Published online Nov 28, 2020. doi: 10.3748/wjg.v26.i44.6929
Peer-review started: April 18, 2020
First decision: July 29, 2020
Revised: August 6, 2020
Accepted: September 17, 2020
Article in press: September 17, 2020
Published online: November 28, 2020
Gastric cancer (GC) is one of the most frequently diagnosed gastrointestinal cancers throughout the world. Novel prognostic biomarkers are required to predict the prognosis of GC.
To identify a multi-long noncoding RNA (lncRNA) prognostic model for GC.
Transcriptome data and clinical data were downloaded from The Cancer Genome Atlas. COX and least absolute shrinkage and selection operator regression analyses were performed to screen for prognosis associated lncRNAs. Receiver operating characteristic curve and Kaplan-Meier survival analyses were applied to evaluate the effectiveness of the model.
The prediction model was established based on the expression of AC007991.4, AC079385.3, and AL109615.2 Based on the model, GC patients were divided into “high risk” and “low risk” groups to compare the differences in survival. The model was re-evaluated with the clinical data of our center.
The 3-lncRNA combination model is an independent prognostic factor for GC.
Core Tip: A model to predict survival of patients with gastric cancer was developed and validated by RNA sequencing and real-time reverse transcription-polymerase chain reaction assays. The model had an excellent performance: The areas under the curves for 3-year and 5-year survival were 0.78 and 0.75, respectively. The C-index was 0.72 (se = 0.022, 95% confidence interval: 0.67-0.76). The model contributed as a poor independent prognostic factor both in disease free survival and overall survival.
- Citation: Zhang J, Piao HY, Wang Y, Lou MY, Guo S, Zhao Y. Development and validation of a three-long noncoding RNA signature for predicting prognosis of patients with gastric cancer. World J Gastroenterol 2020; 26(44): 6929-6944
- URL: https://www.wjgnet.com/1007-9327/full/v26/i44/6929.htm
- DOI: https://dx.doi.org/10.3748/wjg.v26.i44.6929
Gastric cancer (GC), the fourth most frequently diagnosed malignant tumor, has the second highest cancer-related mortality rate worldwide, with high morbidity in Asia[1]. Moreover, GC is still the second most prevalent cancer in China[2]. Most patients are diagnosed at the advanced stage and are prone to chemoresistance and recurrence. It leads to an overall 5-year overall survival rate of less than 25%[3]. Therefore, developing a novel and reliable prognostic stratification system that could be applied to clinical risk assessment would be of great significance for the treatment and follow-up of GC patients.
Long noncoding RNAs (lncRNAs) have limited protein-coding ability, exceeding 200 nucleotides in length[4]. Emerging evidence indicates that lncRNAs are involved in tumor initiation and progression through gene expression regulation from epigenetic to post-transcriptional levels[5-7]. Not surprisingly, some studies have shed light on the role of lncRNAs in GC. GClnc1 spurred tumorigenesis and metastasis by recruiting the WDR5/KAT2A complex as a “scaffold” in GC[8]. HOXC-AS3 could be regulated by abnormal histone modification and functioned as an essential element in GC proliferation and migration[9]. Therefore, the abnormal expression of lncRNAs may indirectly reflect the occurrence and development of GC. As genes do not usually act alone, it is necessary to select suitable lncRNAs and establish a multi-lncRNA prediction model, which may play a pivotal role in evaluating the prognosis of GC.
In the present research, we analyzed the expression of lncRNAs associated with GC patients’ survival in The Cancer Genome Atlas (TCGA). We aimed to develop and validate a useful muti-lncRNA combination prediction model that might be useful in helping predict GC survival by performing COX regression and the least absolute shrinkage and selection operator (LASSO)[10].
RNA sequencing and matching medical data were acquired from TCGA (https://portal.gdc.cancer.gov/). Raw data were obtained through the “RTCGA Toolbox package” (R platform). Overall, 407 samples, including 375 GC and 32 normal tissues, were analyzed. The expression of lncRNAs was acquired from Illumina HiSeq-RNASeq platforms. Moreover, the clinical data of GC patients were also downloaded. R was utilized to evaluate findings.
Package “edgeR” was utilized to evaluate differentially expressed lncRNAs (DELs). P < 0.01 and |logFC| ≥ 2 were utilized as cutoff points. The R package "survival” was used to perform univariate COX regression analysis. Meaningful lncRNAs in univariate COX regression were incorporated into the construction of the least absolute shrinkage and selection operator (LASSO) regression (package “glmnet”) to minimize the overfitting caused by univariate COX regression. Besides, multivariate COX analysis was performed to screen the independent risk factors associated with GC patients’ prognosis. R package “survminer” was used for visualization.
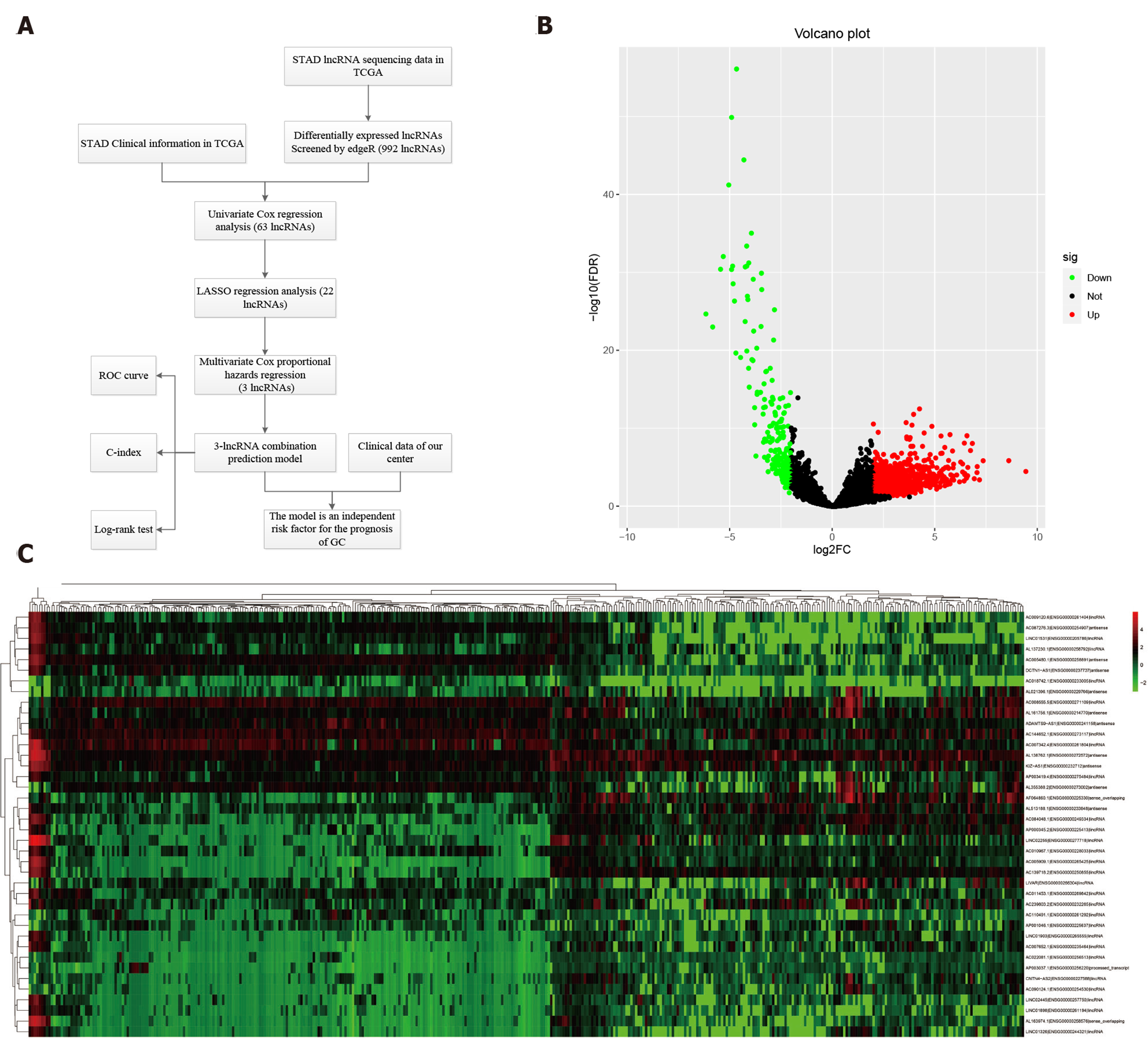
As time-dependent survival receiver operating characteristic (ROC) curve is an essential measure of the predictive power of a prognostic model, we used the R package “survival ROC” to assess the role of lncRNA combination prediction model in predicting 5- and 3-year survival. Kaplan-Meier survival analysis was performed to explore correlations between the lncRNA combination model and the overall survival (OS). This study was carried out according to the flow chart (Figure 1A).
Overall, 200 GC and non-tumorous adjacent tissues were acquired from people undergoing surgery at the Liaoning Province Cancer Hospital and Institute from 2012 and 2014. Individuals were asked to sign an informed consent form before the operation. The hospital’s Ethics Committee approved the study. No preoperative chemotherapy or radiotherapy was performed on enrolled patients. Gastrectomy and D2 lymph node dissection were applied to all patients. Total RNA was extracted from patient tissue samples. Tumor staging was based on the tumor-node-metastasis (TNM) staging system (8th edition). One hundred and forty-two men and 58 women were enrolled in this study; their average age was 65 years (range, 42-78 years). Postoperative adjuvant chemotherapy was applied to stage IIA and above disease. Table 1 shows the clinicopathological parameters of the patients. Follow-up was the same as that in previous studies[9], and the follow-up deadline is December 31, 2019.
| Characteristic | n | Disease-free survival | Overall survival | ||||
| Time (mo) | P value | F | Time (mo) | P value | F | ||
| Age, median, yr | 0.14 | 2.17 | 0.20 | 1.72 | |||
| ≥ 60 | 107 | 42.46 | 49.39 | ||||
| < 60 | 93 | 50.48 | 55.79 | ||||
| Gender | 0.15 | 2.10 | 0.12 | 2.42 | |||
| Male | 142 | 43.86 | 50.23 | ||||
| Female | 58 | 49.58 | 54.82 | ||||
| Bormann type | 0.00 | 44.98 | 0.00 | 43.62 | |||
| I | 15 | 69.93 | 69.93 | ||||
| II | 88 | 45.57 | 51.73 | ||||
| III | 94 | 29.35 | 36.99 | ||||
| IV | 3 | 15.33 | 25.00 | ||||
| Tumor size | 0.00 | 28.46 | 0.00 | 27.42 | |||
| ≥ 5 cm | 87 | 31.51 | 39.48 | ||||
| < 5 cm | 113 | 57.46 | 62.07 | ||||
| Location | 0.09 | 4.91 | 0.13 | 4.11 | |||
| Upper | 44 | 35.22 | 43.33 | ||||
| Middle | 56 | 49.33 | 54.92 | ||||
| Low | 100 | 48.92 | 54.31 | ||||
| Lauren type | 0.62 | 0.96 | 0.63 | 0.93 | |||
| Intestinal | 95 | 45.22 | 51.26 | ||||
| Mixed | 45 | 41.08 | 47.06 | ||||
| Diffuse | 60 | 50.23 | 56.12 | ||||
| Tumor differentiation | 0.16 | 2.00 | 0.19 | 1.71 | |||
| Moderate and well | 66 | 50.80 | 56.28 | ||||
| Poor | 134 | 44.57 | 50.74 | ||||
| Vessel invasion | 0.00 | 16.30 | 0.00 | 14.73 | |||
| Yes | 59 | 31.07 | 38.83 | ||||
| No | 141 | 52.08 | 57.54 | ||||
| Perineural invasion | 0.00 | 11.24 | 0.00 | 11.52 | |||
| Yes | 51 | 28.01 | 37.39 | ||||
| No | 149 | 52.14 | 57.41 | ||||
| TNM stage | 0.00 | 335.89 | 0.00 | 360.28 | |||
| I | 42 | 66.45 | 66.50 | ||||
| II | 47 | 64.36 | 68.85 | ||||
| III | 109 | 18.70 | 28.50 | ||||
| IV | 2 | 5.00 | 8.50 | ||||
| AC007991.4 | 0.05 | 3.77 | 0.05 | 3.95 | |||
| Overexpression | 68 | 53.02 | 58.07 | ||||
| Weak expression | 132 | 42.70 | 49.24 | ||||
| AC079385.3 | 0.00 | 14.23 | 0.00 | 14.71 | |||
| Overexpression | 95 | 36.26 | 43.94 | ||||
| Weak expression | 105 | 55.12 | 59.92 | ||||
| AL109615.2 | 0.00 | 10.48 | 0.02 | 9.87 | |||
| Overexpression | 113 | 39.04 | 46.05 | ||||
| Weak expression | 87 | 56.27 | 60.90 | ||||
| Model | 0.00 | 21.76 | 0.00 | 21.34 | |||
| High risk | 95 | 37.66 | 44.46 | ||||
| Low risk | 105 | 53.91 | 59.27 | ||||
The gastric epithelial cell line GES-1 and gastric cancer cell lines AGS and MKN45 were obtained from China Medical University (Shenyang, China). Cells were maintained in RPMI 1640 supplemented with 10% fetal bovine serum, penicillin, and streptomycin (Invitrogen, United States) at 37 °C at 5% CO2/1% O2. Analyses were performed at least three times.
TRIzol, Promega cDNA core kit, and SYBR Master Mixture were utilized to create cDNA and conduct real-time reverse transcription-polymerase chain reaction (RT-PCR) as those described in previous studies[6,11,12].
Data are shown as the mean ± standard deviation. Student’s t-test, Wilcoxon-signed rank test, and ANOVA were used for statistical analyses through SPSS 23.0 (IBM, NY, United States). The relationship between clinical data and expression of biomarkers was tested by χ2 test or Fisher’s exact test. The log-rank test and COX proportional hazards model were used for survival analysis. P < 0.05 represented statistical significance.
RNA sequencing involved 375 GC and 32 para-carcinoma tissues. EdgeR was used to determine the DELs (P < 0.01 and |logFC| ≥ 2). A total of 992 DELs were obtained, of which 772 were up-regulated, and 220 were down-regulated (Figure 1B and Supplementary Table 1). Moreover, the heatmap elucidated the top 50 DELs (Figure 1C).
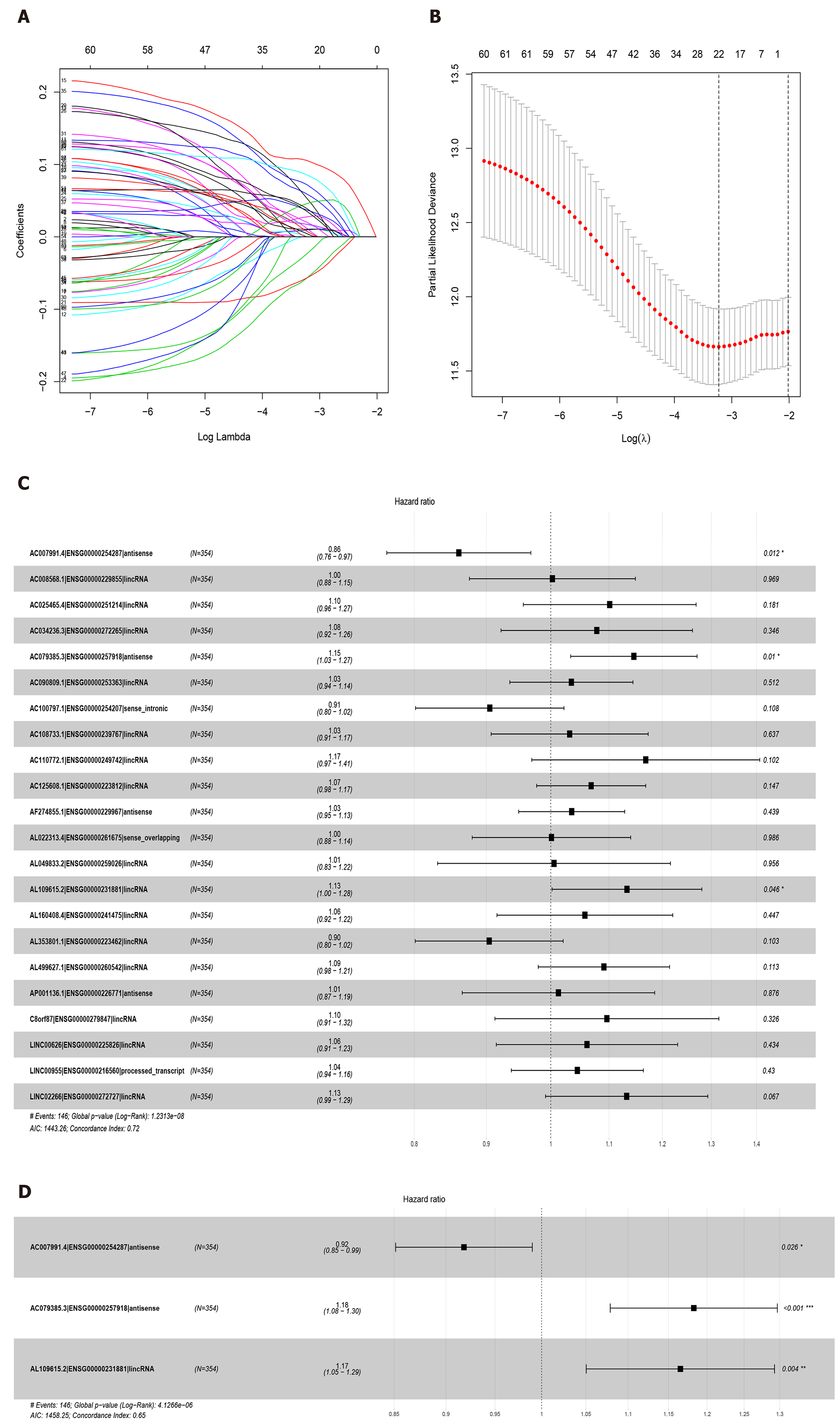
Sixty-three lncRNAs were associated with the survival of GC by univariate COX regression analysis (Supplementary Table 2). The lncRNA-seq expression profile of the 63 lncRNAs and the clinical data were extracted for LASSO regression (Figure 2A). As a result of the LASSO penalized uni-Cox model, 22 lncRNAs were finally identified (Figure 2B) and incorporated into the multivariate COX model to identify independent prognostic factors associated with survival. As a result of LASSO and COX model, AC007991.4 (ENSG00000254287, antisense), AC079385.3 (ENSG00000257918, antisense), and AL109615.2 (ENSG00000231881, lincRNA) were finally identified (Figure 2C and D).
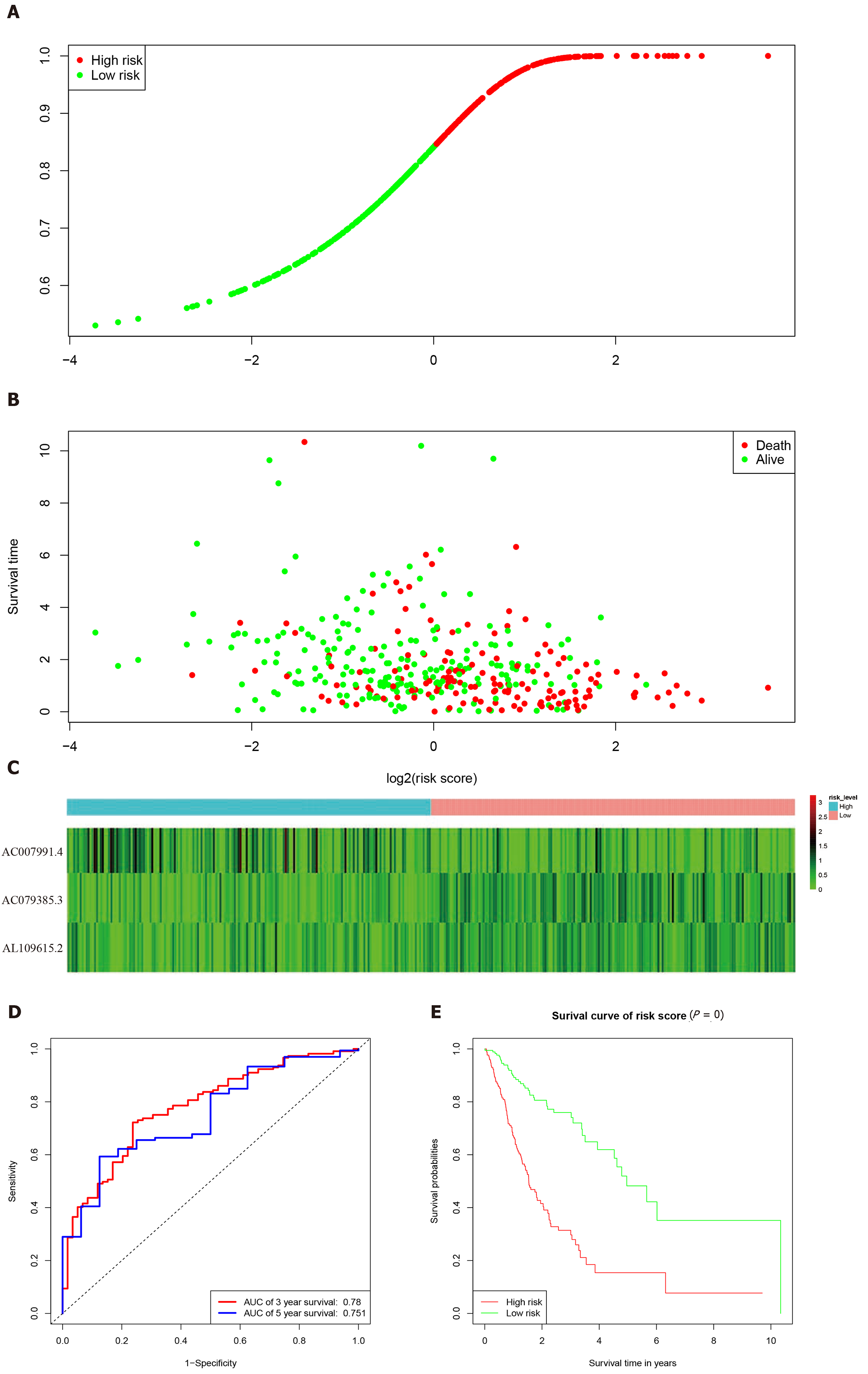
According to the expression levels of the 3-lncRNA combination, all samples were divided into a high-risk group (red, Figure 3A) and low-risk group (green, Figure 3A). The survival time in years is shown in Figure 3B (red dots indicate death, and green dots indicate alive). The heatmap elucidated the expression of the three lncRNAs according to the risk level (Figure 3C). ROC analysis assessed the role of the 3-lncRNA combination prediction model in predicting survival. The areas under the curves for 3-year and 5-year survival were 0.78 and 0.75, respectively (Figure 3D). The C-index was 0.72 (se = 0.022, 95% confidence interval [CI]: 0.67-0.76), indicating that the model was a good predictor of patient survival. The low-risk group had a longer survival time than the high-risk group (Figure 3E, P < 0.001). Based on this, we obtained a 3-lncRNA combination prognostic model for GC patients: “risk score = -0.92 × AC007991.4 + 1.18 × AC079385.3 + 1.17 × AL109615.2”, and prepared for further verification (cutoff value = 6.58).
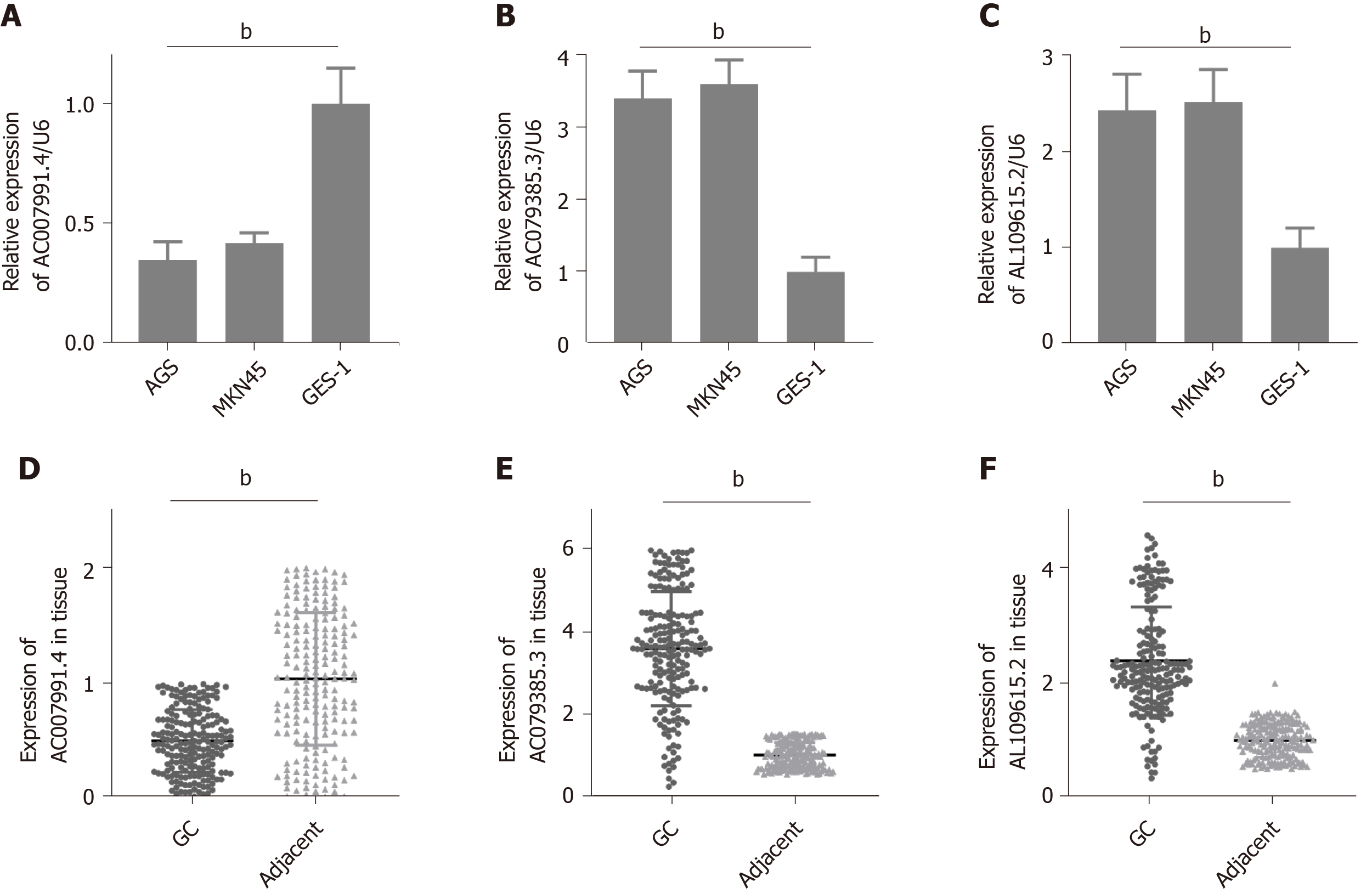
We then tested the expression of AC007991.4, AC079385.3, and AL109615.2 in GC cells and tissues. The results were consistent with those of bioinformatics prediction: AC007991.4 was weakly expressed in both GC cells and tissues (Figure 4A and D), while AC079385.3 and AL109615.2 were overexpressed in GC (Figure 4B, C, E, and F).
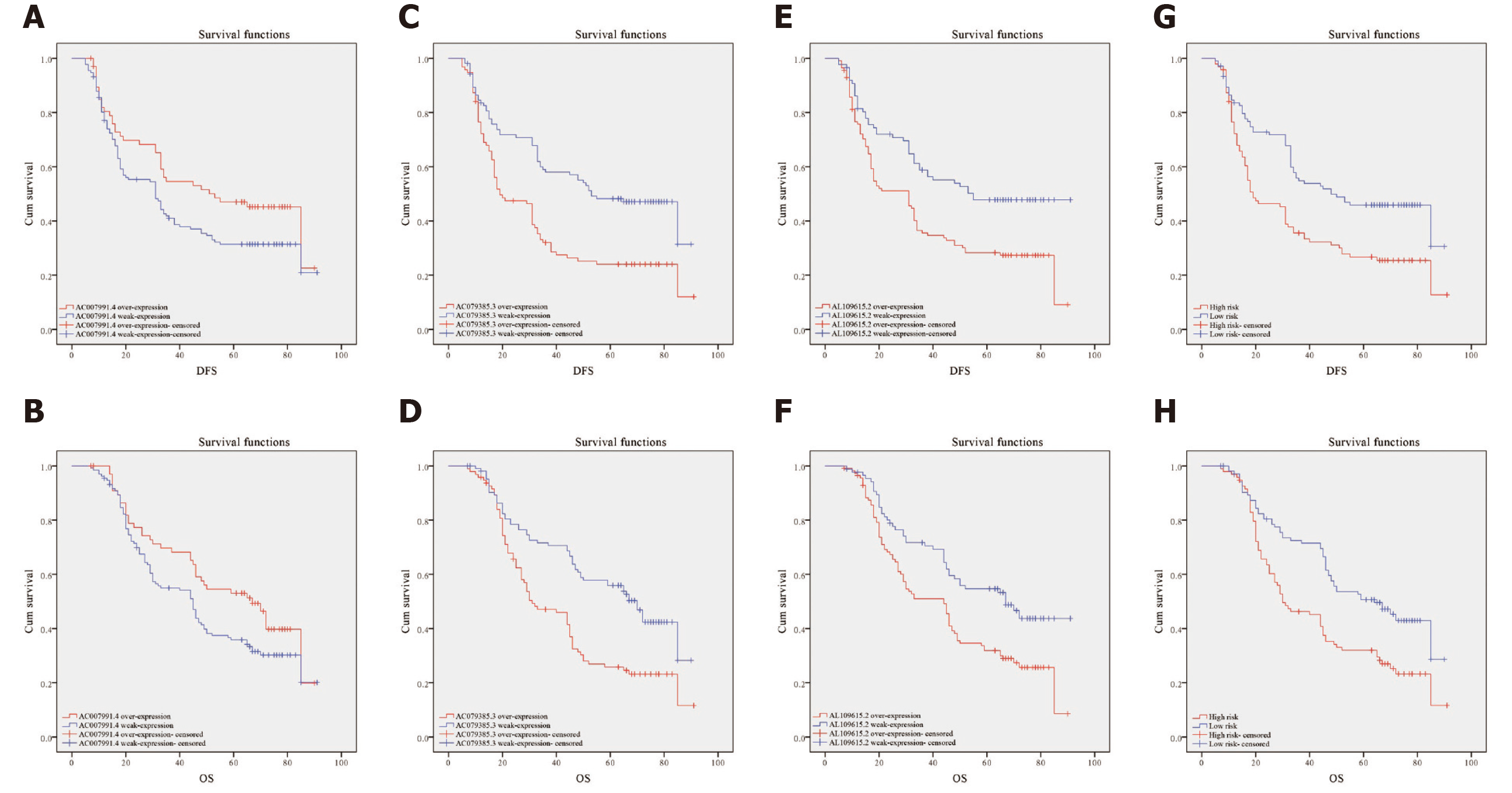
Clinical data of 200 patients were enrolled in this study to verify the effectiveness of the 3-lncRNA combination prediction model. The disease-free survival (DFS) ranged from 5-91 mo, OS ranged from 7-91 mo, and 126 patients died before the end of follow-up. Then, we detected the expression of AC007991.4, AC079385.3, and AC079385.3 in these 200 tissue samples and evaluated the correlation between their expression and survival. As shown in Table 1, the Bormann type, tumor size, vessel invasion, perineural invasion, TNM, AC079385.3, and AL109615.2 were all associated with the poor DFS and OS (P < 0.05). Not surprisingly, “high risk” in the 3-lncRNA combination prediction model was also a poor prognostic factor for DFS (37.66 vs 53.91, P < 0.01, Figure 5) and OS (44.46 vs 59.27, P < 0.01, Figure 5).
The parameters with P < 0.05 in the univariate analysis were included in the COX proportional hazard model. The role of TNM stage in prognosis remained unshaken [DFS: P = 0.00, hazard ratio (HR) = 106.50, 95%CI: 30.76-368.77; OS: P = 0, HR = 94.08, 95%CI: 30.14-293.65, Table 2]. The “high risk” also contributed as a poor independent prognostic factor both in DFS (P = 0.03, HR = 2.38, 95%CI: 1.34-4.23) and OS (P = 0.02, HR = 2.62, 95%CI: 1.44-4.77). In addition, the overexpression of AC079385.3 also foreshadowed the poor of survival of GC (DFS: P = 0.04, HR = 1.67, 95%CI: 1.46-1.99; OS: P = 0.04, HR = 1.72, 95%CI: 1.49-1.96). Its biological function in GC is worth further discussion.
| Variable | Disease-free survival | Overall survival | ||||
| P value | HR | 95%CI | P value | HR | 95%CI | |
| Bormann type | 0.20 | 1.26 | 0.90-1.77 | 0.13 | 1.29 | 0.93-1.91 |
| Tumor size | 0.41 | 0.86 | 0.59-1.24 | 0.40 | 0.85 | 0.59-1.24 |
| Vessel invasion | 0.06 | 0.70 | 0.48-1.01 | 0.16 | 0.76 | 0.53-1.11 |
| Perineural invasion | 0.24 | 0.80 | 0.56-1.16 | 0.09 | 0.73 | 0.50-1.05 |
| TNM stage | 0.00 | 106.50 | 30.76-368.77 | 0.00 | 94.08 | 30.14-293.65 |
| Model | 0.04 | 2.27 | 1.46-3.99 | 0.03 | 2.64 | 1.74-4.27 |
According to the 3-lncRNA combination prediction model and the expression of lncRNAs, we obtained the risk score (risk score = -0.92 × AC007991.4 + 1.18 × AC079385.3 + 1.17 × AL109615.2). Patients were classified as “high risk” and “low risk" according to the cutoff value. The combination was associated with the Bormann type (P = 0, χ2 = 14.29) and TNM stage (P = 0.01, χ2 = 10.85) of GC (Table 3).
| Characteristic | Model | |||
| High risk (%) | Low risk (%) | P value | χ2 | |
| Age, median, yr | 0.11 | 0.74 | ||
| ≥ 60 | 52 (48.6) | 55 (51.4) | ||
| < 60 | 43 (46.2) | 50 (53.8) | ||
| Gender | 0.27 | 1.23 | ||
| Male | 71 (50.0) | 71 (50.0) | ||
| Female | 24 (41.4) | 34 (58.6) | ||
| Bormann type | 0.00 | 14.29 | ||
| I | 2 (13.3) | 13 (86.7) | ||
| II | 36 (40.9) | 52 (59.1) | ||
| III | 56 (59.6) | 38 (40.4) | ||
| IV | 1 (33.3) | 2 (66.7) | ||
| Tumor size | 0.06 | 3.64 | ||
| ≥ 5 cm | 48 (55.2) | 39 (44.8) | ||
| < 5 cm | 47 (41.6) | 66 (58.4) | ||
| Location | 0.77 | 0.52 | ||
| Upper | 23 (52.3) | 21 (47.7) | ||
| Middle | 26 (46.4) | 30 (53.6) | ||
| Low | 46 (46.0) | 54 (54.0) | ||
| Lauren type | 0.82 | 0.39 | ||
| Intestinal | 45 (47.4) | 50 (52.6) | ||
| Mixed | 23 (51.1) | 22 (48.9) | ||
| Diffuse | 27 (45.0) | 33 (55.0) | ||
| Tumor differentiation | 0.20 | 1.72 | ||
| Moderate and well | 27 (40.9) | 39 (59.1) | ||
| Poor | 68 (50.7) | 66 (49.3) | ||
| Vessel invasion | 0.54 | 0.38 | ||
| Yes | 30 (50.8) | 29 (49.2) | ||
| No | 65 (46.1) | 76 (53.9) | ||
| Perineural invasion | 0.06 | 3.52 | ||
| Yes | 30 (58.8) | 21 (41.2) | ||
| No | 65 (43.6) | 84 (56.4) | ||
| TNM stage | 0.01 | 10.85 | ||
| I | 13 (31.0) | 29 (69.0) | ||
| II | 18 (38.3) | 29 (61.7) | ||
| III | 63 (57.8) | 46 (42.2) | ||
| IV | 1 (50.0) | 1 (50.0) | ||
LncRNAs are a class of molecules with functional relevance for gene expression regulation and have been discovered recently. With the expansion of research on lncRNA function, growing evidence demonstrates that a series of lncRNAs are aberrantly expressed in cancer and involved in cancer progression regulation. Moreover, they have increasingly complex functions. As “scaffolds”, they stimulate the interaction between proteins. As “guides”, lncRNAs enable the mixing of protein and genes[13]. As “enhancers”, lncRNAs control transcription of nearby genes. As “decoys”, lncRNAs are bound to microRNAs or proteins[14,15].
Because of the critical regulatory role of lncRNAs in tumorigenesis and development, more and more lncRNAs have been recognized as biomarkers for GC treatment and progenies[12,16]. MAFG-AS1 promoted GC cell proliferation and invasion and might be a valuable prognostic biomarker[16]. LncRNA pcsk2-2:1 could be parceled into serum exosome and acted as a diagnostic biomarker for GC[17]. Fattahi et al[18] summarized the role of famous lncRNAs such as H19, HOTAIR, UCA1, and PVT1 as molecular markers in GC. However, these studies only discussed the predictive value of a single biomarker, which may not be sufficient to predict the prognosis of GC. In the present study, clinical data and RNA-seq data of GC patients were obtained from TCGA as a training set. The patients’ data of our center served as the verification set. P < 0.01 and |logFC| ≥ 2 were used as the cutoff points to obtain the DELs. To improve the regression model’s prediction accuracy, LASSO and COX regression analyses were applied to examine the correlation between the expression of lncRNAs and GC patients’ survival. Compared with previous studies[17], this method is the first to study GC related lncRNA markers; it effectively minimizes the overfitting caused by univariate COX regression[19,20]. LASSO is a kind of regression analysis introduced in statistics and machine learning[21,22]. Compared with the traditional model, LASSO improves the prediction accuracy and interpretability through variable selection and regularization. The LASSO regression evaluation process includes relationship to ridge regression, best subset selection, the connections between lasso coefficient estimates, and soft thresholding[23,24].
Then, we got a 3-lncRNA combination prediction model: AC007991.4, AC079385.3, and AL109615.2. They were all potential prognostic independent risk factors for GC. According to the expression levels of the 3-lncRNA combination, all samples were divided into a high-risk group and low-risk group. This model could predict patients’ poor prognosis in the high expression group in both the training and validation sets. Subsequently, we verified the expression of these three lncRNAs in GC tissues and cells. AC007991.4 was weakly expressed in GC. However, both AC079385.3 and AL109615.2 were potential onco-lncRNAs.
AC007991.4 and AC079385.3 are all antisense lncRNAs, and located at chromosome 8: 39918076-39920890 and chromosome 12: 106714924-106733066. Their molecular functions have not been reported. As a long intergenic non-coding RNA, AL109615.2 locates at chromosome 6: 44058792-44089288, which could competitively bind miR-133b with vascular endothelial growth factor C to induce colorectal cancer cell metastasis[25]. According to the expression of AC007991.4, AC079385.3, and AL109615.2, GC patients were divided into “high risk” and “low risk” groups. Both the area under the ROC curve and C-index suggested that the model has appropriate prediction performance.
Moreover, it was confirmed by the verification set that the prediction model was an independent risk factor for the prognosis of GC. Besides, it was positively correlated with Bormann type and TNM stage in GC. It is also worth pointing out that AC079385.3 was a prognostic risk factor for GC in both the training and validation sets, suggesting that it might play a pivotal role in the model. Its molecular biological function and position in the development of GC require further study.
In conclusion, we present a 3-lncRNA model for evaluating survival in GC patients, which may be an independent prognostic factor. Clinicians can obtain the expression levels of AC007991.4, AC079385.3, and AL109615.2 in tissue samples by RT-PCR and calculate the corresponding risk values (risk score = -0.92 × AC007991.4 + 1.18 × AC079385.3 + 1.17 × AL109615.2, cutoff value = 6.58). And then it can be used to evaluate the prognosis of patients. Besides, we can visualize the model and other related risk factors to assess the modification’s effectiveness in future clinical work.
Gastric cancer (GC) is one of the most frequently diagnosed gastrointestinal cancers throughout the world. It is necessary to identify a multi-long noncoding RNA (lncRNA) prognostic model for GC.
Abnormal expression of lncRNAs may indirectly reflect the occurrence and development of GC. As genes do not usually act alone, it is necessary to select suitable lncRNAs and establish a multi-lncRNA prediction model.
To construct a multi-lncRNA combination model to predict the prognosis of gastric cancer patients.
The RNA-seq dataset and clinical dataset of GC in The Cancer Genome Atlas were used in this study. The least absolute shrinkage and selection operator and COX models were used to identify meaningful modules and hub genes. Clinical data of 200 patients were used to evaluate the clinical significance of the multi-lncRNA combination model via survival analysis.
We found a 3-lncRNA combination prediction model: AC007991.4, AC079385.3, and AL109615.2. It could effectively predict the prognosis of GC. AC079385.3 was found to be a prognostic risk factor for GC, and it may play an important role in the development of GC. Least absolute shrinkage and selection operator improved prediction accuracy and interpretability through variable selection and regularization.
The 3-lncRNA combination model (risk score = -0.92 × AC007991.4 + 1.18 × AC079385.3 + 1.17 × AL109615.2) is an independent prognostic factor for GC.
Clinicians can obtain the expression levels of AC007991.4, AC079385.3, and AL109615.2 in tissue samples by real-time reverse transcription-polymerase chain reaction and calculate the corresponding risk values.
Manuscript source: Unsolicited manuscript
Specialty type: Gastroenterology and hepatology
Country/Territory of origin: China
Peer-review report’s scientific quality classification
Grade A (Excellent): 0
Grade B (Very good): B
Grade C (Good): C
Grade D (Fair): 0
Grade E (Poor): E
P-Reviewer: Kinami S, Tanțău M S-Editor: Gao CC L-Editor: Wang TQ P-Editor: Wang LL
| 1. | Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7-30. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12667] [Cited by in F6Publishing: 13879] [Article Influence: 3469.8] [Reference Citation Analysis (4)] |
| 2. | Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F, Jemal A, Yu XQ, He J. Cancer statistics in China, 2015. CA Cancer J Clin. 2016;66:115-132. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11444] [Cited by in F6Publishing: 12500] [Article Influence: 1562.5] [Reference Citation Analysis (0)] |
| 3. | Van Cutsem E, Sagaert X, Topal B, Haustermans K, Prenen H. Gastric cancer. Lancet. 2016;388:2654-2664. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1282] [Cited by in F6Publishing: 1367] [Article Influence: 170.9] [Reference Citation Analysis (0)] |
| 4. | Schmitt AM, Chang HY. Long Noncoding RNAs in Cancer Pathways. Cancer Cell. 2016;29:452-463. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1994] [Cited by in F6Publishing: 2241] [Article Influence: 280.1] [Reference Citation Analysis (0)] |
| 5. | Zhang T, Piao HY, Guo S, Zhao Y, Wang Y, Zheng ZC, Zhang J. LncRNA PCGEM1 enhances metastasis and gastric cancer invasion through targeting of miR-129-5p to regulate P4HA2 expression. Exp Mol Pathol. 2020;116:104487. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 12] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 6. | Zhang J, Guo S, Piao HY, Wang Y, Wu Y, Meng XY, Yang D, Zheng ZC, Zhao Y. ALKBH5 promotes invasion and metastasis of gastric cancer by decreasing methylation of the lncRNA NEAT1. J Physiol Biochem. 2019;75:379-389. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 130] [Cited by in F6Publishing: 199] [Article Influence: 39.8] [Reference Citation Analysis (0)] |
| 7. | Zhang J, Jin HY, Wu Y, Zheng ZC, Guo S, Wang Y, Yang D, Meng XY, Xu X, Zhao Y. Hypoxia-induced LncRNA PCGEM1 promotes invasion and metastasis of gastric cancer through regulating SNAI1. Clin Transl Oncol. 2019;21:1142-1151. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 46] [Cited by in F6Publishing: 39] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 8. | Sun TT, He J, Liang Q, Ren LL, Yan TT, Yu TC, Tang JY, Bao YJ, Hu Y, Lin Y, Sun D, Chen YX, Hong J, Chen H, Zou W, Fang JY. LncRNA GClnc1 Promotes Gastric Carcinogenesis and May Act as a Modular Scaffold of WDR5 and KAT2A Complexes to Specify the Histone Modification Pattern. Cancer Discov. 2016;6:784-801. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 225] [Cited by in F6Publishing: 293] [Article Influence: 36.6] [Reference Citation Analysis (0)] |
| 9. | Zhang J, Wu Y, Lin YH, Guo S, Ning PF, Zheng ZC, Wang Y, Zhao Y. Prognostic value of hypoxia-inducible factor-1 alpha and prolyl 4-hydroxylase beta polypeptide overexpression in gastric cancer. World J Gastroenterol. 2018;24:2381-2391. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 34] [Cited by in F6Publishing: 35] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 10. | Gao H, Li L, Xiao M, Guo Y, Shen Y, Cheng L, Tang M. Elevated DKK1 expression is an independent unfavorable prognostic indicator of survival in head and neck squamous cell carcinoma. Cancer Manag Res. 2018;10:5083-5089. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 17] [Cited by in F6Publishing: 16] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 11. | Zhang J, Piao HY, Guo S, Wang Y, Zhang T, Zheng ZC, Zhao Y. LINC00163 inhibits the invasion and metastasis of gastric cancer cells as a ceRNA by sponging miR-183 to regulate the expression of AKAP12. Int J Clin Oncol. 2020;25:570-583. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 12] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 12. | Piao HY, Guo S, Wang Y, Zhang J. Exosomal Long Non-Coding RNA CEBPA-AS1 Inhibits Tumor Apoptosis and Functions as a Non-Invasive Biomarker for Diagnosis of Gastric Cancer. Onco Targets Ther. 2020;13:1365-1374. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 32] [Cited by in F6Publishing: 36] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 13. | Chen C, He W, Huang J, Wang B, Li H, Cai Q, Su F, Bi J, Liu H, Zhang B, Jiang N, Zhong G, Zhao Y, Dong W, Lin T. LNMAT1 promotes lymphatic metastasis of bladder cancer via CCL2 dependent macrophage recruitment. Nat Commun. 2018;9:3826. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 169] [Cited by in F6Publishing: 254] [Article Influence: 42.3] [Reference Citation Analysis (0)] |
| 14. | Morris KV, Mattick JS. The rise of regulatory RNA. Nat Rev Genet. 2014;15:423-437. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 908] [Cited by in F6Publishing: 913] [Article Influence: 91.3] [Reference Citation Analysis (0)] |
| 15. | Mattick JS, Rinn JL. Discovery and annotation of long noncoding RNAs. Nat Struct Mol Biol. 2015;22:5-7. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 430] [Cited by in F6Publishing: 467] [Article Influence: 51.9] [Reference Citation Analysis (0)] |
| 16. | Li C, Wu R, Xing Y. MAFG-AS1 is a novel clinical biomarker for clinical progression and unfavorable prognosis in gastric cancer. Cell Cycle. 2020;19:601-609. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 14] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 17. | Cai C, Zhang H, Zhu Y, Zheng P, Xu Y, Sun J, Zhang M, Lan T, Gu B, Li S, Ma P. Serum Exosomal Long Noncoding RNA pcsk2-2:1 As A Potential Novel Diagnostic Biomarker For Gastric Cancer. Onco Targets Ther. 2019;12:10035-10041. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 31] [Cited by in F6Publishing: 46] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 18. | Fattahi S, Kosari-Monfared M, Golpour M, Emami Z, Ghasemiyan M, Nouri M, Akhavan-Niaki H. LncRNAs as potential diagnostic and prognostic biomarkers in gastric cancer: A novel approach to personalized medicine. J Cell Physiol. 2020;235:3189-3206. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 59] [Cited by in F6Publishing: 78] [Article Influence: 15.6] [Reference Citation Analysis (0)] |
| 19. | Yin XH, Yu LP, Zhao XH, Li QM, Liu XP, He L. Development and validation of a 4-gene combination for the prognostication in lung adenocarcinoma patients. J Cancer. 2020;11:1940-1948. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 5] [Cited by in F6Publishing: 6] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 20. | Yang Z, Shang J, Li N, Zhang L, Tang T, Tian G, Chen X. Development and validation of a 10-gene prognostic signature for acute myeloid leukaemia. J Cell Mol Med. 2020;24:4510-4523. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 9] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 21. | Santosa F, Symes WW. Linear Inversion of Band-Limited Reflection Seismograms. SIAM J Sci and Stat Comput. 1986;7:1307-1330. [DOI] [Cited in This Article: ] [Cited by in Crossref: 380] [Cited by in F6Publishing: 406] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 22. | Tibshirani R. Regression Shrinkage and Selection via the Lasso. Journal of the Royal Statistical Society Series B (Methodological). 1996;58:267-288. [DOI] [Cited in This Article: ] [Cited by in Crossref: 8558] [Cited by in F6Publishing: 8680] [Article Influence: 1446.7] [Reference Citation Analysis (0)] |
| 23. | Tibshirani R. The lasso method for variable selection in the Cox model. Stat Med. 1997;16:385-395. [PubMed] [DOI] [Cited in This Article: ] [Cited by in F6Publishing: 42] [Reference Citation Analysis (0)] |
| 24. | Liu Y, Wu WH, Hong S, Fang J, Zhang F, Liu GX, Seo J, Zhang WB. Lasso Proteins: Modular Design, Cellular Synthesis, and Topological Transformation. Angew Chem Int Ed Engl. 2020;Online ahead of print. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 10] [Cited by in F6Publishing: 13] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 25. | Zhou J, Zou Y, Hu G, Lin C, Guo Y, Gao K, Wu M. Facilitating colorectal cancer cell metastasis by targeted binding of long non-coding RNA ENSG00000231881 with miR-133b via VEGFC signaling pathway. Biochem Biophys Res Commun. 2019;509:1-7. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12] [Cited by in F6Publishing: 12] [Article Influence: 2.0] [Reference Citation Analysis (0)] |