[127]. For this reason, it is crucial that during molecular analysis tumour-specific ctDNA is differentiated from non-tumour cfDNA and contamination of blood samples through WBC lysis is kept to a minimum.
Studies have demonstrated that cfDNA levels in cancer patients are generally higher than those of healthy subjects. cfDNA is present in healthy individuals at average concentrations of 30 ng/mL, ranging from 0-100 ng/mL[118]. In cancer patients, given the additional release of cfDNA from tumour cells, the average concentration of cfDNA is much higher, at approximately 180 ng/mL[128]. cfDNA concentration has also been shown to correlate with tumour size, disease stage and metastatic burden[129-131].
In a study of 640 patients with different cancer types at varying stages, Bettegowda et al[131] showed that cfDNA levels were approximately 100 times higher in stage IV disease compared to stage I disease, providing a rough estimate of tumour size based on cfDNA concentration. In another study, aimed at providing a more sensitive metric for estimating tumour size, researchers reported that mutant alleles increased by 6 mutant copies per ml of plasma for every cubic centimetre of tumour in participants with HGSC[129].
In one of the first studies to examine cfDNA concentration in EOC, Kamat et al[130] demonstrated that cfDNA levels were elevated in advanced EOC compared to normal controls. In a more recent study, the same group evaluated the role of preoperative total plasma cfDNA levels in predicting clinical outcomes in patients with EOC[132]. Again, they reported significantly higher cfDNA levels in the EOC group (median 10113 genomic equivalent/mL, GE/mL) compared with benign ovarian tumours (median, 2365 GE/mL; P < 0.001) and unaffected controls (median, 1912 GE/mL; P < 0.001). Moreover, a statistically significant association of cfDNA > 22000 GE/mL with decreased PFS (P < 0.001) was observed, which was superior to CA125 in predicting mortality. Furthermore, the study reported elevated cfDNA levels in patients with early stage disease which was significantly higher compared with those with benign disease and controls (P < 0.01). Shao et al[133] also reported significantly elevated cfDNA levels in advanced stage OC compared to early stage (P < 0.01). In contrast, No et al[134] reported no significant difference between cfDNA levels in EOC patients compared to controls. In this study, preoperative blood samples of 36 EOC patients and 16 benign tumours were analysed using commercially available copy number assay kits to measure cfDNA levels of four genes; beta-2-microglobulin (B2M), member RAS oncogene family (RAB25), claudin 4 (CLDN4) and ATP-binding cassette subfamily F member 2 (ABCF2). This result may be explained, in part, by the fact that, unlike the previous studies, this study used serum instead of plasma as the cfDNA source.
In a study investigating the presence of tumour-specific TP53 sequences in blood and peritoneal fluid in EOC patients, Swisher et al[135] detected 30% (21/69) of patients with confirmed TP53 mutations (exon 2-11) in plasma or serum samples. cfDNA was detected in 93% (28/30) of cases in peritoneal fluid, including six cases with negative cytology. Following multivariate analysis, they concluded that detection of cfDNA was associated with decreased survival (P = 0.02). Dobrzycka et al[136] investigated the prognostic significance of cfDNA and blood plasma p53 antibodies (p53-Ab) in EOC. Serum p53-Ab is predominantly associated with TP53 gene missense mutations and TP53 accumulation in the tumour. cfDNA and p53-Ab were more frequently detected in patients with HGSC (P < 0.001) compared to other EOC subtypes. Prognosis was significantly worse in cfDNA positive patients compared to cfDNA negative (P = 0.022). Similarly, patients who were p53-Ab positive had a significantly worse prognosis than those who were p53-Ab negative (P < 0.001).
Whole exome sequencing (WES) and targeted gene sequencing has been employed to identify tumour-specific mutations in gynaecological cancers[137]. Subsequent monitoring of these mutations using ctDNA and digital PCR technology, in patients with gynaecological cancers, detected the recurrence of cancer, on average, seven months before disease was visible on cross-sectional imaging. Furthermore, undetectable levels of ctDNA at six months following initial treatment was associated with significantly improved progression free and overall survival.
The correlation between ctDNA levels and tumour stage/size highlights the potential prognostic value of ctDNA. Several studies have demonstrated an association between ctDNA and survival outcome in a range of cancers[138].
Assessing the mutational profile of cancer patients is used as a stratification tool to identify those who may be suitable for targeted therapies. For example, in patients with non-small cell lung cancer (NSCLC), the tyrosine kinase inhibitors (TKI) gefitinib and erlotinib are only beneficial to those with an activating mutation (L858R or exon 19 deletion) in the epidermal growth factor receptor (EGFR) gene[139]. Similarly, patients with malignant melanoma will only benefit from BRAF therapy if they harbour an activating BRAF mutation (V600E)[140]. Identifying these mutations and others enables clinicians to alter therapies accordingly and optimise patient care.
Historically, the assessment of mutational status has been carried out using tissue biopsies, however, this practice has some inherent disadvantages. The analysis of a single tissue biopsy taken from a primary tumour or metastatic site is likely to underestimate the mutational landscape of a tumour and can lead to inaccurate classification. Performing several biopsies on one patient can be impractical and extremely invasive. Furthermore, tissue biopsies are associated with complications, such as infection and pain, and often fail to obtain enough material for high quality mutational profiling. Numerous studies, in various malignancies, have shown that these limitations may be overcome by mutational profiling of cfDNA[106].
In CRC, high concordance between tissue and plasma samples have been reported for the detection of KRAS, NRAS and BRAF mutations, with concordance rates of 91.8% reported in one study for RAS mutations[141]. Another study suggested that cfDNA could replace tumour-tissue analysis resulting in considerable reductions in data turnaround time[142].
There are a limited number of studies investigating the role of cfDNA and treatment resistance in EOC. Murtaza et al[143] carried out WES in serial plasma samples to track the genomic evolution of metastatic cancers in response to treatment. Three patients with advanced EOC were included in the study. Quantification of allele fractions in plasma identified mutant alleles association with emerging treatment resistance. This study established a proof-of-principle that WES of ctDNA could complement current invasive biopsy approaches to identify mutations associated with acquired resistant in advanced cancers.
Overall, these studies demonstrate high concordance between tissue biopsies and plasma samples with higher diagnostic accuracy recorded using cfDNA analysis in most cases.
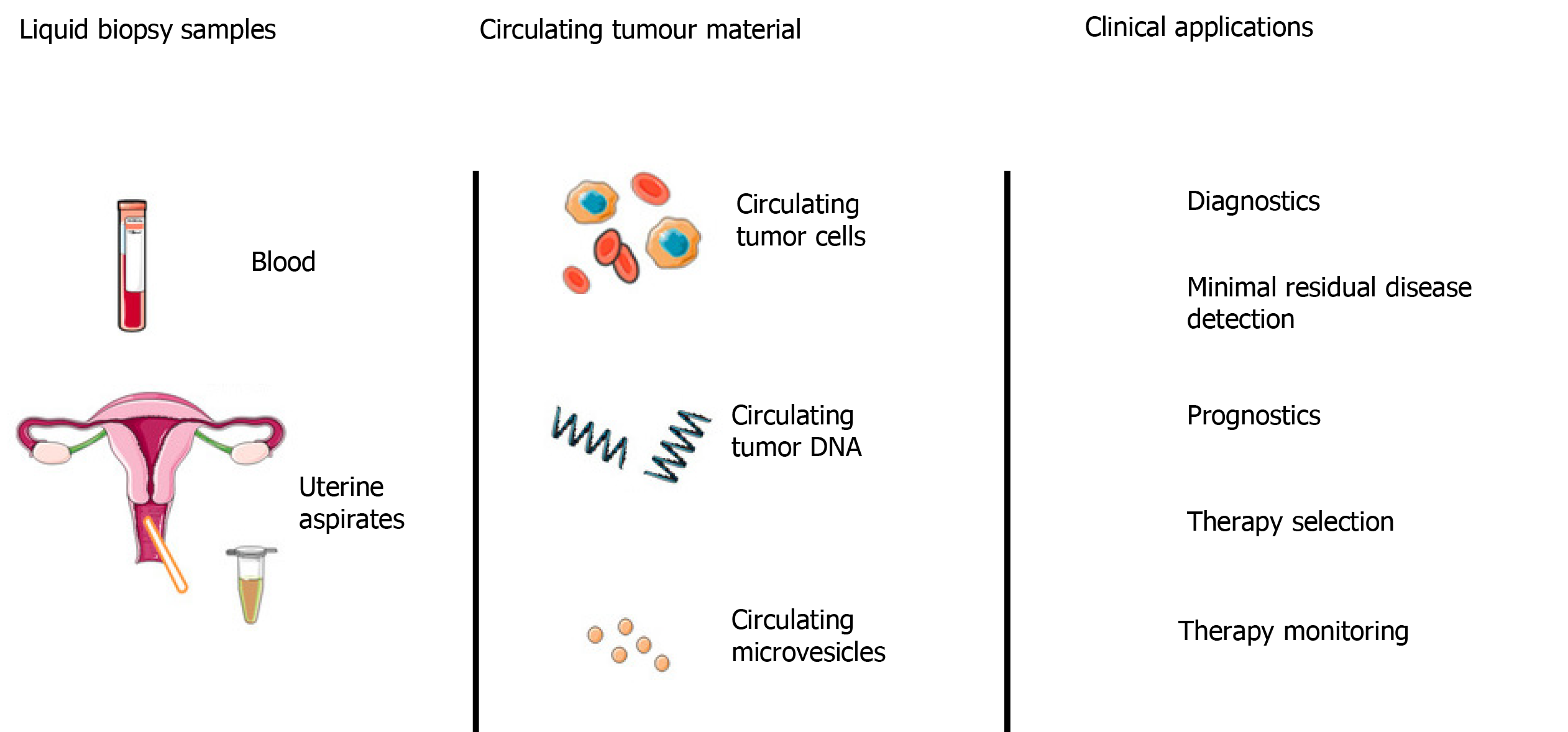
The short half-life of cfDNA, coupled with the minimally invasive nature of venepuncture compared to tissue biopsies makes cfDNA an attractive tool for monitoring treatment response and disease burden (Figure 4). Numerous studies have demonstrated that low levels of cfDNA are associated with a positive treatment response in a range of cancers, including EOC[144]. In contrast, high levels of cfDNA generally correlate with poor response to treatment, treatment resistance, high risk of relapse and poor survival. In the majority of these studies, cfDNA was reported to monitor response to treatment more accurately than traditional methods. In one example, Capizzi et al[145] investigated the role of cfDNA levels in predicting response to chemotherapy in EOC. In this prospective non-randomised clinical trial, 22 patients with advanced EOC (FIGO stage IIIC or IV) undergoing neo-adjuvant chemotherapy were recruited alongside 50 female healthy blood donor controls. Plasma cfDNA levels were quantified before, during and after chemotherapy. Median cfDNA levels were reported to be significantly higher in the cancer group (29.6 ± 22.7 ng/mL) prior to chemotherapy, compared to controls (6.4 ± 4.0 ng/mL), with a sensitivity of 77% and specificity of 96%. A general trend was found between elevated cfDNA levels on completion of chemotherapy and disease progression (P = 0.007), however, the sample size was too small to provide conclusive survival data.
In a retrospective study of 40 patients with relapsed HGSC, Parkinson et al[129] used sequence specific assays to detect predefined TP53 mutations and quantified the TP53 mutant allele frequency (TP53MAF) in cfDNA using digital PCR before, during and after chemotherapy. Pre-treatment ctDNA TP53MAF concentration was positively correlated with total volume of disease, and a decrease of > 60% after one cycle of chemotherapy was associated with longer PFS.
Acquired drug resistance may emerge as a result of de novo mutations or the expansion of a sub-clonal cell population with pre-existing resistance[146]. The underlying mechanisms of acquired resistance are poorly understood. However, longitudinal sampling and analysis of cfDNA can provide valuable insight into the molecular response of cancer during treatment.
cfDNA can be used to monitor the development of resistance by screening for known mutations associated with resistance. Using a digital PCR assay, Ishii et al[147] examined cfDNA in plasma from patients with relapsed NSCLC to identify resistance mutations, namely T790M mutations, associated with EGFR-TKIs. T760M mutation was detected in plasma with a sensitivity of 81.8% and specificity of 85.7%, and overall concordance between plasma and tissue samples was 83.3%. This study showed that digital PCR analysis of plasma is a feasible and accurate alternative to tissue biopsy for detecting T760M mutations in NSCLC patients that become resistant to EGFR-TKIs.
Serial profiling of cfDNA can identify resistant sub-clones before the onset of clinical progression and enable earlier intervention. In a study investigating the acquired resistance to anti-EGFR treatment in CRC, 60% (6/10) patients showed the emergence of secondary KRAS mutations up to four months before an increase in the conventional marker (CEA) was detected, and nine months prior to radiological evidence of relapse[148]. This study also showed that although tumour cells exhibited resistance to EGFR inhibitors, they remained sensitive to a combination of EGFR and MEK inhibitors, enabling early and personalised treatment adjustment.
A key resistance mechanism to platinum-based chemotherapies and PARP inhibitors in BRCA-mutant cancers is the acquisition of BRCA reversion mutations that restore protein function. Lin et al[149] performed targeted next-generation sequencing of cfDNA extracted from plasma collected prior to rucaparib treatment in 112 patients with germline or somatic BRCA-mutant HGSC enrolled in the ARIEL2 study. They found BRCA reversion mutations in cfDNA from 18% (2/11) of platinum-refractory and 13% (5/38) of platinum-resistant patients, compared with 2% (1/48) of platinum-sensitive patients (P = 0.049). Furthermore, patients without BRCA reversion mutations detected in pre-treatment cfDNA had significantly longer rucaparib PFS than those with reversion mutations (median, 9.0 mo vs 18 mo; HR, 0.12; P < 0.0001).
In summary, analysis of cfDNA collected before and after treatment can provide a more comprehensive view of the genetic response of a patients’ tumour, including the dynamic changes in the mutational landscape as well as the heterogeneity that develops due to the selective pressure of therapy.
Another important aspect of cancer management is deciding whether further treatment is required following tumour resection. Decision-making for adjuvant therapy is based on disease stage and certain high-risk clinical or pathological features. As there is, currently, no effective tool of identifying patients with complete tumour resection this practice leads to potential under- and over-treatment with the associated consequences.
cfDNA analysis has shown great promise in the detection of minimal residual disease (MRD) and subsequent risk of recurrence. In one of the first studies to evaluate the use of cfDNA as a biomarker of MRD, Diehl et al[150] showed that tumour derived cfDNA levels decreased by 99% within 24 h after complete surgical resection in CRC, whereas in cases of incomplete resection cfDNA levels did not change significantly and in some cases increased. Furthermore, in subjects with detectable levels of cfDNA after surgery relapse generally occurred within one year. cfDNA levels appeared to be a more reliable and sensitive indicator than the conventional biomarker (carcinoembryonic antigen) in this cohort. More recently, Chaudhuri et al[151] demonstrated the potential of cfDNA deep sequencing analysis in predicting prognosis in patients who had completed potentially curative treatment for early-stage NSCLC with surgery or radical radiotherapy. They retrospectively analysed blood samples from a cohort of 40 patients, with plasma samples taken before treatment and every 2 to 6 mo during follow-up. ctDNA was detectable in 93% (37/40) of patients before any treatment and was detectable in 54% of patients after treatment, all of whom went on to relapse. Detection of ctDNA postoperatively had a very high risk of future relapse (HR, 43.4; 95%CI, 5.7–341), with a median 5.2-mo lead time over clinical progression. Further studies have reported similar findings in a range of malignancies, including EOC[126].
Although these studies are encouraging there remains a large proportion of patients who relapse in whom cfDNA is not detected. Furthermore, the vast majority of these cfDNA assays are patient and mutation specific, limiting their clinical application to wider patient populations. In order to overcome these limitations further research is required to establish highly sensitive cfDNA assays that enable the identification of patients likely to benefit from adjuvant treatments and avoid the adverse effects resulting from unnecessary treatments.
Despite years of research in this area, the diagnosis of early stage cancer remains extremely challenging. Recent research suggests that technological advances in the analysis of cfDNA may provide a solution to these challenges. Studies have shown that cancer-associated mutations can be detected in cfDNA in early-stage disease, before the presence of symptoms and up to 2 years before cancer diagnosis[152-156].
Cervical screening tests have revolutionised the management of cervical cancer by enabling early detection of preinvasive disease. Recently, the traditional Papanicolau smear has been replaced, in many countries, by a liquid-based cytology (LBC) method. Kinde et al[152] exploited this method of DNA collection to develop an assay to detect endometrial and ovarian cancer. Mutational profiling was carried out on 46 cancer patients (24 endometrial cancers and 22 ovarian cancers) for whom LBC specimens were available. The same mutations were detected in the corresponding LBC samples in 100% (24/24) of the endometrial cancers and 41% (9/22) of the ovarian cancers. The same group went on to develop the PapSEEK test using a similar technique to analysis a panel of 18 genes using multiplex PCR[157]. They reported sensitivity of 33% (95%CI, 27%-39%) in the 245 EOC patients tested, including 34% of patients with early stage disease. Specificity was approximately 99% with only 1.4% of 714 women without cancer testing positive. They also analysed plasma in 83 EOC patients for 16 genes of interest and found ctDNA in 43% (95%CI, 33%-55%), with none of the plasma samples from 192 healthy controls testing positive (specificity 100%). When combining LBC samples with matched plasma samples, sensitivity for OC was increased to 63% (95%CI, 51%-73%), including 54% with early stage disease. Although improvements are required before applying this test in routine practice, it highlights the potential to incorporate cfDNA analysis into routine screening tools such as the cervical screening programme.
Non-invasive prenatal testing (NIPT) identifies foetal aneuploidy by sequencing cfDNA in maternal plasma. Pre-symptomatic maternal malignancies have been incidentally detected during NIPT[155]. In a case control study of prospectively collected preoperative HGSC plasma samples, Cohen et al[153] analysed 32 women with HGSC (16 early stage (FIGO I-II) and 16 advanced stage (FIGO III-IV)) and 32 benign controls. Plasma cfDNA was sequenced using a commercial NIPT platform. They detected 40.6% (13/32) HGSC cases using sub-chromosomal analysis, including 38% (6/16) of early stage cases. Although sensitivity was low and correlation with paired tumour DNA was not possible due to a lack of archived specimens, this study established the proof-of-principle that tumour DNA is detectable using NIPT.
The same group developed a blood test to detect eight common cancers through assessment of levels of circulating proteins and mutations in cfDNA[154]. The CancerSEEK® test was used to assess 1005 patients who had been diagnosed with stage I to III cancers of the ovary, liver, stomach, pancreas, oesophagus, colorectum, lung and breast. The median sensitivity of CancerSEEK® among the eight cancer types was 70% (P < 10−96) and ranged from 98% in OC to 33% in breast cancers. At this sensitivity, the specificity was > 99%; with 7 of the 812 controls recorded as positive. Although sensitivity was reported as > 70% for stage II (73%) and stage III (78%) disease, only 43% of stage I cancers were detected. A key concern with this test is its achievable PPV. The prevalence of the eight cancers in healthy individuals > 64 years of age is approximately 1%. Assuming the CancerSEEK® test could achieve 99% sensitivity and specificity, the resulting PPV would be only 50%, which equates to 50% of positive tests being false positives. However, these results imply that combination strategies have the power to greatly improve liquid biopsy analyses.
Gormally et al[156] assessed the significance of plasma DNA mutations for subsequent cancer development in healthy subjects in a large longitudinal prospective study. The study included > 520000 healthy volunteers recruited from 10 European countries. Plasma specimens were tested for TP53 and KRAS2 mutations. Results showed that mutations in TP53 and KRAS2 could be detected in cfDNA of healthy subjects on average 20.8 mo (range, 1.8-44.8) and 14.3 mo (range, 2.6-24.9) before cancer diagnosis, respectively. TP53 and KRAS2 mutations were detected in 3% and 1%, respectively, of subjects who did not develop cancer during follow up. This is an important finding as it highlights the presence of high levels of oncogenic drivers in the plasma of healthy individuals. This has been attributed to a common aging-related phenomenon known as clonal haematopoiesis, in which haematopoietic stem cells or other early blood cell progenitors contribute to the formation of genetically distinct subpopulations of bloods cells[158]. The establishment of a clonal population may occur when a stem or progenitor cell acquires one or more somatic mutations that give it a competitive advantage over other haematopoietic cells. The incidence of clonal haematopoiesis rises with age and is an important consideration when evaluating potential blood-based tumour-specific biomarkers.
Whilst these studies demonstrate the potential of cfDNA as an early detection marker, several significant obstacles need to be overcome before the majority could be used in a clinical setting. The main obstacles to the development of cfDNA based biomarkers are: (1) Low abundance of ctDNA in the blood; and (2) High levels of background non-cancerous cfDNA, mostly shed from WBCs. Highly sensitive technologies are required to accurately detect scarcely abundant alleles within high background levels of nontarget molecules.
Due to this, cfDNA-based cancer biomarkers have yet to make it into the clinical arena. The greatest progress has been observed in obstetric medicine, where cfDNA has been successfully used for fetal Rhesus D genotyping, for the detection of paternally inherited genetic disorders from maternal blood, and revolutionised the identification of fetal aneuploidy through non-invasive pre-natal screening[159]. In the future this level of success may be seen for cancer-specific cfDNA biomarkers.
The use of liquid biopsies is a fast-emerging area of cancer diagnostics, in particular the detection of ctDNA, and multiple cancer types are known to produce quantifiable levels of ctDNA. Studies have shown ctDNA to be useful in monitoring for minimal residual disease, treatment response, chemoresistance, and tumour heterogeneity. ctDNA can be used as an early warning diagnostic tool, either through identification of known genetic aberrations (such as TP53 mutations), or through measurement of cancer-specific DNA methylation (DNAme) of particular genomic loci. However, use of this technology for early diagnosis is very much in its infancy. In EOC it has been shown that TP53 mutations can be identified in ctDNA, using tagged amplicon deep sequencing, in up to 65% of patients with advanced EOC[160]. Once optimised this could be a very useful liquid biopsy for HGSC but it would be important to ascertain the TP53 mutation rate in normal healthy controls as a baseline.
At present, sequencing technology allows for the detection of one mutant allele, e.g., p53, in a background of 1000 wild-type molecules[161]. For this reason, the current focus of our research is on DNAme because it allows for the detection of specific patterns rather than single point mutations, potentially improving the performance characteristics and detection limit of such an assay. There have been numerous reports of alterations in methylation events occurring in EOC, including HGSC[162-164]. This could prove to be an extremely useful mechanism through which HGSCs might be detected earlier and more consistently.
The identification of a biomarker requires a number of phases of development which can be crudely described as; case and control selection, determination of detection limits and assay precision, validation in second/external datasets, statistical interpretation and ROC analysis. The failure to find suitable biomarkers for EOC, despite significant investment in the United Kingdom, Europe and the United States, led to an inquiry into possible causes. This inquiry recommended the use of the PRoBE (prospective-specimen collection, with retrospective-blinded evaluation) design for biomarker discovery and validation because it was felt that biomarkers discovered in clinical sample sets collected at diagnosis from symptomatic patients and controls in hospital settings are unlikely to fully represent the screening population[165]. The PRoBE design involves blinded case-control studies nested within a prospective cohort representing the target population. The specimens and matched clinical data will have been collected prior to knowledge of the outcome (e.g., diagnosis of EOC). Paramount to the successful implementation of the PRoBE design is the knowledge of the accurate calibrator-control for the disease the biomarker is aimed for.
As discussed, the clinicopathological and molecular developments over the last decade has redefined EOC as essentially representing five distinct disease entities. It is now the time to consider identifying disease-specific biomarkers rather than a generic EOC tumour marker. This will likely yield more success and, ultimately, result in increased precision of the biomarker.
HGSC is the major contributor to morbidity and mortality under the EOC “umbrella”. Therefore, an accurate disease-specific biomarker is urgently needed; not just for “screening” but for greater diagnostic accuracy and also for monitoring disease stability/progression during treatment and assessing residual disease post cytoreductive surgery. Our use of cutting-edge, high throughput, molecular assay technologies has helped clarify the underlying molecular profile of this disease[86]. Knowledge like this will guide the identification and validation of novel transcripts that carry the potential to act as disease-specific biomarkers of HGSC.
Manuscript source: Invited manuscript
Specialty type: Oncology
Country/Territory of origin: United Kingdom
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): 0
P-Reviewer: Ju SQ, Lopez-Guerrero J S-Editor: Huang P L-Editor: A P-Editor: Wang LL
| 1. | Cancer Research UK. Ovarian Cancer. Available from: https://www.cancerresearchuk.org/about-cancer/ovarian-cancer. [Cited in This Article: ] |
| 2. | Kurman RJ, Carcangiu ML, Herrington CS. World Health Organisation Classification of Tumours of the Female Reproductive Organs. 4th ed. International Agency for Research on Cancer, 2014. [Cited in This Article: ] |
| 3. | Kurman RJ, Shih IeM. The origin and pathogenesis of epithelial ovarian cancer: a proposed unifying theory. Am J Surg Pathol. 2010;34:433-443. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1183] [Cited by in F6Publishing: 1190] [Article Influence: 85.0] [Reference Citation Analysis (0)] |
| 4. | Kurman RJ, Shih IeM. Molecular pathogenesis and extraovarian origin of epithelial ovarian cancer--shifting the paradigm. Hum Pathol. 2011;42:918-931. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 738] [Cited by in F6Publishing: 767] [Article Influence: 59.0] [Reference Citation Analysis (0)] |
| 5. | Gadducci A, Guerrieri ME, Genazzani AR. New insights on the pathogenesis of ovarian carcinoma: molecular basis and clinical implications. Gynecol Endocrinol. 2012;28:582-586. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 11] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 6. | McCluggage WG. Morphological subtypes of ovarian carcinoma: a review with emphasis on new developments and pathogenesis. Pathology. 2011;43:420-432. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 308] [Cited by in F6Publishing: 326] [Article Influence: 25.1] [Reference Citation Analysis (0)] |
| 7. | Modesitt SC, Tortolero-Luna G, Robinson JB, Gershenson DM, Wolf JK. Ovarian and extraovarian endometriosis-associated cancer. Obstet Gynecol. 2002;100:788-795. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 68] [Cited by in F6Publishing: 110] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 8. | Uekuri C, Shigetomi H, Ono S, Sasaki Y, Matsuura M, Kobayashi H. Toward an understanding of the pathophysiology of clear cell carcinoma of the ovary (Review). Oncol Lett. 2013;6:1163-1173. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 19] [Cited by in F6Publishing: 19] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 9. | Senthong A, Kitkumthorn N, Rattanatanyong P, Khemapech N, Triratanachart S, Mutirangura A. Differences in LINE-1 methylation between endometriotic ovarian cyst and endometriosis-associated ovarian cancer. Int J Gynecol Cancer. 2014;24:36-42. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 25] [Cited by in F6Publishing: 22] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 10. | Samartzis EP, Noske A, Dedes KJ, Fink D, Imesch P. ARID1A mutations and PI3K/AKT pathway alterations in endometriosis and endometriosis-associated ovarian carcinomas. Int J Mol Sci. 2013;14:18824-18849. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 97] [Cited by in F6Publishing: 112] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 11. | Wiegand KC, Shah SP, Al-Agha OM, Zhao Y, Tse K, Zeng T, Senz J, McConechy MK, Anglesio MS, Kalloger SE, Yang W, Heravi-Moussavi A, Giuliany R, Chow C, Fee J, Zayed A, Prentice L, Melnyk N, Turashvili G, Delaney AD, Madore J, Yip S, McPherson AW, Ha G, Bell L, Fereday S, Tam A, Galletta L, Tonin PN, Provencher D, Miller D, Jones SJ, Moore RA, Morin GB, Oloumi A, Boyd N, Aparicio SA, Shih IeM, Mes-Masson AM, Bowtell DD, Hirst M, Gilks B, Marra MA, Huntsman DG. ARID1A mutations in endometriosis-associated ovarian carcinomas. N Engl J Med. 2010;363:1532-1543. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1228] [Cited by in F6Publishing: 1199] [Article Influence: 85.6] [Reference Citation Analysis (0)] |
| 12. | Mao TL, Shih IeM. The roles of ARID1A in gynecologic cancer. J Gynecol Oncol. 2013;24:376-381. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 42] [Cited by in F6Publishing: 45] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 13. | Ahmed AA, Etemadmoghadam D, Temple J, Lynch AG, Riad M, Sharma R, Stewart C, Fereday S, Caldas C, Defazio A, Bowtell D, Brenton JD. Driver mutations in TP53 are ubiquitous in high grade serous carcinoma of the ovary. J Pathol. 2010;221:49-56. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 499] [Cited by in F6Publishing: 551] [Article Influence: 39.4] [Reference Citation Analysis (0)] |
| 14. | Crum CP, McKeon FD, Xian W. The oviduct and ovarian cancer: causality, clinical implications, and "targeted prevention". Clin Obstet Gynecol. 2012;55:24-35. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 54] [Cited by in F6Publishing: 56] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 15. | Vang R, Shih IeM, Kurman RJ. Ovarian low-grade and high-grade serous carcinoma: pathogenesis, clinicopathologic and molecular biologic features, and diagnostic problems. Adv Anat Pathol. 2009;16:267-282. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 396] [Cited by in F6Publishing: 396] [Article Influence: 26.4] [Reference Citation Analysis (0)] |
| 16. | Merritt MA, Cramer DW. Molecular pathogenesis of endometrial and ovarian cancer. Cancer Biomark. 2010;9:287-305. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 46] [Cited by in F6Publishing: 57] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 17. | Etemadmoghadam D, deFazio A, Beroukhim R, Mermel C, George J, Getz G, Tothill R, Okamoto A, Raeder MB, Harnett P, Lade S, Akslen LA, Tinker AV, Locandro B, Alsop K, Chiew YE, Traficante N, Fereday S, Johnson D, Fox S, Sellers W, Urashima M, Salvesen HB, Meyerson M, Bowtell D; AOCS Study Group. Integrated genome-wide DNA copy number and expression analysis identifies distinct mechanisms of primary chemoresistance in ovarian carcinomas. Clin Cancer Res. 2009;15:1417-1427. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 204] [Cited by in F6Publishing: 233] [Article Influence: 15.5] [Reference Citation Analysis (0)] |
| 18. | Cho KR, Shih IeM. Ovarian cancer. Annu Rev Pathol. 2009;4:287-313. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 482] [Cited by in F6Publishing: 520] [Article Influence: 34.7] [Reference Citation Analysis (0)] |
| 19. | Beirne JP, Irwin GW, McIntosh SA, Harley IJG, Harkin DP. The molecular and genetic basis of inherited cancer risk in gynaecology. TOG. 2015;17:233-241. [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 7] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 20. | Weberpals JI, Clark-Knowles KV, Vanderhyden BC. Sporadic epithelial ovarian cancer: clinical relevance of BRCA1 inhibition in the DNA damage and repair pathway. J Clin Oncol. 2008;26:3259-3267. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 50] [Cited by in F6Publishing: 56] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 21. | Tothill RW, Tinker AV, George J, Brown R, Fox SB, Lade S, Johnson DS, Trivett MK, Etemadmoghadam D, Locandro B, Traficante N, Fereday S, Hung JA, Chiew YE, Haviv I; Australian Ovarian Cancer Study Group; Gertig D; DeFazio A; Bowtell DD. Novel molecular subtypes of serous and endometrioid ovarian cancer linked to clinical outcome. Clin Cancer Res. 2008;14:5198-5208. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1023] [Cited by in F6Publishing: 1075] [Article Influence: 67.2] [Reference Citation Analysis (0)] |
| 22. | Bast RC Jr, Feeney M, Lazarus H, Nadler LM, Colvin RB, Knapp RC. Reactivity of a monoclonal antibody with human ovarian carcinoma. J Clin Invest. 1981;68:1331-1337. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1132] [Cited by in F6Publishing: 1095] [Article Influence: 25.5] [Reference Citation Analysis (0)] |
| 23. | Jacobs I, Bast RC Jr. The CA 125 tumour-associated antigen: a review of the literature. Hum Reprod. 1989;4:1-12. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 517] [Cited by in F6Publishing: 506] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 24. | National Institute for Health and Care Excellence. Ovarian cancer: recognition and initial management. Clinical guideline [CG122], 2011. Available from: https://www.nice.org.uk/Guidance/CG122. [Cited in This Article: ] |
| 25. | Sundar SaR, K. Benign and malignant ovarian masses. In: Luesley DMaB, P.N., editor. Obstetrics and Gynaecology: An evidence-based text for MRCOG. 2nd ed. Great Britain: Hodder Arnold, 2010; 814-829. [DOI] [Cited in This Article: ] |
| 26. | Bottoni P, Scatena R. The Role of CA 125 as Tumor Marker: Biochemical and Clinical Aspects. Adv Exp Med Biol. 2015;867:229-244. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 83] [Cited by in F6Publishing: 108] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 27. | Díaz-Padilla I, Razak AR, Minig L, Bernardini MQ, María Del Campo J. Prognostic and predictive value of CA-125 in the primary treatment of epithelial ovarian cancer: potentials and pitfalls. Clin Transl Oncol. 2012;14:15-20. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 17] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 28. | Kristjansdottir B, Levan K, Partheen K, Sundfeldt K. Diagnostic performance of the biomarkers HE4 and CA125 in type I and type II epithelial ovarian cancer. Gynecol Oncol. 2013;131:52-58. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 49] [Cited by in F6Publishing: 49] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 29. | Zhao T, Hu W. CA125 and HE4: Measurement Tools for Ovarian Cancer. Gynecol Obstet Invest. 2016;81:430-435. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 21] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 30. | Ghaemmaghami F, Karimi Zarchi M, Hamedi B. High levels of CA125 (over 1,000 IU/ml) in patients with gynecologic disease and no malignant conditions: three cases and literature review. Arch Gynecol Obstet. 2007;276:559-561. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 27] [Cited by in F6Publishing: 28] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 31. | Abu J, Brown L, Ireland D, Sizeland E. Mesovarian hemangioma presenting as massive ascites, pelvic mass, and elevated CA125. Int J Gynecol Cancer. 2006;16 Suppl 1:412-414. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2] [Cited by in F6Publishing: 2] [Article Influence: 0.1] [Reference Citation Analysis (0)] |
| 32. | Kaneta Y, Nishino R, Asaoka K, Toyoshima K, Ito K, Kitai H. Ovarian hemangioma presenting as pseudo-Meigs' syndrome with elevated CA125. J Obstet Gynaecol Res. 2003;29:132-135. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 17] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 33. | Nossov V, Amneus M, Su F, Lang J, Janco JM, Reddy ST, Farias-Eisner R. The early detection of ovarian cancer: from traditional methods to proteomics. Can we really do better than serum CA-125? Am J Obstet Gynecol. 2008;199:215-223. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 189] [Cited by in F6Publishing: 187] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 34. | Bast RC Jr, Brewer M, Zou C, Hernandez MA, Daley M, Ozols R, Lu K, Lu Z, Badgwell D, Mills GB, Skates S, Zhang Z, Chan D, Lokshin A, Yu Y. Prevention and early detection of ovarian cancer: mission impossible? Recent Results Cancer Res. 2007;174:91-100. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 73] [Cited by in F6Publishing: 78] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 35. | Skates SJ. Ovarian cancer screening: development of the risk of ovarian cancer algorithm (ROCA) and ROCA screening trials. Int J Gynecol Cancer. 2012;22 Suppl 1:S24-S26. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 49] [Cited by in F6Publishing: 56] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 36. | Lu KH, Skates S, Hernandez MA, Bedi D, Bevers T, Leeds L, Moore R, Granai C, Harris S, Newland W, Adeyinka O, Geffen J, Deavers MT, Sun CC, Horick N, Fritsche H, Bast RC Jr. A 2-stage ovarian cancer screening strategy using the Risk of Ovarian Cancer Algorithm (ROCA) identifies early-stage incident cancers and demonstrates high positive predictive value. Cancer. 2013;119:3454-3461. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 90] [Cited by in F6Publishing: 91] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 37. | Pinsky PF, Zhu C, Skates SJ, Black A, Partridge E, Buys SS, Berg CD. Potential effect of the risk of ovarian cancer algorithm (ROCA) on the mortality outcome of the Prostate, Lung, Colorectal and Ovarian (PLCO) trial. Int J Cancer. 2013;132:2127-2133. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 28] [Cited by in F6Publishing: 29] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 38. | Jacobs IJ, Menon U, Ryan A, Gentry-Maharaj A, Burnell M, Kalsi JK, Amso NN, Apostolidou S, Benjamin E, Cruickshank D, Crump DN, Davies SK, Dawnay A, Dobbs S, Fletcher G, Ford J, Godfrey K, Gunu R, Habib M, Hallett R, Herod J, Jenkins H, Karpinskyj C, Leeson S, Lewis SJ, Liston WR, Lopes A, Mould T, Murdoch J, Oram D, Rabideau DJ, Reynolds K, Scott I, Seif MW, Sharma A, Singh N, Taylor J, Warburton F, Widschwendter M, Williamson K, Woolas R, Fallowfield L, McGuire AJ, Campbell S, Parmar M, Skates SJ. Ovarian cancer screening and mortality in the UK Collaborative Trial of Ovarian Cancer Screening (UKCTOCS): a randomised controlled trial. Lancet. 2016;387:945-956. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 698] [Cited by in F6Publishing: 646] [Article Influence: 80.8] [Reference Citation Analysis (0)] |
| 39. | Hellström I, Raycraft J, Hayden-Ledbetter M, Ledbetter JA, Schummer M, McIntosh M, Drescher C, Urban N, Hellström KE. The HE4 (WFDC2) protein is a biomarker for ovarian carcinoma. Cancer Res. 2003;63:3695-3700. [PubMed] [Cited in This Article: ] |
| 40. | Wu L, Dai ZY, Qian YH, Shi Y, Liu FJ, Yang C. Diagnostic value of serum human epididymis protein 4 (HE4) in ovarian carcinoma: a systematic review and meta-analysis. Int J Gynecol Cancer. 2012;22:1106-1112. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 53] [Cited by in F6Publishing: 61] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 41. | Simmons AR, Clarke CH, Badgwell DB, Lu Z, Sokoll LJ, Lu KH, Zhang Z, Bast RC Jr, Skates SJ. Validation of a Biomarker Panel and Longitudinal Biomarker Performance for Early Detection of Ovarian Cancer. Int J Gynecol Cancer. 2016;26:1070-1077. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 32] [Cited by in F6Publishing: 32] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 42. | Romagnolo C, Leon AE, Fabricio ASC, Taborelli M, Polesel J, Del Pup L, Steffan A, Cervo S, Ravaggi A, Zanotti L, Bandiera E, Odicino FE, Scattolo N, Squarcina E, Papadakis C, Maggino T, Gion M. HE4, CA125 and risk of ovarian malignancy algorithm (ROMA) as diagnostic tools for ovarian cancer in patients with a pelvic mass: An Italian multicenter study. Gynecol Oncol. 2016;141:303-311. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 69] [Cited by in F6Publishing: 76] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 43. | Li F, Tie R, Chang K, Wang F, Deng S, Lu W, Yu L, Chen M. Does risk for ovarian malignancy algorithm excel human epididymis protein 4 and CA125 in predicting epithelial ovarian cancer: a meta-analysis. BMC Cancer. 2012;12:258. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 75] [Cited by in F6Publishing: 89] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 44. | Cramer DW, Bast RC Jr, Berg CD, Diamandis EP, Godwin AK, Hartge P, Lokshin AE, Lu KH, McIntosh MW, Mor G, Patriotis C, Pinsky PF, Thornquist MD, Scholler N, Skates SJ, Sluss PM, Srivastava S, Ward DC, Zhang Z, Zhu CS, Urban N. Ovarian cancer biomarker performance in prostate, lung, colorectal, and ovarian cancer screening trial specimens. Cancer Prev Res (Phila). 2011;4:365-374. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 220] [Cited by in F6Publishing: 230] [Article Influence: 17.7] [Reference Citation Analysis (0)] |
| 45. | Moore LE, Pfeiffer RM, Zhang Z, Lu KH, Fung ET, Bast RC Jr. Proteomic biomarkers in combination with CA 125 for detection of epithelial ovarian cancer using prediagnostic serum samples from the Prostate, Lung, Colorectal, and Ovarian (PLCO) Cancer Screening Trial. Cancer. 2012;118:91-100. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 64] [Cited by in F6Publishing: 69] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 46. | Moore RG, McMeekin DS, Brown AK, DiSilvestro P, Miller MC, Allard WJ, Gajewski W, Kurman R, Bast RC Jr, Skates SJ. A novel multiple marker bioassay utilizing HE4 and CA125 for the prediction of ovarian cancer in patients with a pelvic mass. Gynecol Oncol. 2009;112:40-46. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 538] [Cited by in F6Publishing: 566] [Article Influence: 35.4] [Reference Citation Analysis (0)] |
| 47. | Chudecka-Głaz A, Cymbaluk-Płoska A, Jastrzębska J, Menkiszak J. Can ROMA algorithm stratify ovarian tumor patients better when being based on specific age ranges instead of the premenopausal and postmenopausal status? Tumour Biol. 2016;37:8879-8887. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 8] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 48. | Dikmen ZG, Colak A, Dogan P, Tuncer S, Akbiyik F. Diagnostic performances of CA125, HE4, and ROMA index in ovarian cancer. Eur J Gynaecol Oncol. 2015;36:457-462. [PubMed] [Cited in This Article: ] |
| 49. | Fujiwara H, Suzuki M, Takeshima N, Takizawa K, Kimura E, Nakanishi T, Yamada K, Takano H, Sasaki H, Koyama K, Ochiai K. Evaluation of human epididymis protein 4 (HE4) and Risk of Ovarian Malignancy Algorithm (ROMA) as diagnostic tools of type I and type II epithelial ovarian cancer in Japanese women. Tumour Biol. 2015;36:1045-1053. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 32] [Cited by in F6Publishing: 33] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 50. | Jacob F, Meier M, Caduff R, Goldstein D, Pochechueva T, Hacker N, Fink D, Heinzelmann-Schwarz V. No benefit from combining HE4 and CA125 as ovarian tumor markers in a clinical setting. Gynecol Oncol. 2011;121:487-491. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 112] [Cited by in F6Publishing: 128] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 51. | Kadija S, Stefanovic A, Jeremic K, Radojevic MM, Nikolic L, Markovic I, Atanackovic J. The utility of human epididymal protein 4, cancer antigen 125, and risk for malignancy algorithm in ovarian cancer and endometriosis. Int J Gynecol Cancer. 2012;22:238-244. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 40] [Cited by in F6Publishing: 43] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 52. | Montagnana M, Danese E, Ruzzenente O, Bresciani V, Nuzzo T, Gelati M, Salvagno GL, Franchi M, Lippi G, Guidi GC. The ROMA (Risk of Ovarian Malignancy Algorithm) for estimating the risk of epithelial ovarian cancer in women presenting with pelvic mass: is it really useful? Clin Chem Lab Med. 2011;49:521-525. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 107] [Cited by in F6Publishing: 122] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 53. | Van Gorp T, Cadron I, Despierre E, Daemen A, Leunen K, Amant F, Timmerman D, De Moor B, Vergote I. HE4 and CA125 as a diagnostic test in ovarian cancer: prospective validation of the Risk of Ovarian Malignancy Algorithm. Br J Cancer. 2011;104:863-870. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 223] [Cited by in F6Publishing: 241] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 54. | Castle PE. Gynecological cancer: More evidence supporting human papillomavirus testing. Nat Rev Clin Oncol. 2012;9:131-132. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 7] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 55. | Summaries for patients. Screening for ovarian cancer: U.S. Preventive Services Task Force reaffirmation recommendation statement. Ann Intern Med. 2012;157:I-56. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2] [Cited by in F6Publishing: 7] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 56. | Bast RC Jr. Status of tumor markers in ovarian cancer screening. J Clin Oncol. 2003;21:200s-205s. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 141] [Cited by in F6Publishing: 155] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 57. | Sayasneh A, Kaijser J, Preisler J, Smith AA, Raslan F, Johnson S, Husicka R, Ferrara L, Stalder C, Ghaem-Maghami S, Timmerman D, Bourne T. Accuracy of ultrasonography performed by examiners with varied training and experience in predicting specific pathology of adnexal masses. Ultrasound Obstet Gynecol. 2015;45:605-612. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 21] [Cited by in F6Publishing: 22] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 58. | Buys SS, Partridge E, Black A, Johnson CC, Lamerato L, Isaacs C, Reding DJ, Greenlee RT, Yokochi LA, Kessel B, Crawford ED, Church TR, Andriole GL, Weissfeld JL, Fouad MN, Chia D, O'Brien B, Ragard LR, Clapp JD, Rathmell JM, Riley TL, Hartge P, Pinsky PF, Zhu CS, Izmirlian G, Kramer BS, Miller AB, Xu JL, Prorok PC, Gohagan JK, Berg CD; PLCO Project Team. Effect of screening on ovarian cancer mortality: the Prostate, Lung, Colorectal and Ovarian (PLCO) Cancer Screening Randomized Controlled Trial. JAMA. 2011;305:2295-2303. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 870] [Cited by in F6Publishing: 840] [Article Influence: 64.6] [Reference Citation Analysis (0)] |
| 59. | Pinsky PF, Yu K, Kramer BS, Black A, Buys SS, Partridge E, Gohagan J, Berg CD, Prorok PC. Extended mortality results for ovarian cancer screening in the PLCO trial with median 15years follow-up. Gynecol Oncol. 2016;143:270-275. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 94] [Cited by in F6Publishing: 84] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 60. | Shih IeM, Kurman RJ. Ovarian tumorigenesis: a proposed model based on morphological and molecular genetic analysis. Am J Pathol. 2004;164:1511-1518. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 873] [Cited by in F6Publishing: 881] [Article Influence: 44.1] [Reference Citation Analysis (0)] |
| 61. | Fathalla MF. Incessant ovulation--a factor in ovarian neoplasia? Lancet. 1971;2:163. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 769] [Cited by in F6Publishing: 721] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 62. | Casagrande JT, Louie EW, Pike MC, Roy S, Ross RK, Henderson BE. "Incessant ovulation" and ovarian cancer. Lancet. 1979;2:170-173. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 324] [Cited by in F6Publishing: 349] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 63. | Whittemore AS, Harris R, Itnyre J. Characteristics relating to ovarian cancer risk: collaborative analysis of 12 US case-control studies. IV. The pathogenesis of epithelial ovarian cancer. Collaborative Ovarian Cancer Group. Am J Epidemiol. 1992;136:1212-1220. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 186] [Cited by in F6Publishing: 196] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 64. | Rosenblatt KA, Thomas DB. Lactation and the risk of epithelial ovarian cancer. The WHO Collaborative Study of Neoplasia and Steroid Contraceptives. Int J Epidemiol. 1993;22:192-197. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 126] [Cited by in F6Publishing: 133] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 65. | Whittemore AS. Personal characteristics relating to risk of invasive epithelial ovarian cancer in older women in the United States. Cancer. 1993;71:558-565. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 14] [Cited by in F6Publishing: 15] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 66. | Adami HO, Hsieh CC, Lambe M, Trichopoulos D, Leon D, Persson I, Ekbom A, Janson PO. Parity, age at first childbirth, and risk of ovarian cancer. Lancet. 1994;344:1250-1254. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 230] [Cited by in F6Publishing: 231] [Article Influence: 7.7] [Reference Citation Analysis (0)] |
| 67. | Risch HA, Marrett LD, Howe GR. Parity, contraception, infertility, and the risk of epithelial ovarian cancer. Am J Epidemiol. 1994;140:585-597. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 158] [Cited by in F6Publishing: 172] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 68. | Singh N, McCluggage WG, Gilks CB. High-grade serous carcinoma of tubo-ovarian origin: recent developments. Histopathology. 2017;71:339-356. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 51] [Cited by in F6Publishing: 55] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 69. | Salvador S, Gilks B, Köbel M, Huntsman D, Rosen B, Miller D. The fallopian tube: primary site of most pelvic high-grade serous carcinomas. Int J Gynecol Cancer. 2009;19:58-64. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 147] [Cited by in F6Publishing: 154] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 70. | Bannatyne P, Russell P. Early adenocarcinoma of the fallopian tubes. A case for multifocal tumorigenesis. Diagn Gynecol Obstet. 1981;3:49-60. [PubMed] [Cited in This Article: ] |
| 71. | Colgan TJ, Murphy J, Cole DE, Narod S, Rosen B. Occult carcinoma in prophylactic oophorectomy specimens: prevalence and association with BRCA germline mutation status. Am J Surg Pathol. 2001;25:1283-1289. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 279] [Cited by in F6Publishing: 286] [Article Influence: 12.4] [Reference Citation Analysis (0)] |
| 72. | Piek JM, van Diest PJ, Zweemer RP, Jansen JW, Poort-Keesom RJ, Menko FH, Gille JJ, Jongsma AP, Pals G, Kenemans P, Verheijen RH. Dysplastic changes in prophylactically removed Fallopian tubes of women predisposed to developing ovarian cancer. J Pathol. 2001;195:451-456. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 548] [Cited by in F6Publishing: 525] [Article Influence: 22.8] [Reference Citation Analysis (0)] |
| 73. | Aziz S, Kuperstein G, Rosen B, Cole D, Nedelcu R, McLaughlin J, Narod SA. A genetic epidemiological study of carcinoma of the fallopian tube. Gynecol Oncol. 2001;80:341-345. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 162] [Cited by in F6Publishing: 171] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 74. | Zweemer RP, van Diest PJ, Verheijen RH, Ryan A, Gille JJ, Sijmons RH, Jacobs IJ, Menko FH, Kenemans P. Molecular evidence linking primary cancer of the fallopian tube to BRCA1 germline mutations. Gynecol Oncol. 2000;76:45-50. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 143] [Cited by in F6Publishing: 151] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 75. | Brose MS, Rebbeck TR, Calzone KA, Stopfer JE, Nathanson KL, Weber BL. Cancer risk estimates for BRCA1 mutation carriers identified in a risk evaluation program. J Natl Cancer Inst. 2002;94:1365-1372. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 503] [Cited by in F6Publishing: 469] [Article Influence: 21.3] [Reference Citation Analysis (0)] |
| 76. | Finch A, Shaw P, Rosen B, Murphy J, Narod SA, Colgan TJ. Clinical and pathologic findings of prophylactic salpingo-oophorectomies in 159 BRCA1 and BRCA2 carriers. Gynecol Oncol. 2006;100:58-64. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 290] [Cited by in F6Publishing: 302] [Article Influence: 15.9] [Reference Citation Analysis (0)] |
| 77. | Paley PJ, Swisher EM, Garcia RL, Agoff SN, Greer BE, Peters KL, Goff BA. Occult cancer of the fallopian tube in BRCA-1 germline mutation carriers at prophylactic oophorectomy: a case for recommending hysterectomy at surgical prophylaxis. Gynecol Oncol. 2001;80:176-180. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 155] [Cited by in F6Publishing: 161] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 78. | Levine DA, Argenta PA, Yee CJ, Marshall DS, Olvera N, Bogomolniy F, Rahaman JA, Robson ME, Offit K, Barakat RR, Soslow RA, Boyd J. Fallopian tube and primary peritoneal carcinomas associated with BRCA mutations. J Clin Oncol. 2003;21:4222-4227. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 158] [Cited by in F6Publishing: 170] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 79. | Powell CB, Kenley E, Chen LM, Crawford B, McLennan J, Zaloudek C, Komaromy M, Beattie M, Ziegler J. Risk-reducing salpingo-oophorectomy in BRCA mutation carriers: role of serial sectioning in the detection of occult malignancy. J Clin Oncol. 2005;23:127-132. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 294] [Cited by in F6Publishing: 308] [Article Influence: 16.2] [Reference Citation Analysis (0)] |
| 80. | Medeiros F, Muto MG, Lee Y, Elvin JA, Callahan MJ, Feltmate C, Garber JE, Cramer DW, Crum CP. The tubal fimbria is a preferred site for early adenocarcinoma in women with familial ovarian cancer syndrome. Am J Surg Pathol. 2006;30:230-236. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 631] [Cited by in F6Publishing: 591] [Article Influence: 32.8] [Reference Citation Analysis (0)] |
| 81. | Leeper K, Garcia R, Swisher E, Goff B, Greer B, Paley P. Pathologic findings in prophylactic oophorectomy specimens in high-risk women. Gynecol Oncol. 2002;87:52-56. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 236] [Cited by in F6Publishing: 244] [Article Influence: 11.1] [Reference Citation Analysis (0)] |
| 82. | Carcangiu ML, Peissel B, Pasini B, Spatti G, Radice P, Manoukian S. Incidental carcinomas in prophylactic specimens in BRCA1 and BRCA2 germ-line mutation carriers, with emphasis on fallopian tube lesions: report of 6 cases and review of the literature. Am J Surg Pathol. 2006;30:1222-1230. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 105] [Cited by in F6Publishing: 111] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 83. | Olivier RI, van Beurden M, Lubsen MA, Rookus MA, Mooij TM, van de Vijver MJ, van't Veer LJ. Clinical outcome of prophylactic oophorectomy in BRCA1/BRCA2 mutation carriers and events during follow-up. Br J Cancer. 2004;90:1492-1497. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 103] [Cited by in F6Publishing: 109] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 84. | Gilks CB, Irving J, Köbel M, Lee C, Singh N, Wilkinson N, McCluggage WG. Incidental nonuterine high-grade serous carcinomas arise in the fallopian tube in most cases: further evidence for the tubal origin of high-grade serous carcinomas. Am J Surg Pathol. 2015;39:357-364. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 96] [Cited by in F6Publishing: 99] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 85. | Morrison JC, Blanco LZ Jr, Vang R, Ronnett BM. Incidental serous tubal intraepithelial carcinoma and early invasive serous carcinoma in the nonprophylactic setting: analysis of a case series. Am J Surg Pathol. 2015;39:442-453. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 63] [Cited by in F6Publishing: 70] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 86. | Beirne JP, McArt DG, Roddy A, McDermott C, Ferris J, Buckley NE, Coulter P, McCabe N, Eddie SL, Dunne PD, O'Reilly P, Gilmore A, Feeney L, Ewing DL, Drapkin RI, Salto-Tellez M, Kennedy RD, Harley IJG, McCluggage WG, Mullan PB. Defining the molecular evolution of extrauterine high grade serous carcinoma. Gynecol Oncol. 2019;155:305-317. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 8] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 87. | Ducie J, Dao F, Considine M, Olvera N, Shaw PA, Kurman RJ, Shih IM, Soslow RA, Cope L, Levine DA. Molecular analysis of high-grade serous ovarian carcinoma with and without associated serous tubal intra-epithelial carcinoma. Nat Commun. 2017;8:990. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 116] [Cited by in F6Publishing: 125] [Article Influence: 17.9] [Reference Citation Analysis (0)] |
| 88. | Soong TR, Howitt BE, Horowitz N, Nucci MR, Crum CP. The fallopian tube, "precursor escape" and narrowing the knowledge gap to the origins of high-grade serous carcinoma. Gynecol Oncol. 2019;152:426-433. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 49] [Cited by in F6Publishing: 57] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 89. | Kindelberger DW, Lee Y, Miron A, Hirsch MS, Feltmate C, Medeiros F, Callahan MJ, Garner EO, Gordon RW, Birch C, Berkowitz RS, Muto MG, Crum CP. Intraepithelial carcinoma of the fimbria and pelvic serous carcinoma: Evidence for a causal relationship. Am J Surg Pathol. 2007;31:161-169. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 800] [Cited by in F6Publishing: 753] [Article Influence: 44.3] [Reference Citation Analysis (0)] |
| 90. | Loyola A, Huang JY, LeRoy G, Hu S, Wang YH, Donnelly RJ, Lane WS, Lee SC, Reinberg D. Functional analysis of the subunits of the chromatin assembly factor RSF. Mol Cell Biol. 2003;23:6759-6768. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 83] [Cited by in F6Publishing: 87] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 91. | Vignali M, Hassan AH, Neely KE, Workman JL. ATP-dependent chromatin-remodeling complexes. Mol Cell Biol. 2000;20:1899-1910. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 558] [Cited by in F6Publishing: 543] [Article Influence: 22.6] [Reference Citation Analysis (0)] |
| 92. | Zhou W, Han WF, Landree LE, Thupari JN, Pinn ML, Bililign T, Kim EK, Vadlamudi A, Medghalchi SM, El Meskini R, Ronnett GV, Townsend CA, Kuhajda FP. Fatty acid synthase inhibition activates AMP-activated protein kinase in SKOV3 human ovarian cancer cells. Cancer Res. 2007;67:2964-2971. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 113] [Cited by in F6Publishing: 116] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 93. | Grunt TW, Wagner R, Grusch M, Berger W, Singer CF, Marian B, Zielinski CC, Lupu R. Interaction between fatty acid synthase- and ErbB-systems in ovarian cancer cells. Biochem Biophys Res Commun. 2009;385:454-459. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 63] [Cited by in F6Publishing: 65] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 94. | Sehdev AS, Kurman RJ, Kuhn E, Shih IeM. Serous tubal intraepithelial carcinoma upregulates markers associated with high-grade serous carcinomas including Rsf-1 (HBXAP), cyclin E and fatty acid synthase. Mod Pathol. 2010;23:844-855. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 71] [Cited by in F6Publishing: 76] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 95. | Kuhn E, Meeker A, Wang TL, Sehdev AS, Kurman RJ, Shih IeM. Shortened telomeres in serous tubal intraepithelial carcinoma: an early event in ovarian high-grade serous carcinogenesis. Am J Surg Pathol. 2010;34:829-836. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 102] [Cited by in F6Publishing: 111] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 96. | Bernardini MQ, Baba T, Lee PS, Barnett JC, Sfakianos GP, Secord AA, Murphy SK, Iversen E, Marks JR, Berchuck A. Expression signatures of TP53 mutations in serous ovarian cancers. BMC Cancer. 2010;10:237. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 36] [Cited by in F6Publishing: 40] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 97. | Cecconi S, Tatone C, Buccione R, Mangia F, Colonna R. Granulosa cell-oocyte interactions: the phosphorylation of specific proteins in mouse oocytes at the germinal vesicle stage is dependent upon the differentiative state of companion somatic cells. J Exp Zool. 1991;258:249-254. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 21] [Cited by in F6Publishing: 22] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 98. | Roh MH, Yassin Y, Miron A, Mehra KK, Mehrad M, Monte NM, Mutter GL, Nucci MR, Ning G, Mckeon FD, Hirsch MS, Wa X, Crum CP. High-grade fimbrial-ovarian carcinomas are unified by altered p53, PTEN and PAX2 expression. Mod Pathol. 2010;23:1316-1324. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 95] [Cited by in F6Publishing: 98] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 99. | Kuhn E, Kurman RJ, Vang R, Sehdev AS, Han G, Soslow R, Wang TL, Shih IeM. TP53 mutations in serous tubal intraepithelial carcinoma and concurrent pelvic high-grade serous carcinoma--evidence supporting the clonal relationship of the two lesions. J Pathol. 2012;226:421-426. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 263] [Cited by in F6Publishing: 279] [Article Influence: 21.5] [Reference Citation Analysis (0)] |
| 100. | Singh N, Gilks CB, Wilkinson N, McCluggage WG. Assignment of primary site in high-grade serous tubal, ovarian and peritoneal carcinoma: a proposal. Histopathology. 2014;65:149-154. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 55] [Cited by in F6Publishing: 54] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 101. | Singh N, Gilks CB, Wilkinson N, McCluggage WG. The secondary Müllerian system, field effect, BRCA, and tubal fimbria: our evolving understanding of the origin of tubo-ovarian high-grade serous carcinoma and why assignment of primary site matters. Pathology. 2015;47:423-431. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 39] [Cited by in F6Publishing: 39] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 102. | Singh N, Gilks CB, Hirshowitz L, Wilkinson N, McCluggage WG. Adopting a Uniform Approach to Site Assignment in Tubo-Ovarian High-Grade Serous Carcinoma: The Time has Come. Int J Gynecol Pathol. 2016;35:230-237. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 33] [Cited by in F6Publishing: 29] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 103. | Singh N, Gilks CB, Hirschowitz L, Kehoe S, McNeish IA, Miller D, Naik R, Wilkinson N, McCluggage WG. Primary site assignment in tubo-ovarian high-grade serous carcinoma: Consensus statement on unifying practice worldwide. Gynecol Oncol. 2016;141:195-198. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 50] [Cited by in F6Publishing: 53] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 104. | Pantel K, Alix-Panabières C. Circulating tumour cells in cancer patients: challenges and perspectives. Trends Mol Med. 2010;16:398-406. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 449] [Cited by in F6Publishing: 503] [Article Influence: 35.9] [Reference Citation Analysis (0)] |
| 105. | Crowley E, Di Nicolantonio F, Loupakis F, Bardelli A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol. 2013;10:472-484. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1114] [Cited by in F6Publishing: 1213] [Article Influence: 110.3] [Reference Citation Analysis (0)] |
| 106. | Heitzer E, Haque IS, Roberts CES, Speicher MR. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat Rev Genet. 2019;20:71-88. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 579] [Cited by in F6Publishing: 748] [Article Influence: 149.6] [Reference Citation Analysis (0)] |
| 107. | Khoja L, Lorigan P, Dive C, Keilholz U, Fusi A. Circulating tumour cells as tumour biomarkers in melanoma: detection methods and clinical relevance. Ann Oncol. 2015;26:33-39. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 50] [Cited by in F6Publishing: 52] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 108. | Lianidou ES, Markou A, Strati A. The Role of CTCs as Tumor Biomarkers. Adv Exp Med Biol. 2015;867:341-367. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 58] [Cited by in F6Publishing: 65] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 109. | Gold B, Cankovic M, Furtado LV, Meier F, Gocke CD. Do circulating tumor cells, exosomes, and circulating tumor nucleic acids have clinical utility? J Mol Diagn. 2015;17:209-224. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 138] [Cited by in F6Publishing: 145] [Article Influence: 18.1] [Reference Citation Analysis (0)] |
| 110. | Smith B, Selby P, Southgate J, Pittman K, Bradley C, Blair GE. Detection of melanoma cells in peripheral blood by means of reverse transcriptase and polymerase chain reaction. Lancet. 1991;338:1227-1229. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 465] [Cited by in F6Publishing: 490] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 111. | Cristofanilli M, Hayes DF, Budd GT, Ellis MJ, Stopeck A, Reuben JM, Doyle GV, Matera J, Allard WJ, Miller MC, Fritsche HA, Hortobagyi GN, Terstappen LW. Circulating tumor cells: a novel prognostic factor for newly diagnosed metastatic breast cancer. J Clin Oncol. 2005;23:1420-1430. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 820] [Cited by in F6Publishing: 798] [Article Influence: 42.0] [Reference Citation Analysis (0)] |
| 112. | de Bono JS, Scher HI, Montgomery RB, Parker C, Miller MC, Tissing H, Doyle GV, Terstappen LW, Pienta KJ, Raghavan D. Circulating tumor cells predict survival benefit from treatment in metastatic castration-resistant prostate cancer. Clin Cancer Res. 2008;14:6302-6309. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1588] [Cited by in F6Publishing: 1639] [Article Influence: 102.4] [Reference Citation Analysis (0)] |
| 113. | Cohen SJ, Punt CJ, Iannotti N, Saidman BH, Sabbath KD, Gabrail NY, Picus J, Morse M, Mitchell E, Miller MC, Doyle GV, Tissing H, Terstappen LW, Meropol NJ. Relationship of circulating tumor cells to tumor response, progression-free survival, and overall survival in patients with metastatic colorectal cancer. J Clin Oncol. 2008;26:3213-3221. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1354] [Cited by in F6Publishing: 1353] [Article Influence: 84.6] [Reference Citation Analysis (0)] |
| 114. | Liu JF, Kindelberger D, Doyle C, Lowe A, Barry WT, Matulonis UA. Predictive value of circulating tumor cells (CTCs) in newly-diagnosed and recurrent ovarian cancer patients. Gynecol Oncol. 2013;131:352-356. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 55] [Cited by in F6Publishing: 62] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 115. | Zhou Y, Bian B, Yuan X, Xie G, Ma Y, Shen L. Prognostic Value of Circulating Tumor Cells in Ovarian Cancer: A Meta-Analysis. PLoS One. 2015;10:e0130873. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 36] [Cited by in F6Publishing: 42] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 116. | Bankó P, Lee SY, Nagygyörgy V, Zrínyi M, Chae CH, Cho DH, Telekes A. Technologies for circulating tumor cell separation from whole blood. J Hematol Oncol. 2019;12:48. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 130] [Cited by in F6Publishing: 183] [Article Influence: 36.6] [Reference Citation Analysis (0)] |
| 117. | Mandel P, Metais P. Nuclear Acids In Human Blood Plasma. C R Seances Soc Biol Fil. 1948;142:241-243. [PubMed] [Cited in This Article: ] |
| 118. | Fleischhacker M, Schmidt B. Circulating nucleic acids (CNAs) and cancer--a survey. Biochim Biophys Acta. 2007;1775:181-232. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 226] [Cited by in F6Publishing: 418] [Article Influence: 23.2] [Reference Citation Analysis (0)] |
| 119. | Diaz LA Jr, Williams RT, Wu J, Kinde I, Hecht JR, Berlin J, Allen B, Bozic I, Reiter JG, Nowak MA, Kinzler KW, Oliner KS, Vogelstein B. The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers. Nature. 2012;486:537-540. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1259] [Cited by in F6Publishing: 1292] [Article Influence: 107.7] [Reference Citation Analysis (0)] |
| 120. | Francis G, Stein S. Circulating Cell-Free Tumour DNA in the Management of Cancer. Int J Mol Sci. 2015;16:14122-14142. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 70] [Cited by in F6Publishing: 83] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 121. | Stroun M, Lyautey J, Lederrey C, Olson-Sand A, Anker P. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin Chim Acta. 2001;313:139-142. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 516] [Cited by in F6Publishing: 536] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 122. | Anker P, Lefort F, Vasioukhin V, Lyautey J, Lederrey C, Chen XQ, Stroun M, Mulcahy HE, Farthing MJ. K-ras mutations are found in DNA extracted from the plasma of patients with colorectal cancer. Gastroenterology. 1997;112:1114-1120. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 176] [Cited by in F6Publishing: 185] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 123. | Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, Knippers R. DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001;61:1659-1665. [PubMed] [Cited in This Article: ] |
| 124. | Lo YM, Chan KC, Sun H, Chen EZ, Jiang P, Lun FM, Zheng YW, Leung TY, Lau TK, Cantor CR, Chiu RW. Maternal plasma DNA sequencing reveals the genome-wide genetic and mutational profile of the fetus. Sci Transl Med. 2010;2:61ra91. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 634] [Cited by in F6Publishing: 689] [Article Influence: 53.0] [Reference Citation Analysis (0)] |
| 125. | Mouliere F, Robert B, Arnau Peyrotte E, Del Rio M, Ychou M, Molina F, Gongora C, Thierry AR. High fragmentation characterizes tumour-derived circulating DNA. PLoS One. 2011;6:e23418. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 362] [Cited by in F6Publishing: 373] [Article Influence: 28.7] [Reference Citation Analysis (0)] |
| 126. | Bronkhorst AJ, Ungerer V, Holdenrieder S. The emerging role of cell-free DNA as a molecular marker for cancer management. Biomol Detect Quantif. 2019;17:100087. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 318] [Cited by in F6Publishing: 300] [Article Influence: 60.0] [Reference Citation Analysis (0)] |
| 127. | Jung M, Klotzek S, Lewandowski M, Fleischhacker M, Jung K. Changes in concentration of DNA in serum and plasma during storage of blood samples. Clin Chem. 2003;49:1028-1029. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 173] [Cited by in F6Publishing: 176] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 128. | Esposito A, Criscitiello C, Trapani D, Curigliano G. The Emerging Role of "Liquid Biopsies," Circulating Tumor Cells, and Circulating Cell-Free Tumor DNA in Lung Cancer Diagnosis and Identification of Resistance Mutations. Curr Oncol Rep. 2017;19:1. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 33] [Cited by in F6Publishing: 27] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 129. | Parkinson CA, Gale D, Piskorz AM, Biggs H, Hodgkin C, Addley H, Freeman S, Moyle P, Sala E, Sayal K, Hosking K, Gounaris I, Jimenez-Linan M, Earl HM, Qian W, Rosenfeld N, Brenton JD. Exploratory Analysis of TP53 Mutations in Circulating Tumour DNA as Biomarkers of Treatment Response for Patients with Relapsed High-Grade Serous Ovarian Carcinoma: A Retrospective Study. PLoS Med. 2016;13:e1002198. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 202] [Cited by in F6Publishing: 186] [Article Influence: 23.3] [Reference Citation Analysis (0)] |
| 130. | Kamat AA, Sood AK, Dang D, Gershenson DM, Simpson JL, Bischoff FZ. Quantification of total plasma cell-free DNA in ovarian cancer using real-time PCR. Ann N Y Acad Sci. 2006;1075:230-234. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 47] [Cited by in F6Publishing: 50] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 131. | Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, Bartlett BR, Wang H, Luber B, Alani RM, Antonarakis ES, Azad NS, Bardelli A, Brem H, Cameron JL, Lee CC, Fecher LA, Gallia GL, Gibbs P, Le D, Giuntoli RL, Goggins M, Hogarty MD, Holdhoff M, Hong SM, Jiao Y, Juhl HH, Kim JJ, Siravegna G, Laheru DA, Lauricella C, Lim M, Lipson EJ, Marie SK, Netto GJ, Oliner KS, Olivi A, Olsson L, Riggins GJ, Sartore-Bianchi A, Schmidt K, Shih lM, Oba-Shinjo SM, Siena S, Theodorescu D, Tie J, Harkins TT, Veronese S, Wang TL, Weingart JD, Wolfgang CL, Wood LD, Xing D, Hruban RH, Wu J, Allen PJ, Schmidt CM, Choti MA, Velculescu VE, Kinzler KW, Vogelstein B, Papadopoulos N, Diaz LA Jr. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014;6:224ra24. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2770] [Cited by in F6Publishing: 3117] [Article Influence: 311.7] [Reference Citation Analysis (0)] |
| 132. | Kamat AA, Baldwin M, Urbauer D, Dang D, Han LY, Godwin A, Karlan BY, Simpson JL, Gershenson DM, Coleman RL, Bischoff FZ, Sood AK. Plasma cell-free DNA in ovarian cancer: an independent prognostic biomarker. Cancer. 2010;116:1918-1925. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 116] [Cited by in F6Publishing: 123] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 133. | Shao X, He Y, Ji M, Chen X, Qi J, Shi W, Hao T, Ju S. Quantitative analysis of cell-free DNA in ovarian cancer. Oncol Lett. 2015;10:3478-3482. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 35] [Cited by in F6Publishing: 38] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 134. | No JH, Kim K, Park KH, Kim YB. Cell-free DNA level as a prognostic biomarker for epithelial ovarian cancer. Anticancer Res. 2012;32:3467-3471. [PubMed] [Cited in This Article: ] |
| 135. | Swisher EM, Wollan M, Mahtani SM, Willner JB, Garcia R, Goff BA, King MC. Tumor-specific p53 sequences in blood and peritoneal fluid of women with epithelial ovarian cancer. Am J Obstet Gynecol. 2005;193:662-667. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 84] [Cited by in F6Publishing: 96] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 136. | Dobrzycka B, Terlikowski SJ, Kinalski M, Kowalczuk O, Niklinska W, Chyczewski L. Circulating free DNA and p53 antibodies in plasma of patients with ovarian epithelial cancers. Ann Oncol. 2011;22:1133-1140. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 25] [Cited by in F6Publishing: 28] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 137. | Pereira E, Camacho-Vanegas O, Anand S, Sebra R, Catalina Camacho S, Garnar-Wortzel L, Nair N, Moshier E, Wooten M, Uzilov A, Chen R, Prasad-Hayes M, Zakashansky K, Beddoe AM, Schadt E, Dottino P, Martignetti JA. Personalized Circulating Tumor DNA Biomarkers Dynamically Predict Treatment Response and Survival In Gynecologic Cancers. PLoS One. 2015;10:e0145754. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 95] [Cited by in F6Publishing: 117] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 138. | Gorgannezhad L, Umer M, Islam MN, Nguyen NT, Shiddiky MJA. Circulating tumor DNA and liquid biopsy: opportunities, challenges, and recent advances in detection technologies. Lab Chip. 2018;18:1174-1196. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 185] [Cited by in F6Publishing: 141] [Article Influence: 23.5] [Reference Citation Analysis (0)] |
| 139. | Petrelli F, Borgonovo K, Cabiddu M, Barni S. Efficacy of EGFR tyrosine kinase inhibitors in patients with EGFR-mutated non-small-cell lung cancer: a meta-analysis of 13 randomized trials. Clin Lung Cancer. 2012;13:107-114. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 88] [Cited by in F6Publishing: 93] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 140. | Flaherty KT, Puzanov I, Kim KB, Ribas A, McArthur GA, Sosman JA, O'Dwyer PJ, Lee RJ, Grippo JF, Nolop K, Chapman PB. Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med. 2010;363:809-819. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2808] [Cited by in F6Publishing: 2725] [Article Influence: 194.6] [Reference Citation Analysis (0)] |
| 141. | Schmiegel W, Scott RJ, Dooley S, Lewis W, Meldrum CJ, Pockney P, Draganic B, Smith S, Hewitt C, Philimore H, Lucas A, Shi E, Namdarian K, Chan T, Acosta D, Ping-Chang S, Tannapfel A, Reinacher-Schick A, Uhl W, Teschendorf C, Wolters H, Stern J, Viebahn R, Friess H, Janssen KP, Nitsche U, Slotta-Huspenina J, Pohl M, Vangala D, Baraniskin A, Dockhorn-Dworniczak B, Hegewisch-Becker S, Ronga P, Edelstein DL, Jones FS, Hahn S, Fox SB. Blood-based detection of RAS mutations to guide anti-EGFR therapy in colorectal cancer patients: concordance of results from circulating tumor DNA and tissue-based RAS testing. Mol Oncol. 2017;11:208-219. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 96] [Cited by in F6Publishing: 111] [Article Influence: 15.9] [Reference Citation Analysis (0)] |
| 142. | Thierry AR, Pastor B, Jiang ZQ, Katsiampoura AD, Parseghian C, Loree JM, Overman MJ, Sanchez C, Messaoudi SE, Ychou M, Kopetz S. Circulating DNA Demonstrates Convergent Evolution and Common Resistance Mechanisms during Treatment of Colorectal Cancer. Clin Cancer Res. 2017;23:4578-4591. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 66] [Cited by in F6Publishing: 61] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 143. | Murtaza M, Dawson SJ, Tsui DW, Gale D, Forshew T, Piskorz AM, Parkinson C, Chin SF, Kingsbury Z, Wong AS, Marass F, Humphray S, Hadfield J, Bentley D, Chin TM, Brenton JD, Caldas C, Rosenfeld N. Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature. 2013;497:108-112. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1288] [Cited by in F6Publishing: 1252] [Article Influence: 113.8] [Reference Citation Analysis (0)] |
| 144. | Han X, Wang J, Sun Y. Circulating Tumor DNA as Biomarkers for Cancer Detection. Genomics Proteomics Bioinformatics. 2017;15:59-72. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 165] [Cited by in F6Publishing: 146] [Article Influence: 20.9] [Reference Citation Analysis (0)] |
| 145. | Capizzi E, Gabusi E, Grigioni AD, De Iaco P, Rosati M, Zamagni C, Fiorentino M. Quantification of free plasma DNA before and after chemotherapy in patients with advanced epithelial ovarian cancer. Diagn Mol Pathol. 2008;17:34-38. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 23] [Cited by in F6Publishing: 27] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 146. | Greaves M, Maley CC. Clonal evolution in cancer. Nature. 2012;481:306-313. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2054] [Cited by in F6Publishing: 2016] [Article Influence: 168.0] [Reference Citation Analysis (0)] |
| 147. | Ishii H, Azuma K, Sakai K, Kawahara A, Yamada K, Tokito T, Okamoto I, Nishio K, Hoshino T. Digital PCR analysis of plasma cell-free DNA for non-invasive detection of drug resistance mechanisms in EGFR mutant NSCLC: Correlation with paired tumor samples. Oncotarget. 2015; 6:30850-30858. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 57] [Cited by in F6Publishing: 65] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 148. | Misale S, Yaeger R, Hobor S, Scala E, Janakiraman M, Liska D, Valtorta E, Schiavo R, Buscarino M, Siravegna G, Bencardino K, Cercek A, Chen CT, Veronese S, Zanon C, Sartore-Bianchi A, Gambacorta M, Gallicchio M, Vakiani E, Boscaro V, Medico E, Weiser M, Siena S, Di Nicolantonio F, Solit D, Bardelli A. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature. 2012;486:532-536. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1313] [Cited by in F6Publishing: 1373] [Article Influence: 114.4] [Reference Citation Analysis (0)] |
| 149. | Lin KK, Harrell MI, Oza AM, Oaknin A, Ray-Coquard I, Tinker AV, Helman E, Radke MR, Say C, Vo LT, Mann E, Isaacson JD, Maloney L, O'Malley DM, Chambers SK, Kaufmann SH, Scott CL, Konecny GE, Coleman RL, Sun JX, Giordano H, Brenton JD, Harding TC, McNeish IA, Swisher EM. BRCA Reversion Mutations in Circulating Tumor DNA Predict Primary and Acquired Resistance to the PARP Inhibitor Rucaparib in High-Grade Ovarian Carcinoma. Cancer Discov. 2019;9:210-219. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 259] [Cited by in F6Publishing: 243] [Article Influence: 48.6] [Reference Citation Analysis (0)] |
| 150. | Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, Thornton K, Agrawal N, Sokoll L, Szabo SA, Kinzler KW, Vogelstein B, Diaz LA Jr. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985-990. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1719] [Cited by in F6Publishing: 1887] [Article Influence: 111.0] [Reference Citation Analysis (0)] |
| 151. | Chaudhuri AA, Chabon JJ, Lovejoy AF, Newman AM, Stehr H, Azad TD, Khodadoust MS, Esfahani MS, Liu CL, Zhou L, Scherer F, Kurtz DM, Say C, Carter JN, Merriott DJ, Dudley JC, Binkley MS, Modlin L, Padda SK, Gensheimer MF, West RB, Shrager JB, Neal JW, Wakelee HA, Loo BW Jr, Alizadeh AA, Diehn M. Early Detection of Molecular Residual Disease in Localized Lung Cancer by Circulating Tumor DNA Profiling. Cancer Discov. 2017;7:1394-1403. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 419] [Cited by in F6Publishing: 602] [Article Influence: 86.0] [Reference Citation Analysis (0)] |
| 152. | Kinde I, Bettegowda C, Wang Y, Wu J, Agrawal N, Shih IeM, Kurman R, Dao F, Levine DA, Giuntoli R, Roden R, Eshleman JR, Carvalho JP, Marie SK, Papadopoulos N, Kinzler KW, Vogelstein B, Diaz LA Jr. Evaluation of DNA from the Papanicolaou test to detect ovarian and endometrial cancers. Sci Transl Med. 2013;5:167ra4. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 223] [Cited by in F6Publishing: 231] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 153. | Cohen PA, Flowers N, Tong S, Hannan N, Pertile MD, Hui L. Abnormal plasma DNA profiles in early ovarian cancer using a non-invasive prenatal testing platform: implications for cancer screening. BMC Med. 2016;14:126. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 59] [Cited by in F6Publishing: 61] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 154. | Cohen JD, Li L, Wang Y, Thoburn C, Afsari B, Danilova L, Douville C, Javed AA, Wong F, Mattox A, Hruban RH, Wolfgang CL, Goggins MG, Dal Molin M, Wang TL, Roden R, Klein AP, Ptak J, Dobbyn L, Schaefer J, Silliman N, Popoli M, Vogelstein JT, Browne JD, Schoen RE, Brand RE, Tie J, Gibbs P, Wong HL, Mansfield AS, Jen J, Hanash SM, Falconi M, Allen PJ, Zhou S, Bettegowda C, Diaz LA Jr, Tomasetti C, Kinzler KW, Vogelstein B, Lennon AM, Papadopoulos N. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science. 2018;359:926-930. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1752] [Cited by in F6Publishing: 1552] [Article Influence: 258.7] [Reference Citation Analysis (0)] |
| 155. | Bianchi DW, Chudova D, Sehnert AJ, Bhatt S, Murray K, Prosen TL, Garber JE, Wilkins-Haug L, Vora NL, Warsof S, Goldberg J, Ziainia T, Halks-Miller M. Noninvasive Prenatal Testing and Incidental Detection of Occult Maternal Malignancies. JAMA. 2015;314:162-169. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 274] [Cited by in F6Publishing: 273] [Article Influence: 30.3] [Reference Citation Analysis (0)] |
| 156. | Gormally E, Vineis P, Matullo G, Veglia F, Caboux E, Le Roux E, Peluso M, Garte S, Guarrera S, Munnia A, Airoldi L, Autrup H, Malaveille C, Dunning A, Overvad K, Tjønneland A, Lund E, Clavel-Chapelon F, Boeing H, Trichopoulou A, Palli D, Krogh V, Tumino R, Panico S, Bueno-de-Mesquita HB, Peeters PH, Pera G, Martinez C, Dorronsoro M, Barricarte A, Navarro C, Quirós JR, Hallmans G, Day NE, Key TJ, Saracci R, Kaaks R, Riboli E, Hainaut P. TP53 and KRAS2 mutations in plasma DNA of healthy subjects and subsequent cancer occurrence: a prospective study. Cancer Res. 2006;66:6871-6876. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 121] [Cited by in F6Publishing: 127] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 157. | Wang Y, Li L, Douville C, Cohen JD, Yen TT, Kinde I, Sundfelt K, Kjær SK, Hruban RH, Shih IM, Wang TL, Kurman RJ, Springer S, Ptak J, Popoli M, Schaefer J, Silliman N, Dobbyn L, Tanner EJ, Angarita A, Lycke M, Jochumsen K, Afsari B, Danilova L, Levine DA, Jardon K, Zeng X, Arseneau J, Fu L, Diaz LA Jr, Karchin R, Tomasetti C, Kinzler KW, Vogelstein B, Fader AN, Gilbert L, Papadopoulos N. Evaluation of liquid from the Papanicolaou test and other liquid biopsies for the detection of endometrial and ovarian cancers. Sci Transl Med. 2018;10. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 125] [Cited by in F6Publishing: 151] [Article Influence: 30.2] [Reference Citation Analysis (0)] |
| 158. | Jan M, Ebert BL, Jaiswal S. Clonal hematopoiesis. Semin Hematol. 2017;54:43-50. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 79] [Cited by in F6Publishing: 80] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 159. | Dondorp W, de Wert G, Bombard Y, Bianchi DW, Bergmann C, Borry P, Chitty LS, Fellmann F, Forzano F, Hall A, Henneman L, Howard HC, Lucassen A, Ormond K, Peterlin B, Radojkovic D, Rogowski W, Soller M, Tibben A, Tranebjærg L, van El CG, Cornel MC; European Society of Human Genetics; American Society of Human Genetics. Non-invasive prenatal testing for aneuploidy and beyond: challenges of responsible innovation in prenatal screening. Eur J Hum Genet. 2015;23:1438-1450. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 219] [Cited by in F6Publishing: 185] [Article Influence: 20.6] [Reference Citation Analysis (0)] |
| 160. | Forshew T, Murtaza M, Parkinson C, Gale D, Tsui DW, Kaper F, Dawson SJ, Piskorz AM, Jimenez-Linan M, Bentley D, Hadfield J, May AP, Caldas C, Brenton JD, Rosenfeld N. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci Transl Med. 2012;4: 136ra68. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 974] [Cited by in F6Publishing: 948] [Article Influence: 79.0] [Reference Citation Analysis (0)] |
| 161. | Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF, Dunning MJ, Gale D, Forshew T, Mahler-Araujo B, Rajan S, Humphray S, Becq J, Halsall D, Wallis M, Bentley D, Caldas C, Rosenfeld N. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013;368:1199-1209. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1713] [Cited by in F6Publishing: 1600] [Article Influence: 145.5] [Reference Citation Analysis (0)] |
| 162. | Giannopoulou L, Mastoraki S, Buderath P, Strati A, Pavlakis K, Kasimir-Bauer S, Lianidou ES. ESR1 methylation in primary tumors and paired circulating tumor DNA of patients with high-grade serous ovarian cancer. Gynecol Oncol. 2018;150:355-360. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 30] [Cited by in F6Publishing: 36] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 163. | Giannopoulou L, Chebouti I, Pavlakis K, Kasimir-Bauer S, Lianidou ES. RASSF1A promoter methylation in high-grade serous ovarian cancer: A direct comparison study in primary tumors, adjacent morphologically tumor cell-free tissues and paired circulating tumor DNA. Oncotarget. 2017;8:21429-21443. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 38] [Cited by in F6Publishing: 49] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 164. | Widschwendter M, Zikan M, Wahl B, Lempiäinen H, Paprotka T, Evans I, Jones A, Ghazali S, Reisel D, Eichner J, Rujan T, Yang Z, Teschendorff AE, Ryan A, Cibula D, Menon U, Wittenberger T. The potential of circulating tumor DNA methylation analysis for the early detection and management of ovarian cancer. Genome Med. 2017;9:116. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 84] [Cited by in F6Publishing: 102] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 165. | Pepe MS, Feng Z, Janes H, Bossuyt PM, Potter JD. Pivotal evaluation of the accuracy of a biomarker used for classification or prediction: standards for study design. J Natl Cancer Inst. 2008;100:1432-1438. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 540] [Cited by in F6Publishing: 505] [Article Influence: 31.6] [Reference Citation Analysis (0)] |
| 166. | Broncy L, Paterlini-Bréchot P. Circulating Tumor Cells for the Management of Renal Cell Carcinoma. Diagnostics (Basel). 2018;8. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 7] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 167. | Muinelo-Romay L, Casas-Arozamena C, Abal M. Liquid Biopsy in Endometrial Cancer: New Opportunities for Personalized Oncology. Int J Mol Sci. 2018;19. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 66] [Cited by in F6Publishing: 61] [Article Influence: 10.2] [Reference Citation Analysis (0)] |