Published online Apr 8, 2016. doi: 10.4254/wjh.v8.i10.485
Peer-review started: September 15, 2015
First decision: November 13, 2015
Revised: December 4, 2015
Accepted: March 7, 2016
Article in press: March 9, 2016
Published online: April 8, 2016
Hepatocellular carcinoma (HCC) is etiologically linked with hepatitis B virus (HBV) and is the leading cause of death amongst 80% of HBV patients. Among HBV affected patients, genetic factors are also involved in modifying the risk factors of HCC. However, the genetic factors that regulate progression to HCC still remain to be determined. In this review, we discuss several single nucleotide polymorphisms (SNPs) which were reportedly associated with increased or reduced risk of HCC occurrence in patients with chronic HBV infection such as cyclooxygenase (COX)-2 expression specifically at COX-2 -1195G/A in Chinese, Turkish and Egyptian populations, tumor necrosis factor α and the three most commonly studied SNPs: PAT-/+, Lys939Gln (A33512C, rs2228001) and Ala499Val (C21151T, rs2228000). In genome-wide association studies, strong associations have also been found at loci 1p36.22, 11q22.3, 6p21 (rs1419881, rs3997872, rs7453920 and rs7768538), 8p12 (rs2275959 and rs37821974) and 22q11.21. The genes implicated in these studies include HLA-DQB2, HLA-DQA1, TCF19, HLA-C, UBE2L3, LTL, FDX1, MICA, UBE4B and PG. The SNPs found to be associated with the above-mentioned genes still require validation in association studies in order to be considered good prognostic candidates for HCC. Screening of these polymorphisms is very beneficial in clinical experiments to stratify the higher or lower risk for HCC and may help in designing effective and efficient HCC surveillance programs for chronic HBV-infected patients if further genetic vulnerabilities are detected.
Core tip: In this review, we discuss various common associations between hepatitis B virus (HBV) and host polymorphisms. These single nucleotide polymorphisms which have been found to be associated with various genes still require validation in association studies in order to be considered good prognostic candidates for hepatocellular carcinoma (HCC). Screening of these polymorphisms is very beneficial in clinical experiments to stratify the higher or lower risk for HCC and may help in designing effective and efficient HCC surveillance programs for chronic HBV-infected patients if further genetic vulnerabilities are detected.
- Citation: Mathew S, Abdel-Hafiz H, Raza A, Fatima K, Qadri I. Host nucleotide polymorphism in hepatitis B virus-associated hepatocellular carcinoma. World J Hepatol 2016; 8(10): 485-498
- URL: https://www.wjgnet.com/1948-5182/full/v8/i10/485.htm
- DOI: https://dx.doi.org/10.4254/wjh.v8.i10.485
Hepatitis B virus (HBV) infection is the third most common cause of cancer-related deaths in relation to hepatocellular carcinoma (HCC) with a high incidence in Asian countries. HCC is responsible for approximately 660000 deaths worldwide each year and 85%-90% of these deaths are due to primary liver cancers[1]. It is recognized that these cancers are mainly due to HBV infection with 60% of HCC cases seropositive for this virus[2]. Many risk factors including viral factors (e.g., genomic mutations, genotypes, HBV-DNA levels), host factors and unhealthy lifestyles all contribute to the development of liver diseases[3].
Both epigenetic and genetic factors play a role in the malignant transformation of liver cells[4]. Multiple cellular signaling genes are enhanced by the incorporation of HBV into the host’s genome which promotes transactivation of HBx protein[5]. This process activates/inactivates suppressor genes (e.g., p53), oncogenic genes (e.g., c-fos and c-myc), induces loss of heterozygosity and activates transcriptional factors [e.g., nuclear factor kappa-B (NF-κB) and AP-1][6].
However, underlying disease and the duration of severity vary significantly between each phase. Moreover, clinical progression varies between patients. Liver injuries in patients with HBV infection are thought to be the outcome of the host’s immune responses against HBV. For example, cytotoxic T lymphocyte-mediated, an HLA-class I antigen-restricted, response to the HBV antigen expressed on hepatocytes results in necrosis and apoptosis[7].
Several genome wide association studies have identified candidate single nucleotide polymorphisms (SNPs) by comparing the SNPs present in HCC patients and those present in asymptomatic HBV carriers[8]. Therefore, to specifically evaluate genetic factors, it is vital that the controls and patients are well matched regarding these factors to identify the correct SNP. The results of many studies suggest that several SNPs are associated with HBV clearance and persistent infection. Functional analyses are necessary to confirm these results[6,7]. In this review, we discuss several SNPs which are reportedly associated with increased or reduced risk of HCC occurrence in patients with chronic HBV infection[9].
It has been reported previously that SNPs can affect disease progression after HBV infection. Cytokines, such as tumor necrosis factor-α (TNFα) and interleukin (IL)-10, have a significant role in regulating viral infection. Genetic variation of these cytokines is linked with the outcome of HBV infection[10-16].
Several studies have shown that genetic polymorphisms in multiple genes such as TP53[17], IL-6[18], and DNA repair genes[19], are associated with the development of chronic HBC infection, progression of the infection and increased risk of HCC. These may serve as biomarkers in identifying HCC risk[20]. However, these studies were predominantly performed in HBV-positive populations or populations with a high infection rate.
Genetic variation in tumor suppressor genes or oncogenes is capable of altering gene function and, consequently, may contribute to the development of cancer. Significant research has been conducted to investigate the association between polymorphisms in tumor suppressor genes and oncogenes and the risk of HCC; however, the results are controversial.
Cyclooxygenase-2 (COX-2) is involved in many cellular functions, including inflammation, inhibition of apoptosis, carcinogenesis, angiogenesis, invasion and metastasis[21,22]. COX-2 is overexpressed in many cancers including HCC, indicating that there is an association between COX-2 expression and the development of cancer[23,24]. Selective COX-2 inhibitors have been shown to suppress the growth of HCC cells in vitro and in vivo[25]. A polymorphism in the promoter region of the COX-2 gene could functionally upregulate the transcriptional activity of COX-2, indicating a possible mechanism by which COX-2 may contribute to genetic susceptibility to HCC[21]. Several studies have reported that COX-2 point mutations including -1195G/A, -765G/C and +8473T/C were correlated with liver diseases and HBV-related HCC[26]. COX-2-765G/C is related to the risk of skin, esophageal, colorectal, breast and gastric cancers[27-29]. With regard to HCC, contradictory and inconclusive results were found. Some studies have reported a correlation between COX-2-765G/C and HBV-related HCC risk[30-32], but other studies reported that no such correlation exists[26,33,34]. It has been reported that these inconsistent results were possibly due to limited sample sizes and ethnic variation in those studies. COX-2 + 8473T/C is associated with oral and breast cancers[35,36], but is not associated with HCC[37]. A recent meta-analysis by Chen et al[26] on Chinese, Turkish and Egyptian populations, concluded that COX-2-1195G/A may be associated with HCC risk, but not COX-2-765G/C or COX-2 + 847T/C.
IL-1α is a potent pro-inflammatory cytokine and has many different biological functions, including cell survival, proliferation, and anti-apoptosis[38,39]. IL-1β is also reported to inhibit interferon-induced antiviral activity[40] and is assumed to be closely associated with the pathogenesis of chronic hepatitis C. Several polymorphisms of the IL-1 gene that are thought to affect IL-1β production have been reported[41]. -31T SNPs of IL-1β have been shown to enhance IL-1β transcriptional activity[42] and several studies reported that -511C/-31T is a risk factor for the development of cancer and liver diseases[43-45]. Wang et al[41] showed that IL-1β-31 polymorphism was associated with HCC, after controlling for other confounding clinical parameters.
E-cadherin is a transmembrane protein that mediates cell-cell adhesion and is expressed in most normal epithelial cells. Downregulation of E-cadherin may lead to a loss of E-cadherin-mediated adhesion, resulting in increased susceptibility to tumor development and is associated with poor prognosis in various carcinomas including HCC[45-52]. In addition, HBV and HCV reduce E-cadherin expression and promote tumor recurrence in HCC patients. One of the mechanisms that have been proposed for reduced E-cadherin expression is SNPs in the promoter region of the CDH1 gene. CDH1-160 C/A and -347G/GA polymorphisms result in the downregulation of E-cadherin protein and is associated with cancer susceptibility[53]. Several studies demonstrated that CDH1-347 SNPs are significantly associated with HCC risk[52,54-57]. However, the correlation between CDH1-160 SNPs showed conflicting results. Some studies[58,59] have shown that CDH1-160 SNP carriers have an increased risk of prostate and bladder cancer, while others showed that it was not associated with the development of prostate, HCC, colorectal or gastric cancer[60].
Peroxisome proliferator-activated receptor gamma (PPARγ) is a hormone receptor, present in adipose tissue and plays a critical role in the regulation of fatty acid storage and glucose metabolism[61]. PPARγ has been shown to be associated with type 2 diabetes mellitus (T2DM)[62]. PPARγ contains two isoforms, PPARγ1 and PPARγ2 and several variants in the PPARγ gene have been identified[63]. The A allele of PPARγ2 is associated with a significant decrease in the development of T2DM[64]. The relationship between PPAR and HCC is not clear. Although experimental studies have shown that PPAR may have a role in HCC[65,66], the implications of these findings are unclear. Koytak et al[66] investigated the effect of the PPARα L162V polymorphism on clinical outcome in a patient with HCC caused by hepatitis viruses. They concluded that there was a relationship between the PPARα L162V polymorphism and HBV-induced HCC and was associated with advanced HCC. This polymorphism was shown to enhance PPARα transcriptional activity and is associated with lipid abnormalities and an increased body mass index[67-70].
TNFα-inducible protein 3 (TNFαIP3), a cytoplasmic zinc finger protein with ubiquitin-modifying activity, has been shown to inhibit NF-κB activity and TNF-mediated apoptosis[71-74]. TNFαIP3 polymorphisms have been linked to inflammatory, autoimmune and malignant diseases. A recent study reported that there was no association between TNFαIP3 rs2230926 polymorphism and susceptibility to chronic HBV infection or the progression of HBV-related diseases[75].
Cytotoxic T lymphocyte-associated factor 4 (CTLA-4) is a protein receptor expressed in T cells and it functions as a negative regulator of the immune system. Several CTLA-4 gene polymorphisms have been identified including -318C>T, A49G and CT60[76]. CTLA-4 polymorphisms are associated with several autoimmune diseases, including thyroid and liver diseases[77,78]. It has been shown that SNPs in CTLA-4 may be associated with HBV progression and viral persistence[79]. CTLA-4 SNPs can be used as a marker for predicting treatment outcome in chronic HCV-infected patients[80-82].
TNFα is a multifunctional cytokine that regulates the inflammatory reaction and has an important role in the development and progression of a number of diseases, including liver disease[83,84]. It has been suggested that genetic polymorphisms of TNFα may contribute to the pathogenesis of liver diseases, infectious diseases and inflammatory disorders[43,85]. For example, TNFα SNPs affect TNFα production leading to a greater risk of HCC. The polymorphism at site -1031T/C, -863C/A, -857C/T, -376, -308G/A and -238G/A of the TNFα promoter is associated with the outcome of HBV infection and disease progression[86-89].
IL-10 is an important anti-inflammatory cytokine produced in macrophages. Three SNPs in the IL-10 gene promoter, at -1082, -819 and -592, are associated with IL-10 production and secretion by peripheral blood monocytes. It has been shown that IL-10-592 A/C polymorphism was associated with susceptibility to HBV infection[90].
The glutathione S-transferases (GSTs) enzymes play an important role in maintaining the cellular defense mechanism against the effects of reactive oxygen species and various exogenous toxins, and have been shown to be overexpressed in several cancers[91,92]. Deletion polymorphism of GST genes results in diminished enzyme activity leading to the insufficient defense of cells from metabolites and free radicals, elevated concentration of endogenous mutagens and a high risk of various tumors, including HCC[93-96]. GSTs polymorphisms have been shown to be associated with colorectal cancer, lung cancer, squamous cell carcinoma of the head and neck, HBV-related HCC, and various urogenital and gastrointestinal disorders[97-99]. For example, meta-analyses have shown that GSTM1, GSTP1 and GSTT1 are associated with an increased risk of HCC[100,101].
Epidermal growth factor (EGF) and its respective receptor (EGFR) signaling are important regulators of proliferation and the pathogenesis of many human carcinomas[102,103]. Upon ligand binding, the two EGFR domains undergo trans-autophosphorylation at specific tyrosine residues[104]. These phosphotyrosines are recognized by Src homology 2 domain containing proteins[105] and activate a diverse signaling network that includes the RAS/extracellular signal-regulated kinase pathway[106], the phosphatidylinositol 3-kinase pathway[107] and the Janus kinase/Signal transducer and activator of transcription pathway[108].
Activation of EGF has also been shown to be required for hepatocyte growth during liver regeneration[109]. In addition, many viruses such as Epstein Barr virus and HBV can tweak EGF receptor expression in their favor[110-112]. The role of EGF polymorphism has been explored in numerous meta-analyses[113-116] and was shown to be highly associated with susceptibility to HCC[117]. Prominent among these is the EGF + 61A > G transversion (rs4444903) which was shown to regulate expression of the EGF gene[118,119]. This SNP is found in the 5’ untranslated regions of the EGF gene and was shown in cell lines to enhance the stability of EGF mRNA[119]. The G/G allele is associated with higher serum levels of EGF compared with the A/A allele[119,120]. Numerous follow-up studies have validated the positive association between this G/G and G/A genotype with HCC in diverse genetic populations[117,121-123] and thus can be considered a good prognostic marker for the genetically susceptible population.
Murine double minute 2 (MDM2) is a ubiquitin ligase that controls the turnover rate of an important tumor suppressor, p53, which is deleted or mutated in 50% of all human tumors[124]. P53 is also referred to as the guardian of the genome because it can activate DNA repair pathways[125], arrest cell cycle at the G1/S regulation checkpoint[126] or initiate apoptosis if the damage cannot be repaired[127]. All these important networks converge in the active form of p53, which is kept in check by MDM2. The addition of ubiquitin subunits to critical lysine residues transfers the active p53 to 26S proteasome for degradation along with MDM2[128,129]. In addition, the binding of MDM2 can block p53-mediated transactivation functions[130]. The activity of MDM2 protein is equally important in regulating this DNA repair-cell cycle-apoptosis nexus and variation in the expression levels of this protein was shown to have serious consequences in cells or organisms[131]. Bond et al[132] showed that the SNP 309T > G (rs 2279744) located in the promoter region of MDM2 can enhance the transcriptional levels of this protein and subsequent perturbation of p53 functions in the cell. This T > G mutation is thought to generate a binding site on the MDM2 promoter for Sp1 transcription factor[133] and thus enhances the levels of MDM2 protein in the cell.
The positive association between this SNP 309T > G (rs 2279744) in the MDM2 gene and HCC was shown by numerous ethnic-based studies[134-136] and meta-analyses[137,138]. This epidemiological finding together with functional assays of MDM2 levels point to the relevance of MDM2 SNP 309T > G polymorphism as an important player in susceptibility to HCC development.
T cell immunoglobulin mucin-3 (TIM3) negatively regulates the autoimmune and allergic responses and has been linked to T cell dysfunction associated with HBV-related HCC[139]. The 280 aa mature TIM3 is selectively expressed on CD4+ Th1 and CD8+ Tc1 cells, but not on CD4+ Th2 cells[140]. It interacts with its ligand galectin-9 and drives death Th1 T cells[141,142]. Blocking TIM3-mediated signaling restores dysfunctional CD4 and CD8+ T cell-specific adaptive immune responses[143]. TIM3 is upregulated on CD4 and CD8+ T cells in chronic HBV infected individuals[144].
Numerous potential SNPs (-1541C/T, -1516G/T, -882C/T, -574G/T and +4259T/G) in TIM3 have been tested for their association with chronic HBV and HCC[145]. TIM3-1516 G/T (rs10053538) polymorphism has been shown to predispose individuals to cirrhosis and/or HCC[146,147]. One study reported that TIM3 SNPs do not have a functional effect[148], whereas others have reported a significant effect of these TIM3 polymorphic variants[149]. Further studies are needed to determine the functional relevance of this polymorphism.
Xeroderma pigmentosum complementation group C (XPC) protein along with seven other core members (ERCC1, XPA, XPB, XPC, XPD, XPE, XPF and XPG) constitutes the nucleotide excision repair pathway (NER). This pathway is required for the repair of DNA damage including pyrimidine dimers, photo products, chemical adducts and cross-links[150,151]. XPC requires an association with HR23B in order to recognize damaged DNA[152]. The protein HR23B is a human homolog of Saccharomyces cerevisiae RAD23 and binding of XPC-HR23B to a DNA lesion unwinds the helix[153]. The XPA protein can then bind and the whole repair machinery of the NER can be recruited onto the damaged base.
Many studies have investigated the association between XPC sequence variants and cancer risk[154-158]. The three most commonly studied SNPs in the literature are: PAT-/+[159], Lys939Gln (A33512C, rs2228001)[155] and Ala499Val (C21151T, rs2228000)[160]. The poly (AT) insertion/deletion polymorphism (PAT) is located on intron 9 and has been shown to be linked to head and neck cancer risk[161] and to lung cancer[162], but no studies have found an association with HCC risk. The XPC codon Lys939Gln alleles, on the other hand, significantly increased HCC risk[163,164]. The Ala499Val variant homozygous genotype is a risk factor for bladder cancer[158], but has not been studied for HCC.
IL-16 is a pro-inflammatory cytokine and was initially called lymphocyte chemoattractant factor[165]. It can activate a diverse set of immune cells such as CD4+ T cells, monocytes, macrophages, eosinophils and dendritic cells[166-169]. In addition to inducing activation and chemotaxis of immune cells, IL-16 can upregulate the IL-2 receptor[170] and HLA-DR4 expression[171]. Upon CD4 receptor binding, IL-16 signaling increases intracellular calcium and inositol triphosphate, and translocation of protein kinase C from the cytosol to the plasma membrane[172,173]. Moreover, IL-16 can stimulate the production of further pro-inflammatory mediators including LI-1β, IL-6, IL-15 and TNFα, e.g., by monocytes[174] thereby initiating and/or sustaining the inflammatory response.
Genetic polymorphisms in IL-16 have recently been reported and shown to affect susceptibility to a range of cancers including colorectal, gastric and prostate cancer and nasopharyngeal carcinoma[175-178]. Data regarding HCC and IL-16 polymorphisms are scarce in the literature and only two studies were found to have assessed three SNPs (rs11556218T > G, rs4778889T > C, and rs4072111C > T)[179]. In the study by Li et al[180], no association with HCC was found for all three SNPs (rs11556218T/G P = 0.511, rs4072111C/T P = 0.308 and rs4778889T/C P = 0.070). The other study by Thomas et al[178] did not include HCC patients. However, this study did include chronic hepatitis B patients who showed a positive association between rs11556218T > G, a negative association between rs4778889T > C and a positive association between rs4072111C > T polymorphisms and patient susceptibility to chronic hepatitis B infection[179].
Numerous genome-wide association studies (GWAS) have been carried out with chronic HBV and HCC patients to identify novel susceptible loci contributing to disease[6,181-186]. Of these, strong associations were found at 1p36.22, 11q22.3, 6p21 (rs1419881, rs3997872, rs7453920 and rs7768538), 8p12 (rs2275959 and rs37821974) and 22q11.21. The genes implicated in these studies include HLA-DQB2, HLA-DQA1, transcription factor 19 (TCF19), HLA-C, ubiquitin-conjugating enzyme E2 (UBE2L3), LTL, ferredoxin 1 (FDX1), MICA, UBE4B and PG.
HLA-DQ is an MHC class II cell surface receptor found on antigen presenting cells, whereas HLA-C is an MHC class I receptor expressed by all cells. TCF19, as the name suggests, is an important transcription factor during cell cycle G1/S transition[187]. UBE2L3 is a typical E2 ligase that accepts ubiquitin from the E1 complex and transfers it to targeted proteins[188]. Leukocyte telomere length (LTL) has been associated with the risk of developing many malignancies[189] and LTL-related SNPs are potential targets for such GWAS studies. FDX1 is a gene that codes for a small iron-sulfur protein that transfers electrons from NADPH through ferredoxin reductase to mitochondrial cytochrome P450[190]. In addition, it is involved in steroid, vitamin D, and bile acid metabolism[191].
These SNPs found to be associated with the above-mentioned genes still require validation in association studies in order to be considered good prognostic candidates for HCC.
Tumor growth factor beta (TGFβ) is a tumor suppressor gene located on chromosome 19q13.1-13.39. The protein TGFβ is involved in pleiotropic biological processes such as cell growth[192], differentiation[193], extracellular matrix synthesis[194], hematopoiesis[195], angiogenesis[196], and cellular apoptosis[197]. TGFβ1 is one of TGFβ isoforms and is upregulated in HCC tissues correlating with the carcinogenesis and prognosis of HCC[198,199]. TGFβ1 also suppresses HBV replication by reducing hepatocyte nuclear factor-4-alpha[200]. Thus, the relevance of this cytokine and its single nucleotide polymorphism in HBV-associated HCC is of paramount importance.
Seven TGFβ1 polymorphisms have been described in the literature, of which three lie in the upstream region of the gene at positions -988C > A, -800G > A, and -509C > T, one insertion in a nontranslated region at position +72C, two in exon 1 (Leu10Pro and Arg25Pro); and 1 in exon 5 (Thr263Ile)[201]. Numerous studies have investigated the association between these SNPs and HCC[202-205]. There are contrasting reports with some studies reporting a positive association between -509C > T (rs1800469) and HCC risk[206], whereas another study reported a weak or no association[204]. In addition, the Arg25Pro change at +915G/C (rs1800471) was not correlated with HCC risk[207]. The mutation in codon 10 (Leu > Pro) was very strongly correlated with HCC according to one study[208]. There is still limited information regarding other polymorphisms of TGFβ1 and further studies are required to draw firm conclusions on their association with HCC. Table 1 lists the polymorphic genes and their contribution to HCC.
| Polymorphism | Genotype | Significance | Ref. |
| COX-2 | -1195G > A | P < 0.00[26] | He et al[31] |
| -765G > C | P < 0.05[31] and 0.41[26] | Chen et al[26] | |
| +8473T > C | P = 0.83[26] | ||
| IL-1α, β | 511C > T | P = 0.02[41] | Wang et al[41] |
| -31C > T | P = 0.02[41] | ||
| CDH1 | -347G > A | P = 0.171[209] and < 0.05[60] | Li et al[209], Chien et al[60] |
| PPARγ | L162V | P = 0.071[66] | Koytak et al[66] |
| TNFAIP3 | F127C | P = 0.15[75] | Zhang et al[75] |
| TNFα | -1031T/C | P = 0.85[86] | Wei et al[86] |
| -863C/A | P = 0.006[86] | ||
| -857C/T | P = 0.09[86] | ||
| -308G/A | P = 0.046[86] | ||
| -238G/A | P = 0.003[86] | ||
| GST | GSTM1 + GSTT1 | P = 0.001[210] | Liu et al[210] |
| EGF | +61A > G | P < 0.001[117] | Jiang et al[117] |
| MDM2 | 309G > T | P = 0.001[133] | Ezzikouri et al[133] |
| TIM3 | -1516G > T | P = 0.001[146] | Li et al[146] |
| XPC | K939Q | P = 0.001[163] | Long et al[163] |
| 1p36.22, 11q22.3, 6p21, 8p12 22q11.21 | Include genes HLA-DQB2, HLA-DQA1, TCF19, HLA-C, UBE2L3, LTL, FDX1, MICA, UBE4B and PG | P = 1.7 × 10-18 | Al-Qahtani et al[181] |
| P = 4.3 × 10-8 | |||
| P = 0.0266 | |||
| P = 0.0067 | |||
| P = 1.71 × 10-12 | |||
| TGFβ1 | -509C > T | P = 0.01[206] and 0.318[207] | Qi et al[206] |
| R25P | P = 0.472[207] | Hosseini Razavi et al[207] | |
| L10P | P < 0.02[208] | Kim et al[208] |
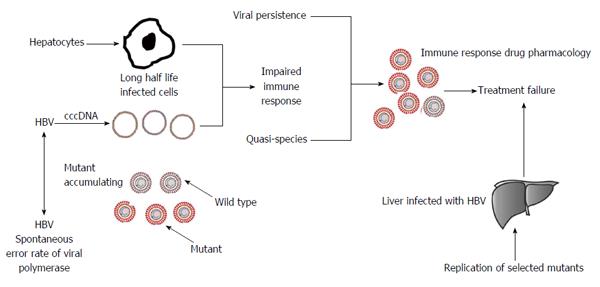
In this article, we discuss the association between the HBV genotype and its mutations in the development of liver cancer and the possibility that individuals with inherited genetic mutations have a hereditary predisposition for HBV-related HCC. Such individuals can inherit a germ-line mutation in one allele of the gene; somatic mutation of the second allele facilitates tumor progression. Although the inherited germ-line mutation may not be adequate to affect tumor development, it is likely that HBV proteins also induce many alterations in the genome. Analysis of the whole transcriptome in these individuals with genetic predisposition would be a useful indicator. It is now well understood that host genetic differences significantly influence susceptibility and resistance to HBV infection and the development of liver cancer, thus it is important to identify these genotype-phenotype associations for better treatment of the disease (Figure 1). Genome-wide sequencing studies have identified numerous germline mutations associated with liver cancer predisposition and large numbers of somatic alterations. It is difficult to assess the difference between background and HBV-related mutations as HBV infection plays an important role in the development of host genetic mutations, due to impairment in the DNA repair process. To elucidate the role of HBV-related genetic variations, researchers have used traditional biological methods to identify genetic mutations. More recently, advanced techniques such as next generation sequencing technology have been used to identify key mutations involved in the development of HCC. Important HCC-associated mutations have been found in key regulatory genes including COX-2, IL-1α and 1β, E-cadherin (CDH1), PPARγ, TNFαIP3, CTLA-4, TNFα, IL-10, GSTM1/GSTT1 Deletion Oxidative stress, EGF, MDM2, TIM3), XPC, IL-16, TGFβ, 1p36.22, 11q22.3, 6p21, 8p12 and 22q11.21 candidate SNPs in GWAS. The association between each locus and the outcome of liver disease is discussed in detail in this article.
Based on these findings, we predict that advanced sequence analysis of host genome will provide us with a better understanding of the viral and host genetic factors involved in the development of HCC. Further studies are needed to evaluate and understand the role of host-HBV interactions in HBV-related HCC to generate effective diagnostic and therapeutic treatments.
P- Reviewer: Chung YH, Vaughan G S- Editor: Wang JL L- Editor: Webster JR E- Editor: Liu SQ
| 1. | El-Serag HB. Epidemiology of viral hepatitis and hepatocellular carcinoma. Gastroenterology. 2012;142:1264-1273.e1. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 2183] [Cited by in F6Publishing: 2337] [Article Influence: 194.8] [Reference Citation Analysis (0)] |
| 2. | Lai CL, Ratziu V, Yuen MF, Poynard T. Viral hepatitis B. Lancet. 2003;362:2089-2094. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 595] [Cited by in F6Publishing: 582] [Article Influence: 27.7] [Reference Citation Analysis (0)] |
| 3. | Yim HJ, Lok AS. Natural history of chronic hepatitis B virus infection: what we knew in 1981 and what we know in 2005. Hepatology. 2006;43:S173-S181. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 364] [Cited by in F6Publishing: 391] [Article Influence: 21.7] [Reference Citation Analysis (1)] |
| 4. | Sherman M. Hepatocellular carcinoma: epidemiology, surveillance, and diagnosis. Semin Liver Dis. 2010;30:3-16. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 284] [Cited by in F6Publishing: 287] [Article Influence: 20.5] [Reference Citation Analysis (0)] |
| 5. | Paterlini-Bréchot P, Saigo K, Murakami Y, Chami M, Gozuacik D, Mugnier C, Lagorce D, Bréchot C. Hepatitis B virus-related insertional mutagenesis occurs frequently in human liver cancers and recurrently targets human telomerase gene. Oncogene. 2003;22:3911-3916. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 239] [Cited by in F6Publishing: 232] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 6. | Kamatani Y, Wattanapokayakit S, Ochi H, Kawaguchi T, Takahashi A, Hosono N, Kubo M, Tsunoda T, Kamatani N, Kumada H. A genome-wide association study identifies variants in the HLA-DP locus associated with chronic hepatitis B in Asians. Nat Genet. 2009;41:591-595. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 396] [Cited by in F6Publishing: 403] [Article Influence: 26.9] [Reference Citation Analysis (0)] |
| 7. | Liaw YF. Hepatitis flares and hepatitis B e antigen seroconversion: implication in anti-hepatitis B virus therapy. J Gastroenterol Hepatol. 2003;18:246-252. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 121] [Cited by in F6Publishing: 127] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 8. | Sokal EM, Paganelli M, Wirth S, Socha P, Vajro P, Lacaille F, Kelly D, Mieli-Vergani G. Management of chronic hepatitis B in childhood: ESPGHAN clinical practice guidelines: consensus of an expert panel on behalf of the European Society of Pediatric Gastroenterology, Hepatology and Nutrition. J Hepatol. 2013;59:814-829. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 151] [Cited by in F6Publishing: 126] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 9. | Cheng HR, Liu CJ, Tseng TC, Su TH, Yang HI, Chen CJ, Kao JH. Host genetic factors affecting spontaneous HBsAg seroclearance in chronic hepatitis B patients. PLoS One. 2013;8:e53008. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 38] [Cited by in F6Publishing: 42] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 10. | Cheong JY, Cho SW, Hwang IL, Yoon SK, Lee JH, Park CS, Lee JE, Hahm KB, Kim JH. Association between chronic hepatitis B virus infection and interleukin-10, tumor necrosis factor-alpha gene promoter polymorphisms. J Gastroenterol Hepatol. 2006;21:1163-1169. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 96] [Cited by in F6Publishing: 97] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 11. | Miyazoe S, Hamasaki K, Nakata K, Kajiya Y, Kitajima K, Nakao K, Daikoku M, Yatsuhashi H, Koga M, Yano M. Influence of interleukin-10 gene promoter polymorphisms on disease progression in patients chronically infected with hepatitis B virus. Am J Gastroenterol. 2002;97:2086-2092. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 111] [Cited by in F6Publishing: 126] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 12. | Du T, Guo XH, Zhu XL, Li JH, Lu LP, Gao JR, Gou CY, Li Z, Liu Y, Li H. Association of TNF-alpha promoter polymorphisms with the outcomes of hepatitis B virus infection in Chinese Han population. J Viral Hepat. 2006;13:618-624. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 49] [Cited by in F6Publishing: 45] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 13. | Wu JF, Wu TC, Chen CH, Ni YH, Chen HL, Hsu HY, Chang MH. Serum levels of interleukin-10 and interleukin-12 predict early, spontaneous hepatitis B virus e antigen seroconversion. Gastroenterology. 2010;138:165-172.e1-3. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 91] [Cited by in F6Publishing: 94] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 14. | Wu JF, Ni YH, Lin YT, Lee TJ, Hsu SH, Chen HL, Tsuei DJ, Hsu HY, Chang MH. Human interleukin-10 genotypes are associated with different precore/core gene mutation patterns in children with chronic hepatitis B virus infection. J Pediatr. 2011;158:808-813. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 20] [Cited by in F6Publishing: 23] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 15. | Xia Q, Zhou L, Liu D, Chen Z, Chen F. Relationship between TNF-<alpha> gene promoter polymorphisms and outcomes of hepatitis B virus infections: a meta-analysis. PLoS One. 2011;6:e19606. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 21] [Cited by in F6Publishing: 24] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 16. | Chatzidaki V, Kouroumalis E, Galanakis E. Hepatitis B virus acquisition and pathogenesis in childhood: host genetic determinants. J Pediatr Gastroenterol Nutr. 2011;52:3-8. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 8] [Cited by in F6Publishing: 8] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 17. | Ortiz-Cuaran S, Villar S, Gouas D, Ferro G, Plymoth A, Khuhaprema T, Kalalak A, Sangrajrang S, Friesen MD, Groopman JD. Association between HBX status, aflatoxin-induced R249S TP53 mutation and risk of hepatocellular carcinoma in a case-control study from Thailand. Cancer Lett. 2013;331:46-51. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 21] [Cited by in F6Publishing: 21] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 18. | Giannitrapani L, Soresi M, Giacalone A, Campagna ME, Marasà M, Cervello M, Marasà S, Montalto G. IL-6 -174G/C polymorphism and IL-6 serum levels in patients with liver cirrhosis and hepatocellular carcinoma. OMICS. 2011;15:183-186. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 35] [Cited by in F6Publishing: 38] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 19. | Gulnaz A, Sayyed AH, Amin F, Khan Au, Aslam MA, Shaikh RS, Ali M. Association of XRCC1, XRCC3, and XPD genetic polymorphism with an increased risk of hepatocellular carcinoma because of the hepatitis B and C virus. Eur J Gastroenterol Hepatol. 2013;25:166-179. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 24] [Cited by in F6Publishing: 23] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 20. | Su C, Lin Y, Niu J, Cai L. Association between polymorphisms in tumor suppressor genes and oncogenes and risk of hepatocellular carcinoma: a case-control study in an HCC epidemic area within the Han Chinese population. Med Oncol. 2014;31:356. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 9] [Cited by in F6Publishing: 10] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 21. | Wu H, Wu X, Wan G, Zhang S. Associations between Cox-2 rs20417 and rs5275 polymorphisms and the risk of hepatocellular carcinoma: a meta analysis. Int J Clin Exp Pathol. 2014;7:6898-6905. [PubMed] [Cited in This Article: ] |
| 22. | Miyashita M, Ito T, Sakaki M, Kajiwara A, Nozawa H, Hiroishi K, Kobayashi M, Kumada H, Imawari M. Genetic polymorphism in cyclooxygenase-2 promoter affects hepatic inflammation and fibrosis in patients with chronic hepatitis C. J Viral Hepat. 2012;19:608-614. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 19] [Cited by in F6Publishing: 21] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 23. | Rouzer CA, Marnett LJ. Endocannabinoid oxygenation by cyclooxygenases, lipoxygenases, and cytochromes P450: cross-talk between the eicosanoid and endocannabinoid signaling pathways. Chem Rev. 2011;111:5899-5921. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 240] [Cited by in F6Publishing: 223] [Article Influence: 17.2] [Reference Citation Analysis (0)] |
| 24. | Pazhang Y, Ahmadian S, Javadifar N, Shafiezadeh M. COX-2 and survivin reduction may play a role in berberine-induced apoptosis in human ductal breast epithelial tumor cell line. Tumour Biol. 2012;33:207-214. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 9] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 25. | Yin J, Liu B, Li B, Liu Z, Xie X, Lv Z, Gao S, Guang J. The cyclooxygenase-2 inhibitor celecoxib attenuates hepatocellular carcinoma growth and c-Met expression in an orthotopic mouse model. Oncol Res. 2011;19:131-139. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 12] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 26. | Chen Z, Zhu J, Huang C, Lian F, Wu G, Zhao Y. The association between three cyclooxygenase-2 polymorphisms and hepatocellular carcinoma risk: a meta-analysis. PLoS One. 2015;10:e0118251. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3] [Cited by in F6Publishing: 3] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 27. | Aubin F, Courivaud C, Bamoulid J, Loupy A, Deschamps M, Ferrand C, Le Corre D, Tiberghien P, Chalopin JM, Legendre C. Influence of cyclooxygenase-2 (COX-2) gene promoter polymorphism at position -765 on skin cancer after renal transplantation. J Invest Dermatol. 2010;130:2134-2136. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 4] [Cited by in F6Publishing: 4] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 28. | Ben Nasr H, Chahed K, Bouaouina N, Chouchane L. PTGS2 (COX-2) -765 G>C functional promoter polymorphism and its association with risk and lymph node metastasis in nasopharyngeal carcinoma. Mol Biol Rep. 2009;36:193-200. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 20] [Cited by in F6Publishing: 21] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 29. | Sitarz R, Leguit RJ, de Leng WW, Polak M, Morsink FM, Bakker O, Maciejewski R, Offerhaus GJ, Milne AN. The COX-2 promoter polymorphism -765 G>C is associated with early-onset, conventional and stump gastric cancers. Mod Pathol. 2008;21:685-690. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 28] [Cited by in F6Publishing: 35] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 30. | Xu DK, Zhang XM, Zhao P, Cai JC, Zhao D, Tan W, Guo YL, Lin DX. [Association between single nucleotide polymorphisms in the promoter of cyclooxygenase COX-2 gene and hereditary susceptibility to pancreatic cancer]. Zhonghua Yi Xue Za Zhi. 2008;88:1961-1965. [PubMed] [Cited in This Article: ] |
| 31. | He J, Zhang Q, Ren Z, Li Y, Li X, Zhou W, Zhang H, Meng W, Yan J, He W. Cyclooxygenase-2 -765 G/C polymorphisms and susceptibility to hepatitis B-related liver cancer in Han Chinese population. Mol Biol Rep. 2012;39:4163-4168. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12] [Cited by in F6Publishing: 12] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 32. | Akkız H, Bayram S, Bekar A, Akgöllü E, Ülger Y. Functional polymorphisms of cyclooxygenase-2 gene and risk for hepatocellular carcinoma. Mol Cell Biochem. 2011;347:201-208. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 26] [Cited by in F6Publishing: 25] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 33. | Gharib AF, Karam RA, Abd El Rahman TM, Elsawy WH. COX-2 polymorphisms -765G→C and -1195A→G and hepatocellular carcinoma risk. Gene. 2014;543:234-236. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 7] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 34. | Chang WS, Yang MD, Tsai CW, Cheng LH, Jeng LB, Lo WC, Lin CH, Huang CY, Bau DT. Association of cyclooxygenase 2 single-nucleotide polymorphisms and hepatocellular carcinoma in Taiwan. Chin J Physiol. 2012;55:1-7. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 17] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 35. | Langsenlehner U, Yazdani-Biuki B, Eder T, Renner W, Wascher TC, Paulweber B, Weitzer W, Samonigg H, Krippl P. The cyclooxygenase-2 (PTGS2) 8473T>C polymorphism is associated with breast cancer risk. Clin Cancer Res. 2006;12:1392-1394. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 67] [Cited by in F6Publishing: 66] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 36. | Upadhyay R, Jain M, Kumar S, Ghoshal UC, Mittal B. Functional polymorphisms of cyclooxygenase-2 (COX-2) gene and risk for esophageal squmaous cell carcinoma. Mutat Res. 2009;663:52-59. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 40] [Cited by in F6Publishing: 41] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 37. | Pan F, Tian J, Pan Y, Zhang Y. Lack of association of the cyclooxygenase 8473 T>C polymorphism with lung cancer: evidence from 9841 subjects. Asian Pac J Cancer Prev. 2011;12:1941-1945. [PubMed] [Cited in This Article: ] |
| 38. | Tilg H, Diehl AM. Cytokines in alcoholic and nonalcoholic steatohepatitis. N Engl J Med. 2000;343:1467-1476. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 688] [Cited by in F6Publishing: 661] [Article Influence: 27.5] [Reference Citation Analysis (0)] |
| 39. | Roshak AK, Jackson JR, McGough K, Chabot-Fletcher M, Mochan E, Marshall LA. Manipulation of distinct NFkappaB proteins alters interleukin-1beta-induced human rheumatoid synovial fibroblast prostaglandin E2 formation. J Biol Chem. 1996;271:31496-31501. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 139] [Cited by in F6Publishing: 142] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 40. | Tian Z, Shen X, Feng H, Gao B. IL-1 beta attenuates IFN-alpha beta-induced antiviral activity and STAT1 activation in the liver: involvement of proteasome-dependent pathway. J Immunol. 2000;165:3959-3965. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 55] [Cited by in F6Publishing: 56] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 41. | Wang Y, Kato N, Hoshida Y, Yoshida H, Taniguchi H, Goto T, Moriyama M, Otsuka M, Shiina S, Shiratori Y. Interleukin-1beta gene polymorphisms associated with hepatocellular carcinoma in hepatitis C virus infection. Hepatology. 2003;37:65-71. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 134] [Cited by in F6Publishing: 143] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 42. | El-Omar EM, Carrington M, Chow WH, McColl KE, Bream JH, Young HA, Herrera J, Lissowska J, Yuan CC, Rothman N. Interleukin-1 polymorphisms associated with increased risk of gastric cancer. Nature. 2000;404:398-402. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1690] [Cited by in F6Publishing: 1614] [Article Influence: 67.3] [Reference Citation Analysis (0)] |
| 43. | Roy N, Mukhopadhyay I, Das K, Pandit P, Majumder PP, Santra A, Datta S, Banerjee S, Chowdhury A. Genetic variants of TNFα, IL10, IL1β, CTLA4 and TGFβ1 modulate the indices of alcohol-induced liver injury in East Indian population. Gene. 2012;509:178-188. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 17] [Cited by in F6Publishing: 16] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 44. | Takamatsu M, Yamauchi M, Maezawa Y, Saito S, Maeyama S, Uchikoshi T. Genetic polymorphisms of interleukin-1beta in association with the development of alcoholic liver disease in Japanese patients. Am J Gastroenterol. 2000;95:1305-1311. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 46] [Cited by in F6Publishing: 41] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 45. | Endo K, Ueda T, Ueyama J, Ohta T, Terada T. Immunoreactive E-cadherin, alpha-catenin, beta-catenin, and gamma-catenin proteins in hepatocellular carcinoma: relationships with tumor grade, clinicopathologic parameters, and patients’ survival. Hum Pathol. 2000;31:558-565. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 127] [Cited by in F6Publishing: 137] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 46. | Huang GT, Lee HS, Chen CH, Sheu JC, Chiou LL, Chen DS. Correlation of E-cadherin expression and recurrence of hepatocellular carcinoma. Hepatogastroenterology. 1999;46:1923-1927. [PubMed] [Cited in This Article: ] |
| 47. | Conacci-Sorrell M, Zhurinsky J, Ben-Ze’ev A. The cadherin-catenin adhesion system in signaling and cancer. J Clin Invest. 2002;109:987-991. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 409] [Cited by in F6Publishing: 405] [Article Influence: 18.4] [Reference Citation Analysis (0)] |
| 48. | Valizadeh A, Karayiannakis AJ, el-Hariry I, Kmiot W, Pignatelli M. Expression of E-cadherin-associated molecules (alpha-, beta-, and gamma-catenins and p120) in colorectal polyps. Am J Pathol. 1997;150:1977-1984. [PubMed] [Cited in This Article: ] |
| 49. | Shiozaki H, Tahara H, Oka H, Miyata M, Kobayashi K, Tamura S, Iihara K, Doki Y, Hirano S, Takeichi M. Expression of immunoreactive E-cadherin adhesion molecules in human cancers. Am J Pathol. 1991;139:17-23. [PubMed] [Cited in This Article: ] |
| 50. | Pignatelli M, Ansari TW, Gunter P, Liu D, Hirano S, Takeichi M, Klöppel G, Lemoine NR. Loss of membranous E-cadherin expression in pancreatic cancer: correlation with lymph node metastasis, high grade, and advanced stage. J Pathol. 1994;174:243-248. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 158] [Cited by in F6Publishing: 171] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 51. | Bringuier PP, Umbas R, Schaafsma HE, Karthaus HF, Debruyne FM, Schalken JA. Decreased E-cadherin immunoreactivity correlates with poor survival in patients with bladder tumors. Cancer Res. 1993;53:3241-3245. [PubMed] [Cited in This Article: ] |
| 52. | Umbas R, Schalken JA, Aalders TW, Carter BS, Karthaus HF, Schaafsma HE, Debruyne FM, Isaacs WB. Expression of the cellular adhesion molecule E-cadherin is reduced or absent in high-grade prostate cancer. Cancer Res. 1992;52:5104-5109. [PubMed] [Cited in This Article: ] |
| 53. | Lee HH, Uen YH, Tian YF, Sun CS, Sheu MJ, Kuo HT, Koay LB, Lin CY, Tzeng CC, Cheng CJ. Wnt-1 protein as a prognostic biomarker for hepatitis B-related and hepatitis C-related hepatocellular carcinoma after surgery. Cancer Epidemiol Biomarkers Prev. 2009;18:1562-1569. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 27] [Cited by in F6Publishing: 34] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 54. | Carter BS, Ewing CM, Ward WS, Treiger BF, Aalders TW, Schalken JA, Epstein JI, Isaacs WB. Allelic loss of chromosomes 16q and 10q in human prostate cancer. Proc Natl Acad Sci USA. 1990;87:8751-8755. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 301] [Cited by in F6Publishing: 318] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 55. | Cleton-Jansen AM, Moerland EW, Kuipers-Dijkshoorn NJ, Callen DF, Sutherland GR, Hansen B, Devilee P, Cornelisse CJ. At least two different regions are involved in allelic imbalance on chromosome arm 16q in breast cancer. Genes Chromosomes Cancer. 1994;9:101-107. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 96] [Cited by in F6Publishing: 103] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 56. | Ribeiro-Filho LA, Franks J, Sasaki M, Shiina H, Li LC, Nojima D, Arap S, Carroll P, Enokida H, Nakagawa M. CpG hypermethylation of promoter region and inactivation of E-cadherin gene in human bladder cancer. Mol Carcinog. 2002;34:187-198. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 66] [Cited by in F6Publishing: 68] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 57. | Matsumura T, Makino R, Mitamura K. Frequent down-regulation of E-cadherin by genetic and epigenetic changes in the malignant progression of hepatocellular carcinomas. Clin Cancer Res. 2001;7:594-599. [PubMed] [Cited in This Article: ] |
| 58. | Zhang X, Ma X, Zhu QG, Li LC, Chen Z, Ye ZQ. Association between a C/A single nucleotide polymorphism of the E-cadherin gene promoter and transitional cell carcinoma of the bladder. J Urol. 2003;170:1379-1382. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 22] [Cited by in F6Publishing: 23] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 59. | Verhage BA, van Houwelingen K, Ruijter TE, Kiemeney LA, Schalken JA. Single-nucleotide polymorphism in the E-cadherin gene promoter modifies the risk of prostate cancer. Int J Cancer. 2002;100:683-685. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 42] [Cited by in F6Publishing: 47] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 60. | Chien MH, Yeh KT, Li YC, Hsieh YH, Lin CH, Weng MS, Kuo WH, Yang SF. Effects of E-cadherin (CDH1) gene promoter polymorphisms on the risk and clinicopathological development of hepatocellular carcinoma. J Surg Oncol. 2011;104:299-304. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 18] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 61. | Semple RK, Chatterjee VK, O’Rahilly S. PPAR gamma and human metabolic disease. J Clin Invest. 2006;116:581-589. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 602] [Cited by in F6Publishing: 600] [Article Influence: 33.3] [Reference Citation Analysis (0)] |
| 62. | Gouda HN, Sagoo GS, Harding AH, Yates J, Sandhu MS, Higgins JP. The association between the peroxisome proliferator-activated receptor-gamma2 (PPARG2) Pro12Ala gene variant and type 2 diabetes mellitus: a HuGE review and meta-analysis. Am J Epidemiol. 2010;171:645-655. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 150] [Cited by in F6Publishing: 161] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 63. | Tönjes A, Stumvoll M. The role of the Pro12Ala polymorphism in peroxisome proliferator-activated receptor gamma in diabetes risk. Curr Opin Clin Nutr Metab Care. 2007;10:410-414. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 33] [Cited by in F6Publishing: 34] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 64. | Huguenin GV, Rosa G. The Ala allele in the PPAR-gamma2 gene is associated with reduced risk of type 2 diabetes mellitus in Caucasians and improved insulin sensitivity in overweight subjects. Br J Nutr. 2010;104:488-497. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 28] [Cited by in F6Publishing: 30] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 65. | Gonzalez FJ. The peroxisome proliferator-activated receptor alpha (PPARalpha): role in hepatocarcinogenesis. Mol Cell Endocrinol. 2002;193:71-79. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 53] [Cited by in F6Publishing: 54] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 66. | Koytak ES, Mizrak D, Bektaş M, Verdi H, Arslan Ergül A, Idilman R, Cinar K, Yurdaydin C, Ersõz S, Karayalçin K. PPAR-alpha L162V polymorphism in human hepatocellular carcinoma. Turk J Gastroenterol. 2008;19:245-249. [PubMed] [Cited in This Article: ] |
| 67. | Flavell DM, Pineda Torra I, Jamshidi Y, Evans D, Diamond JR, Elkeles RS, Bujac SR, Miller G, Talmud PJ, Staels B. Variation in the PPARalpha gene is associated with altered function in vitro and plasma lipid concentrations in Type II diabetic subjects. Diabetologia. 2000;43:673-680. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 135] [Cited by in F6Publishing: 138] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 68. | Vohl MC, Lepage P, Gaudet D, Brewer CG, Bétard C, Perron P, Houde G, Cellier C, Faith JM, Després JP. Molecular scanning of the human PPARa gene: association of the L162v mutation with hyperapobetalipoproteinemia. J Lipid Res. 2000;41:945-952. [PubMed] [Cited in This Article: ] |
| 69. | Tai ES, Demissie S, Cupples LA, Corella D, Wilson PW, Schaefer EJ, Ordovas JM. Association between the PPARA L162V polymorphism and plasma lipid levels: the Framingham Offspring Study. Arterioscler Thromb Vasc Biol. 2002;22:805-810. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 106] [Cited by in F6Publishing: 109] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 70. | Robitaille J, Brouillette C, Houde A, Lemieux S, Pérusse L, Tchernof A, Gaudet D, Vohl MC. Association between the PPARalpha-L162V polymorphism and components of the metabolic syndrome. J Hum Genet. 2004;49:482-489. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 76] [Cited by in F6Publishing: 80] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 71. | Jäättelä M, Mouritzen H, Elling F, Bastholm L. A20 zinc finger protein inhibits TNF and IL-1 signaling. J Immunol. 1996;156:1166-1173. [PubMed] [Cited in This Article: ] |
| 72. | Lee EG, Boone DL, Chai S, Libby SL, Chien M, Lodolce JP, Ma A. Failure to regulate TNF-induced NF-kappaB and cell death responses in A20-deficient mice. Science. 2000;289:2350-2354. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1133] [Cited by in F6Publishing: 1140] [Article Influence: 47.5] [Reference Citation Analysis (0)] |
| 73. | Boone DL, Turer EE, Lee EG, Ahmad RC, Wheeler MT, Tsui C, Hurley P, Chien M, Chai S, Hitotsumatsu O. The ubiquitin-modifying enzyme A20 is required for termination of Toll-like receptor responses. Nat Immunol. 2004;5:1052-1060. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 854] [Cited by in F6Publishing: 863] [Article Influence: 43.2] [Reference Citation Analysis (0)] |
| 74. | Hitotsumatsu O, Ahmad RC, Tavares R, Wang M, Philpott D, Turer EE, Lee BL, Shiffin N, Advincula R, Malynn BA. The ubiquitin-editing enzyme A20 restricts nucleotide-binding oligomerization domain containing 2-triggered signals. Immunity. 2008;28:381-390. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 252] [Cited by in F6Publishing: 268] [Article Influence: 16.8] [Reference Citation Analysis (0)] |
| 75. | Zhang P, Li N, Zhu Q, Li F, Yang C, Zeng X, Lv Y, Zhou Z, Han Q, Liu Z. Association between TNFAIP3 nonsynonymous single-nucleotide polymorphism rs2230926 and chronic hepatitis B virus infection in a Chinese Han population. Virol J. 2015;12:33. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 6] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 76. | Danilovic DL, Mendes-Correa MC, Lima EU, Zambrini H, K Barros R, Marui S. Correlations of CTLA-4 gene polymorphisms and hepatitis C chronic infection. Liver Int. 2012;32:803-808. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 18] [Cited by in F6Publishing: 19] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 77. | Tomer Y, Davies TF. Searching for the autoimmune thyroid disease susceptibility genes: from gene mapping to gene function. Endocr Rev. 2003;24:694-717. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 291] [Cited by in F6Publishing: 275] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 78. | Kristiansen OP, Larsen ZM, Pociot F. CTLA-4 in autoimmune diseases--a general susceptibility gene to autoimmunity? Genes Immun. 2000;1:170-184. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 305] [Cited by in F6Publishing: 285] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 79. | Chen M, Chang Y, Tang F, Xie QH, Li J, Yang H, He XX, Lin JS. Influence of cytotoxic T lymphocyte-associated antigen 4 polymorphisms on the outcomes of hepatitis B virus infection. Mol Med Rep. 2014;9:645-652. [PubMed] [Cited in This Article: ] |
| 80. | Yee LJ, Perez KA, Tang J, van Leeuwen DJ, Kaslow RA. Association of CTLA4 polymorphisms with sustained response to interferon and ribavirin therapy for chronic hepatitis C virus infection. J Infect Dis. 2003;187:1264-1271. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 48] [Cited by in F6Publishing: 45] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 81. | Schott E, Witt H, Hinrichsen H, Neumann K, Weich V, Bergk A, Halangk J, Müller T, Tinjala S, Puhl G. Gender-dependent association of CTLA4 polymorphisms with resolution of hepatitis C virus infection. J Hepatol. 2007;46:372-380. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 36] [Cited by in F6Publishing: 39] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 82. | Nischalke HD, Vogel M, Mauss S, Baumgarten A, Lutz T, Danta M, Naumann U, Coenen M, Sauerbruch T, Rockstroh JK. The cytotoxic lymphocyte antigen 4 polymorphisms affect response to hepatitis C virus-specific therapy in HIV(+) patients with acute and chronic hepatitis C virus co-infection. AIDS. 2010;24:2001-2007. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 11] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 83. | O’Shea RS, Dasarathy S, McCullough AJ. Alcoholic liver disease. Am J Gastroenterol. 2010;105:14-32; quiz 33. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 169] [Cited by in F6Publishing: 194] [Article Influence: 13.9] [Reference Citation Analysis (0)] |
| 84. | Schuppan D, Afdhal NH. Liver cirrhosis. Lancet. 2008;371:838-851. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1446] [Cited by in F6Publishing: 1428] [Article Influence: 89.3] [Reference Citation Analysis (0)] |
| 85. | Rosen HR, Lentz JJ, Rose SL, Rabkin J, Corless CL, Taylor K, Chou S. Donor polymorphism of tumor necrosis factor gene: relationship with variable severity of hepatitis C recurrence after liver transplantation. Transplantation. 1999;68:1898-1902. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 33] [Cited by in F6Publishing: 33] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 86. | Wei Y, Liu F, Li B, Chen X, Ma Y, Yan L, Wen T, Xu M, Wang W, Yang J. Polymorphisms of tumor necrosis factor-alpha and hepatocellular carcinoma risk: a HuGE systematic review and meta-analysis. Dig Dis Sci. 2011;56:2227-2236. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 36] [Cited by in F6Publishing: 34] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 87. | Wilson AG, Symons JA, McDowell TL, McDevitt HO, Duff GW. Effects of a polymorphism in the human tumor necrosis factor alpha promoter on transcriptional activation. Proc Natl Acad Sci USA. 1997;94:3195-3199. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1608] [Cited by in F6Publishing: 1741] [Article Influence: 64.5] [Reference Citation Analysis (0)] |
| 88. | Machado MV, Martins A, Almeida R, Marques-Vidal P, Gonçalves MS, Camilo ME, Cortez-Pinto H. Does the simultaneous tumor necrosis factor receptor 2, tumor necrosis factor promoter gene polymorphism represent a higher risk for alcoholic liver disease? Eur J Gastroenterol Hepatol. 2009;21:201-205. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 6] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 89. | Cookson S, Constantini PK, Clare M, Underhill JA, Bernal W, Czaja AJ, Donaldson PT. Frequency and nature of cytokine gene polymorphisms in type 1 autoimmune hepatitis. Hepatology. 1999;30:851-856. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 138] [Cited by in F6Publishing: 142] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 90. | Grove J, Daly AK, Bassendine MF, Gilvarry E, Day CP. Interleukin 10 promoter region polymorphisms and susceptibility to advanced alcoholic liver disease. Gut. 2000;46:540-545. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 129] [Cited by in F6Publishing: 133] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 91. | Strange RC, Spiteri MA, Ramachandran S, Fryer AA. Glutathione-S-transferase family of enzymes. Mutat Res. 2001;482:21-26. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 560] [Cited by in F6Publishing: 564] [Article Influence: 24.5] [Reference Citation Analysis (0)] |
| 92. | Mohammadzadeh GS, Nasseri Moghadam S, Rasaee MJ, Zaree AB, Mahmoodzadeh H, Allameh A. Measurement of glutathione S-transferase and its class-pi in plasma and tissue biopsies obtained after laparoscopy and endoscopy from subjects with esophagus and gastric cancer. Clin Biochem. 2003;36:283-288. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 16] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 93. | Parl FF. Glutathione S-transferase genotypes and cancer risk. Cancer Lett. 2005;221:123-129. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 135] [Cited by in F6Publishing: 129] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 94. | McIlwain CC, Townsend DM, Tew KD. Glutathione S-transferase polymorphisms: cancer incidence and therapy. Oncogene. 2006;25:1639-1648. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 381] [Cited by in F6Publishing: 378] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 95. | Hayes JD, Strange RC. Glutathione S-transferase polymorphisms and their biological consequences. Pharmacology. 2000;61:154-166. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 683] [Cited by in F6Publishing: 651] [Article Influence: 27.1] [Reference Citation Analysis (0)] |
| 96. | Sun L, Xi B, Yu L, Gao XC, Shi DJ, Yan YK, Xu DJ, Han Q, Wang C. Association of glutathione S-transferases polymorphisms (GSTM1 and GSTT1) with senile cataract: a meta-analysis. Invest Ophthalmol Vis Sci. 2010;51:6381-6386. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 35] [Cited by in F6Publishing: 37] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 97. | Chen SY, Wang LY, Lunn RM, Tsai WY, Lee PH, Lee CS, Ahsan H, Zhang YJ, Chen CJ, Santella RM. Polycyclic aromatic hydrocarbon-DNA adducts in liver tissues of hepatocellular carcinoma patients and controls. Int J Cancer. 2002;99:14-21. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 82] [Cited by in F6Publishing: 91] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 98. | Zhong S, Tang MW, Yeo W, Liu C, Lo YM, Johnson PJ. Silencing of GSTP1 gene by CpG island DNA hypermethylation in HBV-associated hepatocellular carcinomas. Clin Cancer Res. 2002;8:1087-1092. [PubMed] [Cited in This Article: ] |
| 99. | Yu MW, Yang SY, Pan IJ, Lin CL, Liu CJ, Liaw YF, Lin SM, Chen PJ, Lee SD, Chen CJ. Polymorphisms in XRCC1 and glutathione S-transferase genes and hepatitis B-related hepatocellular carcinoma. J Natl Cancer Inst. 2003;95:1485-1488. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 82] [Cited by in F6Publishing: 88] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 100. | Yu L, Wang CY, Xi B, Sun L, Wang RQ, Yan YK, Zhu LY. GST polymorphisms are associated with hepatocellular carcinoma risk in Chinese population. World J Gastroenterol. 2011;17:3248-3256. [PubMed] [Cited in This Article: ] |
| 101. | Wang B, Huang G, Wang D, Li A, Xu Z, Dong R, Zhang D, Zhou W. Null genotypes of GSTM1 and GSTT1 contribute to hepatocellular carcinoma risk: evidence from an updated meta-analysis. J Hepatol. 2010;53:508-518. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 88] [Cited by in F6Publishing: 101] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 102. | Normanno N, Bianco C, De Luca A, Maiello MR, Salomon DS. Target-based agents against ErbB receptors and their ligands: a novel approach to cancer treatment. Endocr Relat Cancer. 2003;10:1-21. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 229] [Cited by in F6Publishing: 216] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 103. | Abd El-Rehim DM, Pinder SE, Paish CE, Bell JA, Rampaul RS, Blamey RW, Robertson JF, Nicholson RI, Ellis IO. Expression and co-expression of the members of the epidermal growth factor receptor (EGFR) family in invasive breast carcinoma. Br J Cancer. 2004;91:1532-1542. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 185] [Cited by in F6Publishing: 192] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 104. | Böni-Schnetzler M, Pilch PF. Mechanism of epidermal growth factor receptor autophosphorylation and high-affinity binding. Proc Natl Acad Sci USA. 1987;84:7832-7836. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 143] [Cited by in F6Publishing: 175] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 105. | Rotin D, Margolis B, Mohammadi M, Daly RJ, Daum G, Li N, Fischer EH, Burgess WH, Ullrich A, Schlessinger J. SH2 domains prevent tyrosine dephosphorylation of the EGF receptor: identification of Tyr992 as the high-affinity binding site for SH2 domains of phospholipase C gamma. EMBO J. 1992;11:559-567. [PubMed] [Cited in This Article: ] |
| 106. | Lowenstein EJ, Daly RJ, Batzer AG, Li W, Margolis B, Lammers R, Ullrich A, Skolnik EY, Bar-Sagi D, Schlessinger J. The SH2 and SH3 domain-containing protein GRB2 links receptor tyrosine kinases to ras signaling. Cell. 1992;70:431-442. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1184] [Cited by in F6Publishing: 1239] [Article Influence: 38.7] [Reference Citation Analysis (0)] |
| 107. | Zhang Y, Wang L, Zhang M, Jin M, Bai C, Wang X. Potential mechanism of interleukin-8 production from lung cancer cells: an involvement of EGF-EGFR-PI3K-Akt-Erk pathway. J Cell Physiol. 2012;227:35-43. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 70] [Cited by in F6Publishing: 83] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 108. | Aaronson DS, Horvath CM. A road map for those who don’t know JAK-STAT. Science. 2002;296:1653-1655. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 967] [Cited by in F6Publishing: 952] [Article Influence: 43.3] [Reference Citation Analysis (0)] |
| 109. | Kiso S, Kawata S, Tamura S, Inui Y, Yoshida Y, Sawai Y, Umeki S, Ito N, Yamada A, Miyagawa J. Liver regeneration in heparin-binding EGF-like growth factor transgenic mice after partial hepatectomy. Gastroenterology. 2003;124:701-707. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 63] [Cited by in F6Publishing: 66] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 110. | Kung CP, Meckes DG, Raab-Traub N. Epstein-Barr virus LMP1 activates EGFR, STAT3, and ERK through effects on PKCdelta. J Virol. 2011;85:4399-4408. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 103] [Cited by in F6Publishing: 116] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 111. | Miyaki M, Sato C, Sakai K, Konishi M, Tanaka K, Muraoka M, Kikuchi-Yanoshita R, Nadaoka Y, Kanda H, Kitagawa T. Malignant transformation and EGFR activation of immortalized mouse liver epithelial cells caused by HBV enhancer-X from a human hepatocellular carcinoma. Int J Cancer. 2000;85:518-522. [PubMed] [Cited in This Article: ] |
| 112. | Chen YJ, Chien PH, Chen WS, Chien YF, Hsu YY, Wang LY, Chen JY, Lin CW, Huang TC, Yu YL. Hepatitis B Virus-Encoded X Protein Downregulates EGFR Expression via Inducing MicroRNA-7 in Hepatocellular Carcinoma Cells. Evid Based Complement Alternat Med. 2013;2013:682380. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 29] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 113. | Chaleshi V, Haghighi MM, Javadi GR, Fatemi SR, Vahedi M, Zali MR. The effect of 5’untranslated region polymorphism in EGF gene, rs4444903, on colorectal cancer. Gastroenterol Hepatol Bed Bench. 2013;6:129-135. [PubMed] [Cited in This Article: ] |
| 114. | Peng Q, Li S, Qin X, Lao X, Chen Z, Zhang X, Chen J. EGF +61A/G polymorphism contributes to increased gastric cancer risk: evidence from a meta-analysis. Cancer Cell Int. 2014;14:134. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3] [Cited by in F6Publishing: 3] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 115. | Li YL, Tian Z, Zhao L, Zhang CL. Association between the EGF rs4444903 polymorphism and liver cancer susceptibility: a meta-analysis and meta-regression. Genet Mol Res. 2014;13:8066-8079. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3] [Cited by in F6Publishing: 3] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 116. | Hu M, Shi H, Xu Z, Liu W. Association between epidermal growth factor gene rs4444903 polymorphism and risk of glioma. Tumour Biol. 2013;34:1879-1885. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 8] [Cited by in F6Publishing: 9] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 117. | Jiang G, Yu K, Shao L, Yu X, Hu C, Qian P, Xie H, Li J, Zheng J, Zheng S. Association between epidermal growth factor gene +61A/G polymorphism and the risk of hepatocellular carcinoma: a meta-analysis based on 16 studies. BMC Cancer. 2015;15:314. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 18] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 118. | Shahbazi M, Pravica V, Nasreen N, Fakhoury H, Fryer AA, Strange RC, Hutchinson PE, Osborne JE, Lear JT, Smith AG. Association between functional polymorphism in EGF gene and malignant melanoma. Lancet. 2002;359:397-401. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 197] [Cited by in F6Publishing: 212] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 119. | Tanabe KK, Lemoine A, Finkelstein DM, Kawasaki H, Fujii T, Chung RT, Lauwers GY, Kulu Y, Muzikansky A, Kuruppu D. Epidermal growth factor gene functional polymorphism and the risk of hepatocellular carcinoma in patients with cirrhosis. JAMA. 2008;299:53-60. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 148] [Cited by in F6Publishing: 160] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 120. | Yuan JM, Fan Y, Ognjanovic S, Wang R, Van Den Berg D, Govindarajan S, Yu MC. Genetic polymorphisms of epidermal growth factor in relation to risk of hepatocellular carcinoma: two case-control studies. BMC Gastroenterol. 2013;13:32. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 13] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 121. | Suenaga M, Yamada S, Fujii T, Fuchs BC, Okumura N, Kanda M, Kobayashi D, Tanaka C, Nakayama G, Sugimoto H. A functional polymorphism in the epidermal growth factor gene predicts hepatocellular carcinoma risk in Japanese hepatitis C patients. Onco Targets Ther. 2013;6:1805-1812. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 14] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 122. | Abbas E, Shaker O, Abd El Aziz G, Ramadan H, Esmat G. Epidermal growth factor gene polymorphism 61A/G in patients with chronic liver disease for early detection of hepatocellular carcinoma: a pilot study. Eur J Gastroenterol Hepatol. 2012;24:458-463. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1] [Cited by in F6Publishing: 7] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 123. | Zhong JH, You XM, Gong WF, Ma L, Zhang Y, Mo QG, Wu LC, Xiao J, Li LQ. Epidermal growth factor gene polymorphism and risk of hepatocellular carcinoma: a meta-analysis. PLoS One. 2012;7:e32159. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 35] [Cited by in F6Publishing: 40] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 124. | Rivlin N, Brosh R, Oren M, Rotter V. Mutations in the p53 Tumor Suppressor Gene: Important Milestones at the Various Steps of Tumorigenesis. Genes Cancer. 2011;2:466-474. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 536] [Cited by in F6Publishing: 624] [Article Influence: 48.0] [Reference Citation Analysis (0)] |
| 125. | Meek DW. The p53 response to DNA damage. DNA Repair (Amst). 2004;3:1049-1056. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 260] [Cited by in F6Publishing: 241] [Article Influence: 12.7] [Reference Citation Analysis (0)] |
| 126. | Pellegata NS, Antoniono RJ, Redpath JL, Stanbridge EJ. DNA damage and p53-mediated cell cycle arrest: a reevaluation. Proc Natl Acad Sci USA. 1996;93:15209-15214. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 134] [Cited by in F6Publishing: 141] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 127. | Soengas MS, Alarcón RM, Yoshida H, Giaccia AJ, Hakem R, Mak TW, Lowe SW. Apaf-1 and caspase-9 in p53-dependent apoptosis and tumor inhibition. Science. 1999;284:156-159. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 480] [Cited by in F6Publishing: 470] [Article Influence: 18.8] [Reference Citation Analysis (0)] |
| 128. | Moll UM, Petrenko O. The MDM2-p53 interaction. Mol Cancer Res. 2003;1:1001-1008. [PubMed] [Cited in This Article: ] |
| 129. | Rodriguez MS, Desterro JM, Lain S, Lane DP, Hay RT. Multiple C-terminal lysine residues target p53 for ubiquitin-proteasome-mediated degradation. Mol Cell Biol. 2000;20:8458-8467. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 281] [Cited by in F6Publishing: 276] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 130. | Haupt Y, Maya R, Kazaz A, Oren M. Mdm2 promotes the rapid degradation of p53. Nature. 1997;387:296-299. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3222] [Cited by in F6Publishing: 3262] [Article Influence: 120.8] [Reference Citation Analysis (0)] |
| 131. | Bond GL, Hu W, Levine AJ. MDM2 is a central node in the p53 pathway: 12 years and counting. Curr Cancer Drug Targets. 2005;5:3-8. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 233] [Cited by in F6Publishing: 250] [Article Influence: 13.2] [Reference Citation Analysis (0)] |
| 132. | Bond GL, Hu W, Bond EE, Robins H, Lutzker SG, Arva NC, Bargonetti J, Bartel F, Taubert H, Wuerl P. A single nucleotide polymorphism in the MDM2 promoter attenuates the p53 tumor suppressor pathway and accelerates tumor formation in humans. Cell. 2004;119:591-602. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 926] [Cited by in F6Publishing: 954] [Article Influence: 50.2] [Reference Citation Analysis (0)] |
| 133. | Ezzikouri S, El Feydi AE, Afifi R, El Kihal L, Benazzouz M, Hassar M, Marchio A, Pineau P, Benjelloun S. MDM2 SNP309T>G polymorphism and risk of hepatocellular carcinoma: a case-control analysis in a Moroccan population. Cancer Detect Prev. 2009;32:380-385. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 29] [Cited by in F6Publishing: 29] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 134. | Di Vuolo V, Buonaguro L, Izzo F, Losito S, Botti G, Buonaguro FM, Tornesello ML. TP53 and MDM2 gene polymorphisms and risk of hepatocellular carcinoma among Italian patients. Infect Agent Cancer. 2011;6:13. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 19] [Cited by in F6Publishing: 20] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 135. | Dharel N, Kato N, Muroyama R, Moriyama M, Shao RX, Kawabe T, Omata M. MDM2 promoter SNP309 is associated with the risk of hepatocellular carcinoma in patients with chronic hepatitis C. Clin Cancer Res. 2006;12:4867-4871. [PubMed] [Cited in This Article: ] |
| 136. | Yoon YJ, Chang HY, Ahn SH, Kim JK, Park YK, Kang DR, Park JY, Myoung SM, Kim do Y, Chon CY. MDM2 and p53 polymorphisms are associated with the development of hepatocellular carcinoma in patients with chronic hepatitis B virus infection. Carcinogenesis. 2008;29:1192-1196. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 96] [Cited by in F6Publishing: 103] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 137. | Liu GY, Jiang DK, Shen SQ, Yu L. MDM2 SNP309T>G polymorphism with hepatocellular carcinoma risk: a meta-analysis. Arch Med Res. 2011;42:149-155. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 21] [Cited by in F6Publishing: 21] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 138. | Peng Q, Lao X, Chen Z, Lai H, Deng Y, Wang J, Mo C, Sui J, Wu J, Zhai L. TP53 and MDM2 gene polymorphisms, gene-gene interaction, and hepatocellular carcinoma risk: evidence from an updated meta-analysis. PLoS One. 2013;8:e82773. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 21] [Cited by in F6Publishing: 26] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 139. | Li H, Wu K, Tao K, Chen L, Zheng Q, Lu X, Liu J, Shi L, Liu C, Wang G. Tim-3/galectin-9 signaling pathway mediates T-cell dysfunction and predicts poor prognosis in patients with hepatitis B virus-associated hepatocellular carcinoma. Hepatology. 2012;56:1342-1351. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 322] [Cited by in F6Publishing: 357] [Article Influence: 29.8] [Reference Citation Analysis (0)] |
| 140. | Monney L, Sabatos CA, Gaglia JL, Ryu A, Waldner H, Chernova T, Manning S, Greenfield EA, Coyle AJ, Sobel RA. Th1-specific cell surface protein Tim-3 regulates macrophage activation and severity of an autoimmune disease. Nature. 2002;415:536-541. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1124] [Cited by in F6Publishing: 1194] [Article Influence: 54.3] [Reference Citation Analysis (0)] |
| 141. | Zhu C, Anderson AC, Schubart A, Xiong H, Imitola J, Khoury SJ, Zheng XX, Strom TB, Kuchroo VK. The Tim-3 ligand galectin-9 negatively regulates T helper type 1 immunity. Nat Immunol. 2005;6:1245-1252. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1334] [Cited by in F6Publishing: 1459] [Article Influence: 76.8] [Reference Citation Analysis (0)] |
| 142. | Sánchez-Fueyo A, Tian J, Picarella D, Domenig C, Zheng XX, Sabatos CA, Manlongat N, Bender O, Kamradt T, Kuchroo VK. Tim-3 inhibits T helper type 1-mediated auto- and alloimmune responses and promotes immunological tolerance. Nat Immunol. 2003;4:1093-1101. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 519] [Cited by in F6Publishing: 552] [Article Influence: 26.3] [Reference Citation Analysis (0)] |
| 143. | Golden-Mason L, Palmer BE, Kassam N, Townshend-Bulson L, Livingston S, McMahon BJ, Castelblanco N, Kuchroo V, Gretch DR, Rosen HR. Negative immune regulator Tim-3 is overexpressed on T cells in hepatitis C virus infection and its blockade rescues dysfunctional CD4+ and CD8+ T cells. J Virol. 2009;83:9122-9130. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 338] [Cited by in F6Publishing: 368] [Article Influence: 24.5] [Reference Citation Analysis (0)] |
| 144. | Wu W, Shi Y, Li J, Chen F, Chen Z, Zheng M. Tim-3 expression on peripheral T cell subsets correlates with disease progression in hepatitis B infection. Virol J. 2011;8:113. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 58] [Cited by in F6Publishing: 63] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 145. | Li Z, Liu Z, Zhang G, Han Q, Li N, Zhu Q, Lv Y, Chen J, Xing F, Wang Y. TIM3 gene polymorphisms in patients with chronic hepatitis B virus infection: impact on disease susceptibility and hepatocellular carcinoma traits. Tissue Antigens. 2012;80:151-157. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 20] [Cited by in F6Publishing: 23] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 146. | Li Z, Li N, Zhu Q, Zhang G, Han Q, Zhang P, Xun M, Wang Y, Zeng X, Yang C. Genetic variations of PD1 and TIM3 are differentially and interactively associated with the development of cirrhosis and HCC in patients with chronic HBV infection. Infect Genet Evol. 2013;14:240-246. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 38] [Cited by in F6Publishing: 41] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 147. | Wang L, Zhao C, Peng Q, Shi J, Gu G. Expression levels of CD28, CTLA-4, PD-1 and Tim-3 as novel indicators of T-cell immune function in patients with chronic hepatitis B virus infection. Biomed Rep. 2014;2:270-274. [PubMed] [Cited in This Article: ] |
| 148. | Zhang J, Daley D, Akhabir L, Stefanowicz D, Chan-Yeung M, Becker AB, Laprise C, Paré PD, Sandford AJ. Lack of association of TIM3 polymorphisms and allergic phenotypes. BMC Med Genet. 2009;10:62. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 11] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 149. | DeKruyff RH, Bu X, Ballesteros A, Santiago C, Chim YL, Lee HH, Karisola P, Pichavant M, Kaplan GG, Umetsu DT. T cell/transmembrane, Ig, and mucin-3 allelic variants differentially recognize phosphatidylserine and mediate phagocytosis of apoptotic cells. J Immunol. 2010;184:1918-1930. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 208] [Cited by in F6Publishing: 225] [Article Influence: 16.1] [Reference Citation Analysis (0)] |
| 150. | Sugasawa K, Ng JM, Masutani C, Iwai S, van der Spek PJ, Eker AP, Hanaoka F, Bootsma D, Hoeijmakers JH. Xeroderma pigmentosum group C protein complex is the initiator of global genome nucleotide excision repair. Mol Cell. 1998;2:223-232. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 628] [Cited by in F6Publishing: 632] [Article Influence: 24.3] [Reference Citation Analysis (0)] |
| 151. | de Laat WL, Jaspers NG, Hoeijmakers JH. Molecular mechanism of nucleotide excision repair. Genes Dev. 1999;13:768-785. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 811] [Cited by in F6Publishing: 798] [Article Influence: 31.9] [Reference Citation Analysis (0)] |
| 152. | Thoma BS, Vasquez KM. Critical DNA damage recognition functions of XPC-hHR23B and XPA-RPA in nucleotide excision repair. Mol Carcinog. 2003;38:1-13. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 95] [Cited by in F6Publishing: 100] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 153. | Rouillon C, White MF. The XBP-Bax1 helicase-nuclease complex unwinds and cleaves DNA: implications for eukaryal and archaeal nucleotide excision repair. J Biol Chem. 2010;285:11013-11022. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 27] [Cited by in F6Publishing: 27] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 154. | Hollander MC, Philburn RT, Patterson AD, Velasco-Miguel S, Friedberg EC, Linnoila RI, Fornace AJ. Deletion of XPC leads to lung tumors in mice and is associated with early events in human lung carcinogenesis. Proc Natl Acad Sci USA. 2005;102:13200-13205. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 105] [Cited by in F6Publishing: 105] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 155. | Hu Z, Wang Y, Wang X, Liang G, Miao X, Xu Y, Tan W, Wei Q, Lin D, Shen H. DNA repair gene XPC genotypes/haplotypes and risk of lung cancer in a Chinese population. Int J Cancer. 2005;115:478-483. [PubMed] [Cited in This Article: ] |
| 156. | Vogel U, Overvad K, Wallin H, Tjønneland A, Nexø BA, Raaschou-Nielsen O. Combinations of polymorphisms in XPD, XPC and XPA in relation to risk of lung cancer. Cancer Lett. 2005;222:67-74. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 72] [Cited by in F6Publishing: 65] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 157. | Zhu Y, Lai M, Yang H, Lin J, Huang M, Grossman HB, Dinney CP, Wu X. Genotypes, haplotypes and diplotypes of XPC and risk of bladder cancer. Carcinogenesis. 2007;28:698-703. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 33] [Cited by in F6Publishing: 37] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 158. | Qiu L, Wang Z, Shi X, Wang Z. Associations between XPC polymorphisms and risk of cancers: A meta-analysis. Eur J Cancer. 2008;44:2241-2253. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 60] [Cited by in F6Publishing: 64] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 159. | Khan SG, Metter EJ, Tarone RE, Bohr VA, Grossman L, Hedayati M, Bale SJ, Emmert S, Kraemer KH. A new xeroderma pigmentosum group C poly(AT) insertion/deletion polymorphism. Carcinogenesis. 2000;21:1821-1825. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 104] [Cited by in F6Publishing: 110] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 160. | Khan SG, Muniz-Medina V, Shahlavi T, Baker CC, Inui H, Ueda T, Emmert S, Schneider TD, Kraemer KH. The human XPC DNA repair gene: arrangement, splice site information content and influence of a single nucleotide polymorphism in a splice acceptor site on alternative splicing and function. Nucleic Acids Res. 2002;30:3624-3631. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 130] [Cited by in F6Publishing: 131] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 161. | Zhang D, Chen C, Fu X, Gu S, Mao Y, Xie Y, Huang Y, Li Y. A meta-analysis of DNA repair gene XPC polymorphisms and cancer risk. J Hum Genet. 2008;53:18-33. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 41] [Cited by in F6Publishing: 36] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 162. | Jin B, Dong Y, Zhang X, Wang H, Han B. Association of XPC polymorphisms and lung cancer risk: a meta-analysis. PLoS One. 2014;9:e93937. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 19] [Cited by in F6Publishing: 19] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 163. | Long XD, Ma Y, Zhou YF, Ma AM, Fu GH. Polymorphism in xeroderma pigmentosum complementation group C codon 939 and aflatoxin B1-related hepatocellular carcinoma in the Guangxi population. Hepatology. 2010;52:1301-1309. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 58] [Cited by in F6Publishing: 59] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 164. | Yao JG, Huang XY, Long XD. Interaction of DNA repair gene polymorphisms and aflatoxin B1 in the risk of hepatocellular carcinoma. Int J Clin Exp Pathol. 2014;7:6231-6244. [PubMed] [Cited in This Article: ] |
| 165. | Cruikshank W, Center DM. Modulation of lymphocyte migration by human lymphokines. II. Purification of a lymphotactic factor (LCF). J Immunol. 1982;128:2569-2574. [PubMed] [Cited in This Article: ] |
| 166. | Ferland C, Flamand N, Davoine F, Chakir J, Laviolette M. IL-16 activates plasminogen-plasmin system and promotes human eosinophil migration into extracellular matrix via CCR3-chemokine-mediated signaling and by modulating CD4 eosinophil expression. J Immunol. 2004;173:4417-4424. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 24] [Cited by in F6Publishing: 25] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 167. | Bandeira-Melo C, Sugiyama K, Woods LJ, Phoofolo M, Center DM, Cruikshank WW, Weller PF. IL-16 promotes leukotriene C(4) and IL-4 release from human eosinophils via CD4- and autocrine CCR3-chemokine-mediated signaling. J Immunol. 2002;168:4756-4763. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 52] [Cited by in F6Publishing: 55] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 168. | Liu Y, Cruikshank WW, O’Loughlin T, O’Reilly P, Center DM, Kornfeld H. Identification of a CD4 domain required for interleukin-16 binding and lymphocyte activation. J Biol Chem. 1999;274:23387-23395. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 45] [Cited by in F6Publishing: 46] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 169. | Krautwald S. IL-16 activates the SAPK signaling pathway in CD4+ macrophages. J Immunol. 1998;160:5874-5879. [PubMed] [Cited in This Article: ] |
| 170. | Cruikshank WW, Greenstein JL, Theodore AC, Center DM. Lymphocyte chemoattractant factor induces CD4-dependent intracytoplasmic signaling in lymphocytes. J Immunol. 1991;146:2928-2934. [PubMed] [Cited in This Article: ] |
| 171. | Cruikshank WW, Berman JS, Theodore AC, Bernardo J, Center DM. Lymphokine activation of T4+ T lymphocytes and monocytes. J Immunol. 1987;138:3817-3823. [PubMed] [Cited in This Article: ] |
| 172. | Parada NA, Cruikshank WW, Danis HL, Ryan TC, Center DM. IL-16- and other CD4 ligand-induced migration is dependent upon protein kinase C. Cell Immunol. 1996;168:100-106. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 40] [Cited by in F6Publishing: 43] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 173. | Ryan TC, Cruikshank WW, Kornfeld H, Collins TL, Center DM. The CD4-associated tyrosine kinase p56lck is required for lymphocyte chemoattractant factor-induced T lymphocyte migration. J Biol Chem. 1995;270:17081-17086. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 62] [Cited by in F6Publishing: 63] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 174. | Mathy NL, Scheuer W, Lanzendörfer M, Honold K, Ambrosius D, Norley S, Kurth R. Interleukin-16 stimulates the expression and production of pro-inflammatory cytokines by human monocytes. Immunology. 2000;100:63-69. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 155] [Cited by in F6Publishing: 168] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 175. | Gao LB, Liang WB, Xue H, Rao L, Pan XM, Lv ML, Bai P, Fang WL, Liu J, Liao M. Genetic polymorphism of interleukin-16 and risk of nasopharyngeal carcinoma. Clin Chim Acta. 2009;409:132-135. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 23] [Cited by in F6Publishing: 20] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 176. | Gao LB, Rao L, Wang YY, Liang WB, Li C, Xue H, Zhou B, Sun H, Li Y, Lv ML. The association of interleukin-16 polymorphisms with IL-16 serum levels and risk of colorectal and gastric cancer. Carcinogenesis. 2009;30:295-299. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 64] [Cited by in F6Publishing: 73] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 177. | Qin X, Peng Q, Lao X, Chen Z, Lu Y, Lao X, Mo C, Sui J, Wu J, Zhai L. The association of interleukin-16 gene polymorphisms with IL-16 serum levels and risk of nasopharyngeal carcinoma in a Chinese population. Tumour Biol. 2014;35:1917-1924. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 20] [Cited by in F6Publishing: 20] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 178. | Thomas G, Jacobs KB, Yeager M, Kraft P, Wacholder S, Orr N, Yu K, Chatterjee N, Welch R, Hutchinson A. Multiple loci identified in a genome-wide association study of prostate cancer. Nat Genet. 2008;40:310-315. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 699] [Cited by in F6Publishing: 771] [Article Influence: 48.2] [Reference Citation Analysis (0)] |
| 179. | Romani S, Hosseini SM, Mohebbi SR, Kazemian S, Derakhshani S, Khanyaghma M, Azimzadeh P, Sharifian A, Zali MR. Interleukin-16 gene polymorphisms are considerable host genetic factors for patients’ susceptibility to chronic hepatitis B infection. Hepat Res Treat. 2014;2014:790753. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 10] [Cited by in F6Publishing: 13] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 180. | Li S, Deng Y, Chen ZP, Huang S, Liao XC, Lin LW, Li H, Peng T, Qin X, Zhao JM. Genetic polymorphism of interleukin-16 influences susceptibility to HBV-related hepatocellular carcinoma in a Chinese population. Infect Genet Evol. 2011;11:2083-2088. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 34] [Cited by in F6Publishing: 39] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 181. | Al-Qahtani A, Khalak HG, Alkuraya FS, Al-hamoudi W, Alswat K, Al Balwi MA, Al Abdulkareem I, Sanai FM, Abdo AA. Genome-wide association study of chronic hepatitis B virus infection reveals a novel candidate risk allele on 11q22.3. J Med Genet. 2013;50:725-732. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 22] [Cited by in F6Publishing: 23] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 182. | Chan KY, Wong CM, Kwan JS, Lee JM, Cheung KW, Yuen MF, Lai CL, Poon RT, Sham PC, Ng IO. Genome-wide association study of hepatocellular carcinoma in Southern Chinese patients with chronic hepatitis B virus infection. PLoS One. 2011;6:e28798. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 47] [Cited by in F6Publishing: 54] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 183. | Chen K, Shi W, Xin Z, Wang H, Zhu X, Wu X, Li Z, Li H, Liu Y. Replication of genome wide association studies on hepatocellular carcinoma susceptibility loci in a Chinese population. PLoS One. 2013;8:e77315. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 29] [Cited by in F6Publishing: 35] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 184. | Hu Z, Liu Y, Zhai X, Dai J, Jin G, Wang L, Zhu L, Yang Y, Liu J, Chu M. New loci associated with chronic hepatitis B virus infection in Han Chinese. Nat Genet. 2013;45:1499-1503. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 111] [Cited by in F6Publishing: 118] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 185. | Chang SW, Fann CS, Su WH, Wang YC, Weng CC, Yu CJ, Hsu CL, Hsieh AR, Chien RN, Chu CM. A genome-wide association study on chronic HBV infection and its clinical progression in male Han-Taiwanese. PLoS One. 2014;9:e99724. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 44] [Cited by in F6Publishing: 43] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 186. | Pan W, Cheng G, Xing H, Shi J, Lu C, Wei J, Li L, Zhou C, Yuan Q, Zhou L. Leukocyte telomere length-related rs621559 and rs398652 genetic variants influence risk of HBV-related hepatocellular carcinoma. PLoS One. 2014;9:e110863. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 5] [Cited by in F6Publishing: 6] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 187. | Krautkramer KA, Linnemann AK, Fontaine DA, Whillock AL, Harris TW, Schleis GJ, Truchan NA, Marty-Santos L, Lavine JA, Cleaver O. Tcf19 is a novel islet factor necessary for proliferation and survival in the INS-1 β-cell line. Am J Physiol Endocrinol Metab. 2013;305:E600-E610. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 26] [Cited by in F6Publishing: 28] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 188. | Hoeller D, Hecker CM, Wagner S, Rogov V, Dötsch V, Dikic I. E3-independent monoubiquitination of ubiquitin-binding proteins. Mol Cell. 2007;26:891-898. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 119] [Cited by in F6Publishing: 121] [Article Influence: 7.1] [Reference Citation Analysis (0)] |
| 189. | Xing J, Ajani JA, Chen M, Izzo J, Lin J, Chen Z, Gu J, Wu X. Constitutive short telomere length of chromosome 17p and 12q but not 11q and 2p is associated with an increased risk for esophageal cancer. Cancer Prev Res (Phila). 2009;2:459-465. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 63] [Cited by in F6Publishing: 63] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 190. | Imamichi Y, Mizutani T, Ju Y, Matsumura T, Kawabe S, Kanno M, Yazawa T, Miyamoto K. Transcriptional regulation of human ferredoxin 1 in ovarian granulosa cells. Mol Cell Endocrinol. 2013;370:1-10. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 23] [Cited by in F6Publishing: 21] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 191. | Sheftel AD, Stehling O, Pierik AJ, Elsässer HP, Mühlenhoff U, Webert H, Hobler A, Hannemann F, Bernhardt R, Lill R. Humans possess two mitochondrial ferredoxins, Fdx1 and Fdx2, with distinct roles in steroidogenesis, heme, and Fe/S cluster biosynthesis. Proc Natl Acad Sci USA. 2010;107:11775-11780. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 190] [Cited by in F6Publishing: 243] [Article Influence: 17.4] [Reference Citation Analysis (0)] |
| 192. | Huang SS, Huang JS. TGF-beta control of cell proliferation. J Cell Biochem. 2005;96:447-462. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 163] [Cited by in F6Publishing: 168] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 193. | Massagué J, Xi Q. TGF-β control of stem cell differentiation genes. FEBS Lett. 2012;586:1953-1958. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 125] [Cited by in F6Publishing: 121] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 194. | Sethi A, Mao W, Wordinger RJ, Clark AF. Transforming growth factor-beta induces extracellular matrix protein cross-linking lysyl oxidase (LOX) genes in human trabecular meshwork cells. Invest Ophthalmol Vis Sci. 2011;52:5240-5250. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 139] [Cited by in F6Publishing: 150] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 195. | Larsson J, Blank U, Helgadottir H, Björnsson JM, Ehinger M, Goumans MJ, Fan X, Levéen P, Karlsson S. TGF-beta signaling-deficient hematopoietic stem cells have normal self-renewal and regenerative ability in vivo despite increased proliferative capacity in vitro. Blood. 2003;102:3129-3135. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 123] [Cited by in F6Publishing: 126] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 196. | Ferrari G, Cook BD, Terushkin V, Pintucci G, Mignatti P. Transforming growth factor-beta 1 (TGF-beta1) induces angiogenesis through vascular endothelial growth factor (VEGF)-mediated apoptosis. J Cell Physiol. 2009;219:449-458. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 225] [Cited by in F6Publishing: 240] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 197. | Sledzińska A, Hemmers S, Mair F, Gorka O, Ruland J, Fairbairn L, Nissler A, Müller W, Waisman A, Becher B. TGF-β signalling is required for CD4+ T cell homeostasis but dispensable for regulatory T cell function. PLoS Biol. 2013;11:e1001674. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 73] [Cited by in F6Publishing: 75] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 198. | Okumoto K, Hattori E, Tamura K, Kiso S, Watanabe H, Saito K, Saito T, Togashi H, Kawata S. Possible contribution of circulating transforming growth factor-beta1 to immunity and prognosis in unresectable hepatocellular carcinoma. Liver Int. 2004;24:21-28. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 65] [Cited by in F6Publishing: 70] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 199. | Sacco R, Leuci D, Tortorella C, Fiore G, Marinosci F, Schiraldi O, Antonaci S. Transforming growth factor beta1 and soluble Fas serum levels in hepatocellular carcinoma. Cytokine. 2000;12:811-814. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 36] [Cited by in F6Publishing: 37] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 200. | Hong MH, Chou YC, Wu YC, Tsai KN, Hu CP, Jeng KS, Chen ML, Chang C. Transforming growth factor-β1 suppresses hepatitis B virus replication by the reduction of hepatocyte nuclear factor-4α expression. PLoS One. 2012;7:e30360. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 40] [Cited by in F6Publishing: 42] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 201. | Cambien F, Ricard S, Troesch A, Mallet C, Générénaz L, Evans A, Arveiler D, Luc G, Ruidavets JB, Poirier O. Polymorphisms of the transforming growth factor-beta 1 gene in relation to myocardial infarction and blood pressure. The Etude Cas-Témoin de l’Infarctus du Myocarde (ECTIM) Study. Hypertension. 1996;28:881-887. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 264] [Cited by in F6Publishing: 282] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 202. | Ben-Ari Z, Mor E, Papo O, Kfir B, Sulkes J, Tambur AR, Tur-Kaspa R, Klein T. Cytokine gene polymorphisms in patients infected with hepatitis B virus. Am J Gastroenterol. 2003;98:144-150. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 167] [Cited by in F6Publishing: 188] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 203. | Kwon OS, Song SH, Ju KT, Chung MG, Park DK, Kim SS, Kim YS, Koo YS, Kim YK, Choi DJ. [Polymorphism in codons 10 and 25 of the transforming growth factor-beta1 gene in Korean population and in patients with liver cirrhosis and hepatocellular carcinoma]. Korean J Gastroenterol. 2003;42:212-219. [PubMed] [Cited in This Article: ] |
| 204. | Migita K, Miyazoe S, Maeda Y, Daikoku M, Abiru S, Ueki T, Yano K, Nagaoka S, Matsumoto T, Nakao K. Cytokine gene polymorphisms in Japanese patients with hepatitis B virus infection--association between TGF-beta1 polymorphisms and hepatocellular carcinoma. J Hepatol. 2005;42:505-510. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 108] [Cited by in F6Publishing: 122] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 205. | Shi HZ, Ren P, Lu QJ, Niedrgethmnn M, Wu GY. Association between EGF, TGF-β1 and TNF-α gene polymorphisms and hepatocellular carcinoma. Asian Pac J Cancer Prev. 2012;13:6217-6220. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 17] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 206. | Qi P, Chen YM, Wang H, Fang M, Ji Q, Zhao YP, Sun XJ, Liu Y, Gao CF. -509C>T polymorphism in the TGF-beta1 gene promoter, impact on the hepatocellular carcinoma risk in Chinese patients with chronic hepatitis B virus infection. Cancer Immunol Immunother. 2009;58:1433-1440. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 47] [Cited by in F6Publishing: 47] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 207. | Hosseini Razavi A, Azimzadeh P, Mohebbi SR, Hosseini SM, Romani S, Khanyaghma M, Hatami Y, Sharifian A, Zali MR. Lack of Association Between Transforming Growth Factor Beta 1 -509C/T and +915G/C Polymorphisms and Chronic Hepatitis B in Iranian Patients. Hepat Mon. 2014;14:e13100. [PubMed] [Cited in This Article: ] |
| 208. | Kim YJ, Lee HS, Im JP, Min BH, Kim HD, Jeong JB, Yoon JH, Kim CY, Kim MS, Kim JY. Association of transforming growth factor-beta1 gene polymorphisms with a hepatocellular carcinoma risk in patients with chronic hepatitis B virus infection. Exp Mol Med. 2003;35:196-202. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 63] [Cited by in F6Publishing: 73] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 209. | Li XD, Wu LM, Xie HY, Xu X, Zhou L, Liang TB, Wang WL, Shen Y, Zhang M, Zheng SS. No association exists between E-cadherin gene polymorphism and tumor recurrence in patients with hepatocellular carcinoma after transplantation. Hepatobiliary Pancreat Dis Int. 2007;6:254-258. [PubMed] [Cited in This Article: ] |
| 210. | Liu K, Zhang L, Lin X, Chen L, Shi H, Magaye R, Zou B, Zhao J. Association of GST genetic polymorphisms with the susceptibility to hepatocellular carcinoma (HCC) in Chinese population evaluated by an updated systematic meta-analysis. PLoS One. 2013;8:e57043. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 14] [Cited by in F6Publishing: 16] [Article Influence: 1.5] [Reference Citation Analysis (0)] |