Published online Apr 28, 2018. doi: 10.3748/wjg.v24.i16.1708
Peer-review started: March 27, 2018
First decision: April 3, 2018
Revised: April 10, 2018
Accepted: April 16, 2018
Article in press: April 16, 2018
Published online: April 28, 2018
The annual number of deaths caused by hepatitis B virus (HBV)-related disease, including cirrhosis and hepatocellular carcinoma (HCC), is estimated as 887000. The reported prevalence of HBV reverse transcriptase (RT) mutation prior to treatment is varied and the impact of preexisting mutations on the treatment of naïve patients remains controversial, and primarily depends on geographic factors, HBV genotypes, HBeAg serostatus, HBV viral loads, disease progression, intergenotypic recombination and co-infection with HIV. Different sensitivity of detection methodology used could also affect their prevalence results. Several genotype-dependent HBV RT positions that can affect the emergence of drug resistance have also been reported. Eight mutations in RT (rtL80I, rtD134N, rtN139K/T/H, rtY141F, rtM204I/V, rtF221Y, rtI224V, and rtM309K) are significantly associated with HCC progression. HBeAg-negative status, low viral load, and genotype C infection are significantly related to a higher frequency and prevalence of preexisting RT mutations. Preexisting mutations are most frequently found in the A-B interdomain of RT which overlaps with the HBsAg “a” determinant region, mutations of which can lead to simultaneous viral immune escape. In conclusion, the presence of baseline RT mutations can affect drug treatment outcomes and disease progression in HBV-infected populations via modulation of viral fitness and host-immune responses.
Core tip: The prevalence of preexisting reverse transcriptase (RT) mutations in treatment-naïve patients largely depends on geographic factors, HBV genotypes, HBeAg serostatus, hepatitis B virus (HBV) viral loads, disease progression, intergenotypic recombination, co-infection with HIV and the method used for detecting the mutation. Genotype-dependent polymorphic amino acid substitutions in RT may affect the emergence of drug resistance, and genotype C exhibits relatively elevated spontaneous RT mutation rates. HBeAg-negative status and low viral loads are significantly associated with a higher frequency and prevalence of HBV preexisting RT mutations. Preexisting mutations are most frequently found in the A-B interdomain of RT, mutations of which can lead to simultaneous viral immune escape.
- Citation: Choi YM, Lee SY, Kim BJ. Naturally occurring hepatitis B virus reverse transcriptase mutations related to potential antiviral drug resistance and liver disease progression. World J Gastroenterol 2018; 24(16): 1708-1724
- URL: https://www.wjgnet.com/1007-9327/full/v24/i16/1708.htm
- DOI: https://dx.doi.org/10.3748/wjg.v24.i16.1708
Although an effective and safe vaccine against hepatitis B virus (HBV) has been available since 1982[1], approximately 257 million people are chronic carriers of the virus. The annual number of deaths caused by HBV related diseases, including cirrhosis and hepatocellular carcinoma (HCC), was estimated as 887000 in 2015 (WHO, 2017)[2].
Reverse transcriptase (RT) conducts the major enzymatic activity required for viral replication. Nucleos(t)ide analogs (NAs) such as lamivudine[3], adefovir dipivoxil[4], entecavir[5], telbivudine[6], and tenofovir[7], for treatment of HBV infection, mainly target RT and function as reverse transcriptase inhibitors by mimicking natural nucleosides and integrating within the DNA molecules to interfere with viral replication[8,9]. However, due to the lack of proof reading ability of RT, the error rate for viral genome replication is as high as 10-7 per nucleotide, which is 10-fold greater than that of other DNA viruses[10], resulting in the emergence of antiviral-drug resistance mutations[11-15]. These NA-resistant (NAr) mutants are the greatest challenge for treatment of HBV because they change the conformational structure of RT and lower the effectiveness of NAs by impeding their binding[16]. In addition, RT partially overlaps with HBV surface antigen (HBsAg) and RT mutation may simultaneously generate HBsAg mutations, which can alter the antigenicity, immune recognition, replication capacity, and virulence of HBV[17-19].
The reported prevalence of preexisting HBV polymerase RT mutations is varied and the impact of preexisting RT mutations on treatment-naïve patients remains controversial. In addition, the relationship between preexisting RT mutations and advanced liver diseases, such as cirrhosis and HCC, has not been fully investigated[20]. Therefore, this review focuses primarily on factors affecting the prevalence and types of preexisting RT mutations in treatment-naïve patients and the relationship between these mutations and disease progression.
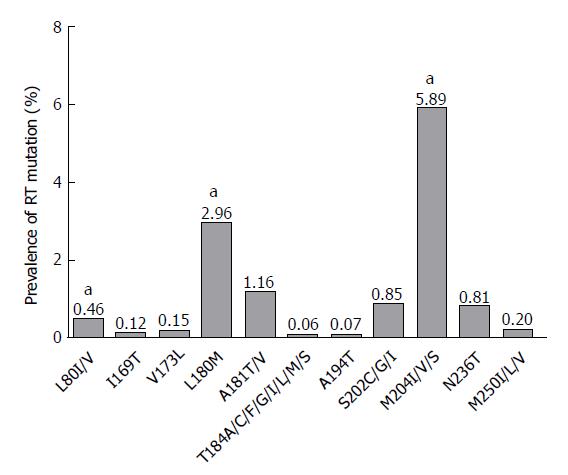
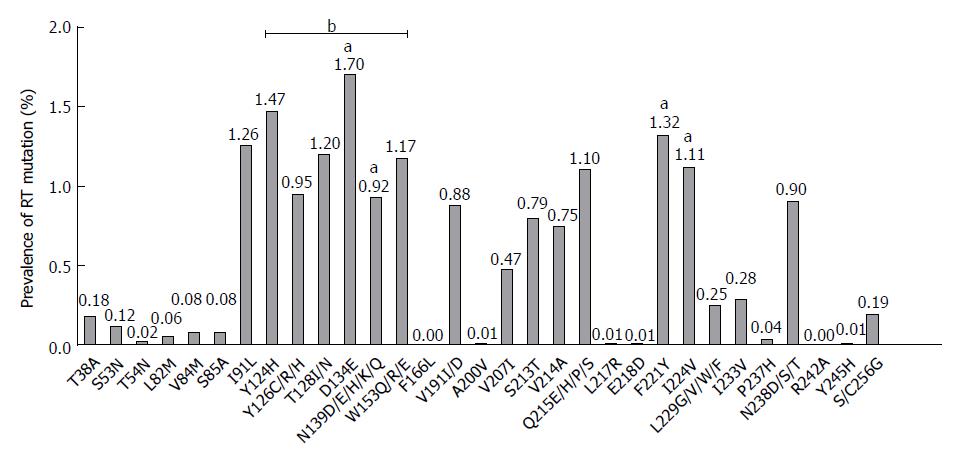
Liu et al[21] identified pre-existing HBV RT mutations in 42 potential NAr RT positions from 192 treatment-naïve Chinese patients and arranged them into following four mutation categories: primary drug resistance (Category 1); secondary/compensatory mutation (Category 2); putative NAr (Category 3); and pretreatment (Category 4) (Table 1). To understand the global prevalence of these 42 naturally occurring NAr resistance mutations of RT, we reviewed a total of 50 previous studies[12,20-68] and collated their results (Figure 1). These include 32 articles published from institutions based in Asia (12 published from China, four from Iran, four from Turkey, four from India, three from Japan, two from Taiwan, and one each from Korea, Jordan, and Indonesia), 11 articles published from institutions based in Europe (six from Italy, two from Germany, and one each from Austria, Ireland, and Spain), four articles published from institutions based in North America (three from United States and one from Canada), two articles published from institutions based in South America (both from Brazil), and one article published from an institution in South Africa (Supplementary Table 1). Among the 50 studies, 36[12,20,21,32-58,60-65] used direct PCR sequencing methods, 11[22-28,59,66-68] used the INNO-LiPA line assay, and 3[29-31] detected RT mutations by ultra-deep pyrosequencing (UDPS). Seventeen articles[21,26,27,30,31,34,35,37-39,42,43,46-48,53,58] included treatment-naïve patients infected with genotypes B and C, one study[32] with genotypes A and D, eleven studies[22,28,29,36,50,51,55,56,60,63,68] with genotype D, one study[33] with genotype C, and fifteen studies[20,23-25,40,41,44,45,52,54,57,61,64,65,67] with more than three genotypes (e.g., A, C, and D or A, B, C, and D). In five studies, genotypes of patients were not mentioned. Our literature-based study demonstrated that preexisting RT mutations were also found in treatment-naïve patients at 40 of 42 preciously identified NAr RT positions, the two exceptions were rtF242A, a pretreatment mutation and rtF166L, a lamivudine (LMV)-associated putative mutation. The distribution and overall incidence of RT mutations is presented in Figures 1 and 2.
| Mutation | RT mutation type | Change in HBsAg3 | Drug resistance | Genotype | Location | Ref. |
| Primary1 | I169T | ETV | B, C | China | [20,53] | |
| A181T/V | sW172 stop | LMV, LdT, ADV, TNF | A, B, C, D | Canada, Italy, China, United States | [24,25,30,39,45,51,53] | |
| T184A/C/F/G/I/L/M/S | ETV, LMV | A, B, C, D | Canada, China, South Korea | [24,27,33] | ||
| A194T | no change in HBsAg | ADV, TNF | B, C, D | Indonesia, Italy | [38,51] | |
| S202C/G/I | ETV, LMV | A, B, C, D | Canada, China, Spain, Italy | [24,27,30,57,59] | ||
| M204I/V/S | V-sI195M, I- sW196S/L/Stop | LMV, LdT, ADV, TNF, ETV | A, B, C, D | Canada, Italy, South Korea, China, Indonesia, United States, Turkey | [24,25,27,30,33-36,38, 39,47,52,53] | |
| N236T | ADV, TNF | B, C | China | [27,30,47] | ||
| M250I/L/V | ETV | A, B, C, D | Canada, Italy, China, United States | [24,25,27,45] | ||
| Secondary1 | L80I/V | no change in HBsAg | LMV | A, B, C, D | Canada, Italy, South Korea, Indonesia, China | [24,25,33,38,47,53] |
| V173L | sE164D | LMV | A, B, C, D | China, Canada, Italy | [20,24,25,30,39,47] | |
| L180M | no change in HBsAg | LMV, ETV, LdT | A, B, C, D | Canada, South Korea, China, Italy, Indonesia | [24,25,27,30,33,38, 39,47,53] | |
| Putative2 | S53N | LMV | B, C | China, South Korea | [21,33,58] | |
| T54N | sP46T | ADV | A, B, C, D | Italy | [83] | |
| L82M | LMV | C | South Korea | [33] | ||
| V84M, S85A | ADV | B, C | South Korea, China | [33,53,127] | ||
| I91L | no change in HBsAg | LMV | B, C, D | China, Indonesia, India | [21,38,54,58] | |
| Y126C/R/H | C-sT118A | ADV | A, B, C, D | South Korea, China, India | [33,54,58] | |
| T128I, T128N | N-sG145R, I-sP120S | LMV | B, C, D | Indonesia, China, India | [38,53,54,58] | |
| N139D/E/Q | A | India | [54] | |||
| W153Q/R/E | Q- sP120T/ sG145R, E-sD144E/G145R | LMV | B, C, D | South Korea, Indonesia, China, India | [33,38,54,58] | |
| F166L | sF158Y | LMV | ||||
| V191I | sW182 stop | LMV, ADV | B, C, D | South Korea, China, Italy | [33,39,51,58] | |
| A200V | sL192F | LMV | B, C | South Korea, China | [27,33,42] | |
| V207I | sW199 stop, sM198I | LMV | A, B, C, D | Germany, China, Italy | [27,32,39,83] | |
| S213T | LMV, ETV | A, B, C, D | China, India | [39,40,42,58] | ||
| V214A | T-sS204R | ADV | B, C, D | South Korea, Italy, China, | [30,33,53,83] | |
| Q215E/H/P/S | LMV, ADV | A, B, C, D | China, Italy, India, Turkey | [30,39,51,53-55] | ||
| L217R, E218D, F221Y | D-sS210I/T | ADV | A, B, C, D | South Korea, China | [21,33,42,58] | |
| L229G/V/W | LMV | B, C | South Korea, China | [33,42,58] | ||
| L229F | F-sC221L, V-sF220L | LMV | B, C, D | China, India | [54,58,128] | |
| I233V | ADV | A, C, D | Indonesia, Italy, India, China, South Korea, Germany | [25,38,40,58,78,79] | ||
| P237H, N238D/S/T, Y245H | N/A | ADV | B, C | South Korea, China | [21,33,39,58] | |
| S/C256G | LMV, ETV | B, C | China, Indonesia | [21,38,58] | ||
| Pretreatment2 | T38A, T38K | K-sQ30K | B, C | South Korea, China | [33,58] | |
| Y124H/D/N | B, C, | China, South Korea | [21,33,58] | |||
| D134E/N | sI126S/N | B, C, D | China, South Korea, Indonesia | [21,33,38,58] | ||
| N139K/H | K-sT131N, T-sT131P, H-sG139N | A, B, C | South Korea, India, Indonesia, China | [33,38,54,58] | ||
| I224V | No change in HBsAg | B, C, | China, South Korea | [21,33,39,58] | ||
| R242A |
Primary drug resistance mutations are amino acid changes that cause direct NA resistance by decreasing viral susceptibility to NAs[69-71]. Mutated RT positions known to induce primary drug resistance are rt169, rt181, rt184, rt194, rt202, rt204, rt236, and rt250. The mutations rtA181T/V, rtM204I, and rtM204V also cause the simultaneous HBsAg mutations, sW172 stop, sW196S/L/Stop, and sI195M, respectively[17,72] (Table 1). Our literature based pooled incidence data showed that of primary drug resistance mutations, M204I/V is the most frequently encountered in treatment-naïve patients[14,28,33] (5.89%), which was far more than the pooled mutation rate of rtA181T/V, rtS202C/G/I and rtN236T (incidence: 1.16%, 0.85% and 0.81%, respectively). Mutation of rtI169T (0.12%), rtT184G (0.06%), rtA194T (0.07%), and rtM250V/L (0.20%) had a very low pooled incidence (Figure 1). A systematic review by Zhang et al[73] revealed that the global incidence of rtM204I/V/S is 4.85%. Several other studies[24,28,34-37] have also reported the frequent incidence of rtM204I/V/S in treatment-naïve patients. For examples, Kobayashi et al[34], Lee et al[35], Tuncbilek et al[36], Fung et al[24], and Huang et al[37] reported rtM204I/V/S mutation frequencies in Japanese, Taiwanese, Turkish, Canadian, and Chinese treatment-naïve patients reached of 27.8%, 57%, 7.8%, 12%, and 26.9%, respectively.
Secondary, or compensatory, mutations refer to amino acid substitutions that compensate for replication defects caused by primary drug resistance mutations and may reduce drug susceptibility by restoring viral replication fitness[38,69,74]. The mutations rtL80I/V, rtV173L, rtL180M are known for secondary resistance mutations. Our literature based incidence data showed that rtL180M had the highest natural incidence (2.96%), which was higher than the pooled mutation rate of rtL80I/V and V173L (incidence:0.46% and 0.15%, respectively) (Figure 1). Similarly, Zhang et al[73] reported that the overall frequency of rtL180M mutation is 2.67%. Other studies, including Fung et al[24], Yamani et al[38], and Mirandola et al[25] reported that the prevalence rates of rtL180M were 10.0%, 2.08%, and 1.18% in Chinese, Indonesian, and Italian HBV carriers, respectively. The rtL80I/V mutation also occurs frequently in treatment-naïve patients. Yamani et al[38], Kim et al[33], and Mirandola et al[25] found pre-existing rtL80I/V mutation frequencies of 1.07%, 3.82%, and 0.78%, respectively. Notably, Kim et al[33] reported that rtL80I/V was the most frequently encountered preexisting mutation of secondary drug resistance mutations in South Korea (3.8%, 5/131 patients), even higher than rtL180M frequency (2.3%, 3/131 patients). Another compensatory RT mutation, rtV173L, was also detected in several studies of treatment-naïve patients, where Zheng et al[20], Wang et al[39], and Mirandola et al[25] reported that it occurred in 0.6%, 0.56%, and 0.39% of their patients, respectively.
RT mutations which have been identified as associated with drug resistance, but have not been confirmed experimentally in vitro, are defined as putative NA resistant mutations[75-77]. A total of 26 types of RT mutations, including rtS53N, rtT54N, rtL82M, rtV84M, rtS85A, rtI91L, rtY126C, rtT128I/N, rtN139D, rtW153Q, rtF166L, rtV191I, rtA200V, rtV207I, rtS213T, rtV214A, rtQ215P/S, rtL217R, rtE218D, rtF221Y, rtL229G/V/W, rtI233V, rtP237H, rtN238D/S/T, rtY245H, and rtS/C256G, are considered putative drug resistance mutations (Category 3) (Table 1)[21]. Other amino acid substitutions, which were detected before treatment, but for which the association between their occurrence and drug resistance has not been evaluated, are defined as pretreatment mutations, these include rtT38A, rtY124H, rtD134E/N, rtN139K/H, rtI224V, and rtR242A (Category 4) (Table 1)[21,38].
Recently, it has been proven through in vitro and in vivo experiments that several putative or pretreatment mutations, including rtL229F, rtS13T, and rtI233V, can also contribute to the development of drug resistance[40,78,79]. In addition, several studies have reported that treatment-naïve patients with only putative RT mutations, and without primary or secondary changes, developed drug resistance since treatment initiation[41]. Our literature based pooled incidence data showed that several putative or pretreatment mutations, including rtI91L, mutations in 6 positions of A-B interdomain (rtY124H, rtY126C/R/H, rtT128I/N, rtD134E/N, rtN139D/E/H/K/Q and rtW153Q/R/E), rtF221Y and rtI224V, were encountered with high frequency from the treatment naïve patients[20,21,33,38,39,42]. Of these, a RT mutation in the A-B interdomain, rtD134E/N, which also cause a simultaneous sI126N/S mutation of the HBsAg “a” determinant, was found to have the highest frequency in treatment-naïve patients (1.70%) (Figure 2). Of note, the following four putative or pretreatment mutations found in treatment naïve patients, rtD134E/N, rtN139D/E/H/K/Q, rtF221Y, and rtI224V, are also reported as associated with progression of severe liver diseases, such as HCC and cirrhosis (described below). Moreover, some of these mutations also overlapped with genotype-dependent polymorphic sites, as described in the next section.
To date, a total of 10 HBV genotypes (A-J) and several sub-genotypes have been identified; genotypes are separated from each other by sequence differences of more than 8% by phylogenetic analysis, based on whole genome sequences[80,81]. HBV genotypes, including genotypes A-J and the various sub-genotypes, are associated with several distinct traits, including geographical distribution, host ethnicity, and pathogenicity[82]. Since specific mutational patterns of mutation can be restricted by structural/functional constraints to particular genotypes, HBV genotype can influence the evolution frequency, or types, of mutations associated with NAr in treatment-naïve patients, as described by Liu et al[21]. Moreover, HBV genotypes can affect LAM-resistant mutations in the YMDD motif of viral RT in patients with chronic infections after long term drug treatment[42,43,83]. Two recent studies[83,84] reported that genotype A favors the rtM204V mutation, while HBV-B, C, and D select for rtM204I at higher rates, compared with rtM204V. Moreover, Liu et al[21] identified eight genotype-dependent AA polymorphic positions, (i.e., rt53, rt91, rt124, rt134, rt221, rt224, rt238, and rt256) useful as signature for B- and C-genotypes with mutations at the 42 positions associated with NAr, and which are important for the distinction between mutations and mere polymorphisms during genotypic mutation analysis of samples from infected patients. These specific, genotype-dependent AA polymorphic positions in RT functional domains, (i.e., rt91 in domain A, rt238 in domain D and rt256 in domain E) affect drug treatment outcomes[21,85]. Liu et al[21] also showed that rtL91 and rtI91 were generally favored by genotypes B and C, respectively, and that rtS256 was prevalent in both genotypes, with rtC256 more common in genotype C than in genotype B. rtL91 and rtC256 were also more closely correlated with failure of extended LMV therapy, compared with other polymorphisms (rtI91and rtS256) leading to the suggestion that they may represent potential pretreatment markers[86].
Mirandola et al[83] further extended the eight genotype dependent polymorphic-sites suggested by Liu et al[21] into a total of 27 polymorphic-sites potentially affecting treatment, via sequence analysis of 200 treatment-naïve chronically infected patients from north-east Italy, infected with the six HBV genotypes; A, B, C, D, E, and F. In this study, substitutions at residues rt53, rt54, and rt91 of the 27 genotype-dependent polymorphic sites were the most frequent single AA substitutions, and HBV-DNA levels of patients with these mutations were significantly lower than those of patients with no mutations, suggesting that these changes contributed to reducing viral fitness during infections. The authors also found that patients with multiple mutations were mainly infected with genotype D, rather than other genotypes (A, B, C, E, or F), strongly supporting previous results indicating that HBV-D may have the highest genetic variability among all HBV-genotypes[87].
Yamani et al[38] demonstrated that the distribution of primary and secondary drug resistance mutations was not significantly different between genotypes B and C; however, significant differences were identified in some genotype-dependent polymorphic sites. For example, rtL91I and rtY221F were more common in genotype C, compared to genotype B (P < 0.001) (Table 2). Moreover, some genotype dependent mutations, such as rtM129L, rtD134N, rtM145L, and rtE263H/D/Q, were more frequent in in genotype C than genotype B viruses (P < 0.001). Notably, rtN226H/T was the only pretreatment mutation, which is more common in genotype B than genotype C (P < 0.001). Singla et al[44] also showed that rtL91I and rtM129L are more common in samples from genotype C, than genotype D, infected patients. Overall, these findings indicate that distribution of genotype dependent polymorphic sites in treatment-naive patients could affect drug treatment outcomes via modulation of viral fitness or replication. The distribution of the 29 genotype-dependent polymorphic-sites in the HBV RT region among treatment naïve patients identified in other reports is summarized in Table 2.
| RT position | Drug resistance | Mutations in RT region of four genotypes1 | Polymorphism | Ref. | |||
| A | B | C | D | ||||
| 38 | T (4.4)4 | T (14.0)4 | A/T/T/A | [83] | |||
| 53 | LMV | D/T (1.8)4 | I/N/S/N | [21] | |||
| 54 | ADV | N (2.2)2 | T/T/T/H | [83] | |||
| 84 | I (0.5)2 | V | [127] | ||||
| I (1.5)2 | [21] | ||||||
| 85 | T (0.6)2 | S | [127] | ||||
| 91 | LMV | I (20.0)4 | L (23.5)2 | I (16.7)4 | I/L/I/L | [83] | |
| I (14.3)2 | [44] | ||||||
| 103 | I (100)3 | I (1.67)3 | V | [40] | |||
| 122 | H (47.0)2 | H (6.66)2 | F | [40] | |||
| L/V/I(25.0)2 | [44] | ||||||
| 124 | H (2.2)3 | H (20.0)3 | H (11.8)3 | N/N/Y/H | [83] | ||
| H (3.6)3 | H (6.6)3 | [21] | |||||
| D (5.5)3 | |||||||
| 126 | H (6.7)4 | R (23.7)4 | Y/H/H/H | [83] | |||
| R (17.9)4 | [44] | ||||||
| Y (1.4)4 Q (0.5)4 | [83] | ||||||
| 128 | LMV | N (2.2)2 | N (1.4)2 I (1.4)2 | T | [83] | ||
| I (1.9)2 | [38] | ||||||
| 129 | L (60.0)2 | L (21.4)2 | M | [44] | |||
| L (1.9)2 | L (26.2)2 | [38] | |||||
| L (100.0)2 | L (9.0)2 | L (3.3)2 | [40] | ||||
| 134 | N (40.5)2 | D/N/D/D | [38,54] | ||||
| E (23.1)3 | E (5.0)3 | [21,54] | |||||
| I/S (1.8)3 | E (5.8)3 | [21] | |||||
| 139 | LMV | K (3.7)2 | K (11.9)2 | Q/N/N/N | [38] | ||
| K (2.3)3 | [83] | ||||||
| K (1.5)3 | [21] | ||||||
| 145 | L (3.7)2 | L (40.5)2 | M | [83] | |||
| 153 | LMV | Q (100)2 | W/R/R/R | [40] | |||
| 191 | LMV | V (8.3)2 | V/I/I/V | [39] | |||
| I (4.6)2 | F (7.7)2 | [54] | |||||
| 200 | LMV | V (2.2)2 | A | [83] | |||
| 207 | LMV | M (6.0)2 L (2.3)2 | V | [26] | |||
| I (5.9)2 | [83] | ||||||
| I/L/G (2.1)2 | [39] | ||||||
| 214 | LMV/ADV | A (5.9)2 | A (2.3)2 | V | [83] | ||
| A (0.5)2 I (0.5)2 | I (0.6)2 | A (0.8)2 E (0.7)2 | [127] | ||||
| 215 | ADV | E (7.7)2 | H (5.0)2 S (5.0)2 | Q | [54] | ||
| H (0.9)2 | H (3.0)2 S (4.3)2 | [127] | |||||
| P (2.8)2 S (4.2)2 | [83] | ||||||
| 217 | L (6.7)4 | R (0.9)2 | R/L/L/L | [83] | |||
| 221 | ADV | F (40.5)2 | Y/Y//F/F | [38] | |||
| Y (5.1)2 | [83] | ||||||
| 226 | H/T (33.3)2 | H/T (2.4)2 | N | [38] | |||
| 237 | T (6.4)2 | P | [127] | ||||
| 238 | LMV, ETV | H(1.0)2 W(1.0)2 | Q (3.9)2 | T (8.7)2 | H (2.3)2 | N/H/N/N | [127] |
| Q (1.82)2 | S (2.19)2 T (0.73)2 | [21] | |||||
| T (3.87)2 | [39] | ||||||
| D (2.2)2 T (2.2)2 | D (1.4)2 | [83] | |||||
| 245 | H (1.4)2 | Y | [83] | ||||
| 248 | ADV | H (7.4)3 | H (4.8)3 | N | [38] | ||
| 256 | LMV | G (40.0)2 | G (10.7)2 | S/S/S/C | [44] | ||
| G (20.0)2 | G (3.7)2 | [83] | |||||
| G (5.45)2 | [21] | ||||||
| G (3.7)2 | [26] | ||||||
Mirandola et al[25] identified the different genotype different distributions of antiviral drug resistant RT mutations using INNO LiPA line probe analysis of samples from treatment-naïve patients; RT mutations were detected in 13 (5%) of 255 HBV infected patients. Of these, 10 patients had mutations associated with primary resistance or reduced sensitivity, including three cases with a YMDD mutation (rtM204V), three with the mutation, rtM250L/V, which is associated with ETV resistance, and four with the mutation rtI233V, which is associated with reduced sensitivity to ADV. Notably all the three patients with the rtM204V mutation also had coexisting L180M compensatory mutations, and all were infected with HBV-C genotype viruses, suggesting that naturally occurring LMV-resistant HBV may be more frequent in patients infected with genotype C virus. This hypothesis is strongly supported by the recent report of Kim et al[33] of the high frequency of the YMDD mutation, (rtM204V/I) (6.87%, 9/131 patients), in Korean treatment-naïve patients with HBV genotype C2 infections. Wang et al[39] also reported that RT mutations were only found in genotype C treatment-naïve patients; however, no primary or secondary RT mutations were found in genotype B patients. In addition, a systemic meta-analysis review by Zhang et al[73] showed that rtM204V/I had the highest incidence of 4.89% (95%CI: 4.13%-5.65%) among primary and secondary RT mutations. These authors also found, via the subgroup analysis by genotype, that HBV genotype C had a tendency of toward a higher spontaneous YMDD mutation frequency (19.32%) than genotype B (15.01%) or D (14.79%). The increased spontaneous mutations in the viral genome of HBV genotype C could translate to a higher risk of primary NA resistance in HBV endemic areas, where genotype C infections are prevalent, including China and South Korea.
The majority of studies have consistently reported a significant association between the prevalence of preexisting RT mutations and lower HBV DNA loads, or HBeAg-negative status, in treatment-naïve patients[26,27,34-36,45-47,88] (Table 3). Vutien et al[26] reported that treatment-naïve patients with HBeAg-negative status had higher RT mutation frequencies (78.57%), compared with HBeAg-positive patients (21.42%). These authors also showed that HBeAg-negative patients had significantly lower HBV DNA viral loads compared with HBeAg-positive patients (5.65 log10 IU/mL vs 7.82 log10 IU/mL, respectively). Zhao et al[27] also reported similar results showing that 75% of patients with RT mutations were HBeAg-negative and had lower HBV DNA levels (3.92 log10 IU/mL) whereas 25% of patients with RT mutations were HBeAg-positive with higher HBV DNA loads (5.54 log10 IU/mL). Similarly, Zhu et al[89] found that Chinese patients with chronic HBV carrying preexisting RT mutations had significantly decreased serum baseline HBV DNA loads (P = 0.0363) and blood platelet counts (P = 0.0181) compared with those without RT mutations.
| HBeAg-positive | HBeAg-negative | HBV genotype (%) | Location | Ref. | ||
| Mutations1 | HBV-DNA2 | Mutations1 | HBV-DNA2 | |||
| 3/14 (21.4) | 7.8 | 11/14 (78.6) | 5.7 | B, C, E | California | [26] |
| 6/24 (25.0) | 5.5 | 18/24 (75.0) | 3.9 | B, C, B-C | China | [27] |
| 0/4 (0.0) | 7.2 | 4/4 (100.0) | 4.7 | A, B, C, D, F | California | [45] |
| 3/6 (50.0) | 8.0 | 3/6 (50.0) | 3.2 | D | Turkey | [36] |
| 8/12 (67.0) | 7.9 | 4/12 (33.0) | 6.9 | NA | Taiwan, China | [35] |
| 27/43 (62.8) | 5.7 | 16/43 (37.2) | 4.7 | B, C | China | [46] |
| 8/13 (61.5) | 6.3 | 5/13 (38.5) | 5.4 | B, C | China | [47] |
| 0/5 (0.0) | NA | 5/5 (100.0) | NA | NA | Japan | [34] |
| 0/4 (0.0) | NA | 4/4 (100.0) | NA | NA | Japan | [88] |
Several other studies[34,45,88] also found RT mutations only in HBeAg-negative patients, and the patients were also more likely to have decreased HBV DNA levels compared with those who were HBeAg-positive[45]. Kobayashi et al[34] reported that all asymptomatic HBV carriers with YMDD mutation were HBeAg-negative and eAb-positive, suggesting that sustained host immune pressure may be a major force driving potential NAr mutations. Zhang et al[73] also reported a systemic meta-analysis finding that patients with chronic hepatitis B (CHB) and genotype C infections, who were male and HBeAg-negative tended to have higher spontaneous mutation rates in subgroup analysis. Xu et al[46] reported no significant correlation between pre-existing mutations and the majority of clinical factors including gender, age, HBV genotype, ALT, HBeAg, and HBV DNA loads, in a Chinese population; however, subgroup analysis indicated that pre-existing mutations were strongly associated with lower HBV DNA levels in HBeAg sero-negative, but not HBeAg sero-positive, patients (HBeAg+ vs HBeAg-: 5.74 log10 IU/mL vs 4.72 log10 IU/mL, P = 0.0112). These findings suggest that preexisting RT mutations might lead to lower HBV viral loads in treatment-naïve patients with HBeAg-negative serostatus. Several other studies have reported similar positive associations between the frequency of pre-existing RT mutations and decreased HBV viral loads[33,42,83].
Taken together, there appears to be a clear causal link between preexisting RT mutations and HBeAg-negative status, decreased HBV DNA load, or liver disease progression. This may be because mutations in the RT active domain, could impair enzyme activity, particularly at the HBeAg negative immune clearance stage, thus decreasing the efficacy of virus replication and, resulting in liver disease progression and poor treatment outcomes[17,42,90,91].
Reports of the incidence of preexisting RT mutations in treatment-naïve patients are highly variable, ranging from 0% to 57%[25,26,28,32-35,48,92,93]. This huge discrepancy among studies may be due to differences in factors such as the geographical or ethnic backgrounds of studied patients, sample size, and viral genotype[27]. A number of studies have reported prevalence rate of preexisting RT mutations (primary and secondary RT mutations) of more than 5% in treatment-naïve patients (Table 4). Fung et al[24] found a higher rate of baseline RT mutations (12% M204I/V, 10% L180M) by using the INNO-LIPA v.3 assay. In this study, many patients, most of whom were infected with genotype D, carried rtL180M, rtM204V/I, and rtL80V/I mutations. In addition, Nishijima et al[94] identified a high mutation rate (35.7%) in 14 treatment-naive patients in Japan, using UDPS. Also, a recent study using direct sequencing[33] of samples from 131 treatment-naïve patients infected with genotype C2 reported an overall rate of 12.98% for primary (rtT184A/C/F and rtM204I/V) or compensatory (rtL80I and rtL180M) mutations. According to a systemic meta-analysis review conducted by Zhang et al[73], the overall prevalence of spontaneous mutations among treatment-naïve patients worldwide was 5.73%. The highest pooled prevalence (8.00%) was identified in samples from China, followed by Japan, Turkey, Korea, South America, and Europe at 6.62%, 6.43%, 5.72%, 3.89%, and 2.53%, respectively. Another study of 325 genotype D infected treatment-naïve patients using direct PCR sequencing[50] reported overall incidence of 15.69% for primary and secondary drug resistance mutations, including L80V/I, L180M, M204I/V, and S213T/N.
| Prevalence | Location | No. of cases | Genotype | HBV DNA loads (log10 IU/mL) | RT mutations prevalence | Mutation detecting methods | Ref. |
| HBV DNA RT mutations ≥ 5% | Italy | 255 | A, C, D | 5.0 | 5.0% mutations overall | INNO-Lipa HBV DR v.3 | [25] |
| China | 269 | B, C, B-C | 4.9 | 8.9% mutations overall | INNO-Lipa HBV DR v.3 | [27] | |
| Canada | 209 | A, B, C, D | 7.0 | 12% M204I/V, 10% L180M, 9% L80V/I, 3%V173L | INNO-Lipa HBV DR v.3 | [24] | |
| Turkey | 71 | NA | NA | 18.3% YMDD mutations | INNO-Lipa HBV DR v.1 | [28] | |
| South Korea | 131 | C2 | 6.5 | 12.98% mutations overall | Direct Sequencing | [33] | |
| Turkey | 77 | D | 7.3 | 7.8% YMDD mutations | Direct Sequencing | [36] | |
| China | 213 | B, C | 6.2 | 6.1% mutations overall | Direct Sequencing | [47] | |
| China | 104 | B, C, B-C | 4.5 | 26.9% YMDD mutations | Direct Sequencing | [93] | |
| Japan | 18 | NA | NA | 27.8% YMDD mutations | Direct Sequencing | [34] | |
| Iran | 325 | D | NA | 15.69% mutations overall | Direct Sequencing | [50] | |
| Taiwan, China | 28 | NA | 7.5 | 57% YMDD mutations | Direct Sequencing | [35] | |
| China | 357 | B, C | 6.3 | 16.8% mutations overall | Direct Sequencing | [39] | |
| Meta-analysis (China) | 8156 | B, C, D | NA | 8.00% mutations overall | Record screening | [73] | |
| Japan | 14 | B, C | 4.9 | 35.7% YMDD mutations | Ultra-deep sequencing | [94] | |
| HBV DNA RT mutations < 5% | Iran | 147 | D | NA | None | Direct sequencing | [56] |
| China | 328 | B, C | 6.9 | 3.6% mutations overall | Direct sequencing | [53] | |
| Japan | 20 | NA | NA | None | Direct sequencing | [48] | |
| California | 472 | A, B, C, D, F | 5.3 | < 1% mutations overall | Direct sequencing | [45] | |
| Italy | 100 | NA | NA | None | Direct sequencing | [49] | |
| Italy | 140 | D | 4.0 | 3.5% mutations overall | Direct sequencing | [51] | |
| Germany | 96 | A, D | NA | None | Direct sequencing | [32] | |
| Brazil | 189 | A, C, D, F | 3.2 | overall 6.0% in Northeast/ 0% in North | Direct sequencing | [41] | |
| California | 198 | B, C | 4.2 | 1% mutations in polymerase | INNO-Lipa HBV DR v.3 | [26] |
In contrast, several studies have reported prevalence rates of less than 5 % for pre-existing RT mutations (primary and secondary RT mutations) in treatment-naïve patients (Table 4). For example, using direct sequencing of samples from treatment-naïve patients from the United States, Nguyen et al[45] demonstrated that only four (0.9%) of 472 patients were infected with viruses with primary and secondary mutations (rtA181A/S, rtA194S, and rtM250I). Similarly, Zollner et al[32] screened a total of 96 patients infected with HBV genotypes A and D (52.08% and 47.92%, respectively) using a direct sequencing assay, but found no primary or secondary resistance mutations. Another study by Salpini et al[51] using the direct sequencing method reported that, of 140 treatment-naïve patients infected with genotype D, only 1.4% had primary drug resistance mutations, while 2.1% carried secondary mutations.
Overall, preexisting RT mutation prevalence clearly reflects the geographical distribution of HBV infection. For example, China is an area with high levels of endemic area of HBV infection (8%, according to a national survey in 2006) and also has higher prevalence of pre-existing RT mutations[73]. Meanwhile, in Europe, which has low levels of endemic HBV infection (approximately 2%), there is a low incidence of spontaneous mutations (2.53%)[73]. Since the HBV geographic distribution has also a close relationship with the genotype distribution, the majority of countries in Asia with prevalent genotype B and C infections have high rates of spontaneous RT mutation (≥ 5%)[27,33-35,39,47,73], whereas countries in Europe, where genotype A and D infections are dominant, tended to have low incidences (≤ 5%)[32,49,51].
HBV intergenotypic recombination between different genotypes is regarded as an important strategy for HBV genetic diversity and may impose challenges on vaccine designation and antiviral therapy strategies[95,96]. In particular, the high prevalence of vertical infections in HBV endemic areas, such as Asia or Africa, could lead to a life-long chronic infection[97], resultantly leading to a high probability of co-infection and a high risk for virus recombination[96,98-101]. Previous studies on HBV recombination have identified different types of intergenotypic recombinants in HBV RT, most of which have recombination in RT/S overlapping region[96,98-103]. Of note, a recent study conducted by Liu et al[100] demonstrated that, through full-length HBV RT sequences analysis from 201 Chinese chronic hepatitis B (CHB) patients, 38.10% (24/63) infected with genotype B had recombination with genotype C in the 3′-terminal RT sequences. These authors also showed that these intergenotypic recombinants were associated with enhanced viral DNA load and higher RT point mutation rates, compared with their parental genotype B or C, highlighting the importance of monitoring intergenotypic RT recombinants in HBV endemic areas to ensure optimal management.
Approximately, 10% of HIV-infected persons worldwide are chronically infected with HBV, and co-infection of two viruses is most frequently identified (up to 25%) in sub-Saharan Africa and Asia[104]. HBV and HIV co-infection is a major cause of morbidity and mortality because it could contribute to an increased risk of liver cirrhosis and HCC[105]. In general, previous studies showed a predominance of HBV genotype A in HIV infected individuals, compared with other genotypes[106,107]. In particular, Makondo et al[108] reported that the ratio of genotype A to non-A (97% to 3%) was higher in the HBV/HIV co-infected Southern Africa patients compared with mono-infected individuals. These authors also showed that 10 percent, 3 out of 29 patients prior to the initiation of antiretroviral therapy (ART), had drug resistance mutations rtV173L, rtL180M+rtM204V, and rtV214A. In South Africa, rtM204I has been mainly detected in treatment-naïve HBV/HIV co-infected individuals[12] with rtM204V in treated HBV mono-infected participants[109], suggesting HIV co-infection could affect HBV preexisting RT mutation pattern. A study of South African patients conducted by Selabe et al[12] demonstrated that HBV lamivudine‐resistant strains were detected in three out of 15 treatment-naïve mono‐infected chronic hepatitis B patients, whereas detected in 10 out of 20 treatment-naïve HBV/HIV-coinfected patients. In contrast, a multi-national study of HIV/HBV-coinfected individuals carried out by Thio et al[110] demonstrated that no subject had preexisting RT mutations in the majority population of the quasispecies, suggesting no need for HBV drug-resistance testing prior to starting anti-HBV therapy in HIV-HBV co-infected individuals. It is also supported by a recent study of Ghana patients conducted by Archampong et al[111]. Taken together, geographical factors and HBV genotypes could have effects on the preexisting HBV RT mutation in treatment-naïve HBV/HIV-coinfected patients.
The detection methods used can also have a profound effect on the reported incidence results of preexisting RT mutations. The majority of studies have used direct sequencing methods, which can lead to the underestimation of preexisting RT mutations, due to the relative low sensitivity of these assays. Wang et al[39] reported that the sensitivity of direct sequencing-based protocols declined when circulating viral subspecies (AA substitutions) levels were at ratio below 20%-25%. Similarly, there were several studies have reported discordance in the incidence of pre-existing RT mutations detected by direct sequencing and other screening methods, such as the INNO-LiPA assay, or UDPS. For example, Margeridon-Thermet et al[52] reported that direct sequencing found an average of 5.9 mutations per sample, while UDPS identified an additional 4.6 mutations per sample, which could not be detected by direct sequencing. In that study, two of 17 treatment-naïve patients had mutations which were detected only by UDPS, but not by direct sequencing; one rtM204I mutation with (1.3% mutant ratio) and the other an rtA181T mutation (1.0% ratio). Similarly, Aberle et al[66] also compared the detection efficacies for preexisting RT mutations between the INNO-LiPA assay and direct sequencing. The former identified additional mutations in 8 (14%) of 56 patient samples, which could not be detected using the latter method, indicating the superiority of the former over the latter for RT mutation detection. Overall, these data demonstrate that the method used for detecting the mutations can affect the prevalence estimates of preexisting RT mutations in treatment-naïve patients, which may cause discrepancies among the results of different studies.
Although the clear association between preexisting RT mutations and advanced liver disease has not been fully investigated, several types of HBV mutations in RT have previously been reported as related to the progression of liver diseases, such as cirrhosis and HCC (Table 5). Kim et al[33] compared types and frequencies of pre-existing RT mutations between CHB and HCC treatment-naïve patients. These authors found a significantly higher rate of RT mutations in HCC patients than in those with chronic hepatitis (3.17% vs 2.09%, P = 0.003) and also identified a total of three NAr mutations (rtL80I, rtN139K/T/H, and rtM204I/V) significantly associated with HCC progression. RT mutations rtN139K/T/H and rtM204I/V also cause simultaneous mutations in the overlapped HBsAg coding sequence (sT131N/P and sI195M, or sW196S/L/Stop)[17,21,72]. Of these, the YMDD-motif mutation (rtM204I/V) was found in 9 patients of 131 patients (8 HCC and 1 CHB) with the other two types of mutation, rt204I and rt204V, in 8 and 1 patients, respectively. The other HCC-related mutation (rtL80I) was first identified as a compensatory mutation associated with LMV resistance[69,112]. Its relationship with clinical deterioration is also corroborated by other reports that it was associated with increased viral loads, accompanied by an elevation in serum aminotransferase activity, and exacerbation of liver disease in every case[113]. Interestingly, Kim et al[33] also showed that rtL80I was combined with the rtM204I/V mutation in five of nine rtM204I/V cases, and that patients with L80I had increased HBV replication compared with those without this mutation, suggesting that, together with rtM204I/V, it may contribute to HCC generation in treatment-naïve patients by compensating for the defective replication of caused by rtM204I/V.
| Type of mutation in RT | Chang in HBsAg | Genotype | Location | Disease progression | P value | Ref. |
| rtL80I2 | NC | C | South Korea | HCC | 0.036 | [33] |
| rtD134N4 | sI126S/N | B, C | China | HCC | 0.007 | [114] |
| rtN139K/T/H4 | sT131N/P | C | South Korea | HCC | 0.008 | [33] |
| rtY141F5 | sM307T | Ce | Taiwan | HCC | 0.029 | [37] |
| rtM204I/V1 | sW196L/S/W | C | South Korea | HCC | 0.021 | [33] |
| rtF221Y3 | NA | B,C,D | China | HCC, poor survival rate | 0.028/0.004 | [20,115] |
| rtI224V4 | NC | C | China | HCC | 0.005 | [116] |
| rtM309K5 | NA | C | China | HCC | 0.007 | [116] |
In another study, Yin et al[114] analyzed the association of the mutations of HBV polymerase with postoperative survival in 92 patients with HBV-related HCC using direct sequencing. They discovered three nucleotide sites, one (31st nucleotide) in a spacer region and two [529th (P = 0.007) and 1078th (P = 0.038)] in the RT region, which could be considered independent predictors of postoperative survival in HBV-related HCC. Of the two sites in RT related to HCC outcomes, rtD134N (mutation G529A) was associated with lamivudine resistance, further supporting previous findings of potential correlation between resistance to the anti-HBV nucleoside analog, lamivudine, and HCC prognosis[114]. Since rtD134N also causes an amino acid change in HBsAg (sI126N/V), it can induce changes in the antigenic properties of HBsAg. Further functional studies are necessary to determine whether the rtD134N mutation can induce HCC via modulation of RT activity or through its effects on HBV replication.
Huang et al[37] found seven viral single nucleotide polymorphisms (SNPs) in HBV polymerase, which enhance viral replication and liver disease progression in HBeAg negative subjects. Of these SNPs, rtY141F (Y487F), which is located in the RT region of HBV polymerase was associated with increased viral load and HCC (P = 0.0291). Moreover, rtY141F, a genotype C-related SNP, also led to a simultaneous amino acid change in the overlapping ’a’ determinant region of HBsAg (sM307T). In addition, Li et al[115] and Zheng et al[20] reported that the rtF221Y mutation was strongly related to HCC prognosis after liver resection (hazard ratio, 2.345; P = 0.001). Moreover, the rtF221Y mutation was also associated with poor overall survival (hazard ratio, 2.557; P = 0.004), suggesting that it is a potential independent risk factor and viral marker for HCC. Those results were consistent with the report of Li et al[115], which identified the rtF221Y mutant as an independent risk factor for recurrence of HCC and poor overall survival (P = 0.001 and P = 0.004, respectively). Wu et al[116] also investigated preexisting RT mutations potentially related to HCC in Chinese patients and identified rtI224V and rtM309K as significant risk factors for HCC (P = 0.005 and P = 0.007, respectively).
In addition, the number of RT mutations is associated with the liver disease progression. Zhu et al[89] revealed that patients with multiple RT mutant sites showed a significantly higher rate of liver fibrosis (P = 0.0128), suggesting a link between viral mutation and clinical progression of chronic hepatitis, and also highlighting that the natural accumulation of RT mutations is a process involved in viral survival during chronic liver fibrosis.
Overall, eight mutations in the RT region, namely rtL80I, rtD134N, rtN139K/T/H, rtY141F, rtM204I/V, rtF221Y, rtI224V, and rtM309K, are significantly related to liver disease progression. The majority of HCC-related RT mutations were reported from studies of treatment-naïve patients infected with genotype C HBV. This supports previous reports that HBV genotype C is more likely to lead to severe and aggressive liver disease than other HBV genotypes[112,117-122]. Of note, association of the following three mutations, rtM204V, rtL80I, and rtD134N, with disease progression provides a likely explanation for the positive relationship between lamivudine resistance and liver disease progression.
HBV RT consists of seven functional domains (G, F, A, B, C, D, and E) and five inter-domains (F-A, A-B, B-C, C-D, and D-E) which link the functional domains[18,31,123]. Previous studies[20,21,33,38] reported a higher frequency of preexisting RT mutations in the A-B inter-domain, compared with other regions.
Liu et al[21] revealed that all six sites in the A-B interdomain, rt124, rt126, rt128, rt134, rt139, and rt153, exhibit mutations (6/6, prevalence 100%), indicating high genetic variability of this region compared with other sites within RT domains (sites with mutations: 6/22, 27.27%; P = 0.0014). In this study, the mutation frequency of the A-B inter-domain (44/1152, 3.82%) was also significantly higher than those in other RT domains (Table 6). This result is in line with that reported by Zheng et al[20], who demonstrated that A-B interdomain exhibits higher mutation frequencies (4.3%, 5.3%, 3.6%) than those of other RT domains (1.4%, 1.4%, 1.3%) in Chinese treatment-naive patients with CHB, cirrhosis, and HCC. Specifically, they found that there was a clear tendency toward frequent mutations of the A-B interdomain in patients with cirrhosis suggesting a relationship between mutations in the A-B inter-domain and the development of this condition.
Similarly, Yamani et al[38] also reported that the A-B interdomain had the highest mutation prevalence and frequency (3.51% ± 2.53%) compared with functional domains and non-A-B interdomains (1.45% ± 1.05% and 2.58% ± 0.51%, respectively) in Indonesian treatment-naïve patients (Table 6). Moreover, they found that genotype C had substantially higher mutations rates in the A-B interdomain than genotype B (P < 0.001). Kim et al[33] also revealed that mutations within the A-B interdomain were most frequent in treatment-naïve Korean patients infected with genotype C2, compared with other domains, with 46 of 79 patients (58.22%) with preexisting RT mutations having changes in the A-B inter-domain. In this study, rtD134E/N/C was the most frequently encountered hot spot site among the six A-B inter-domain sites and was mutated in 12/79 patients (15.2%). The authors also showed that the mutation frequency of A-B interdomain (59/786, 7.50%) was higher than that of non A-B interdomain (3.16%) (Table 6). Our pooled incidence also supported the previous notion of higher frequency of persisting RT mutations in A-B interdomain compared with other region in RT (Figure 2).
RT and HBsAg mutations can occur simultaneously, due to the overlap of RT region and HBsAg gene sequences[19,124]. Liu et al[21] reported that 14 of 18 mutated positions in RT overlapped with HBsAg, and that RT mutations at 12 out of 14 RT positions (except those at rt124 and rt126) also led to simultaneous HBsAg mutations of 19 types in 16.67% (32/192) of isolates (Figure 1). Notably, these authors also found that RT mutations in the A-B interdomain could lead to simultaneous AA substitutions sI126A/N/S/, sG130N, sT131N/P, and sG145R of the overlapped ‘a’ determinant of HBsAg, including the most frequently described immune-escape mutation sG145R (1/192, 0.52%)[125,126]. Similarly, Kim et al[33] demonstrated that RT mutations at 10 of 42 NAr positions could lead to 15 types of simultaneous overlapped HBsAg mutations in 32.06% (42/131) patients. Of interest, they also found that the RT mutations at 3 NAr positions (rt134, rt139, and rt153) located in the overlapped HBsAg “a” determinant region from 22 treatment-naïve patients also had simultaneous “a” determinant mutations in two positions, S126 and S131, in 15 patients (15/22, 68.2%) (12 patients with mutations at rt134, leading to 10 changes of AA S126, and 8 patients with mutations at rt139, leading to 5 alterations of AA S131).
Overall, preexisting RT mutations are distributed in a non-random manner, and most frequently found in the A-B interdomain, overlapped with the HBsAg “a” determinant region, than in other domains. Moreover, the A-B interdomain also contains the most abundant mutations, indicating that these positions might be preexisting mutation hotspots in treatment-naïve patients. Of six positions’ mutation in the A-B interdomain, three RT mutations, rtD134E/N, rtN139D/E/H/K/Q, and rtW153E/Q/R, that overlap with HBsAg “a” determinant region are hotspots found most frequently in treatment-naïve patients, which could contribute to HBV viral persistence via generation of immune escape “a” determinant mutants proteins. In general, A-B interdomain mutations are prevalent in patients with genotype C2 infections and could contribute to HBV-associated disease, such as HCC and cirrhosis.
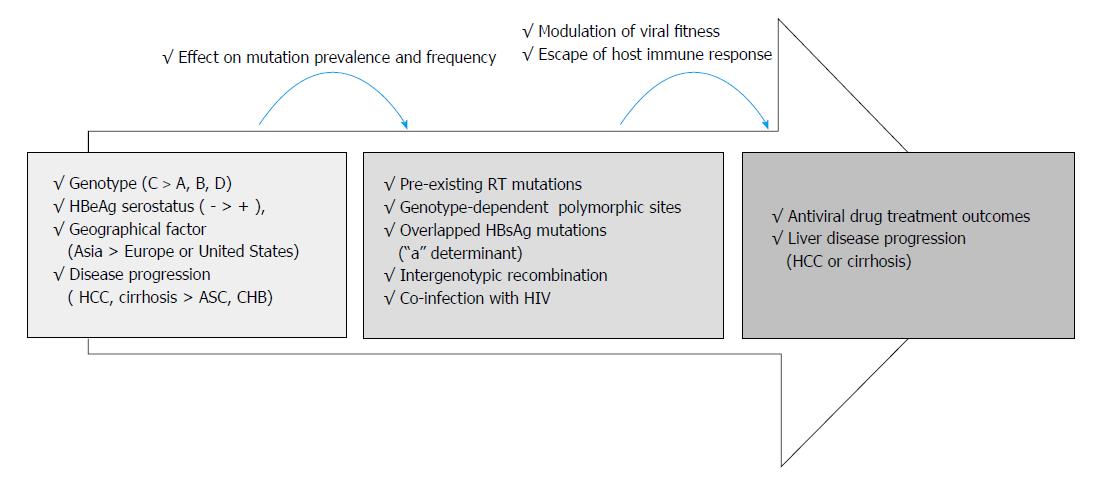
Preexisting HBV RT mutations in treatment-naïve patients are related to potential drug resistance and progression of liver disease, such as HCC or cirrhosis. In addition, genotype-dependent polymorphic amino acid substitution in RT can also affect the emergence of drug resistance and treatment outcomes. The reported prevalence of spontaneous RT mutations in treatment-naïve patients is varied, and largely depends on geographic factors, HBV genotypes, HBeAg serostatus, HBV viral loads, disease progression, intergenoytpic recombination, and co-infection with HIV. Different sensitivity of detection methodology used could also affect their prevalence results. The INNO-LiPA assay and UDPS method detect higher prevalence rates of preexisting RT mutations compared with direct PCR sequencing in treatment-naïve patients. Genotype C infection, HBeAg-negative status, and low viral loads are significantly associated with higher frequencies and prevalence rate of pre-existing HBV RT mutations. Higher frequencies of preexisting RT mutations were also generally associated with liver disease progression, including of HCC and cirrhosis. Eight mutations in RT region, rtL80I, rtD134N, rtN139K/T/H, rtY141F, rtM204I/V, rtF221Y, rtI224V, and rtM309K were significantly associated with progression of HCC in treatment-naïve patients. Of RT domains, preexisting RT mutations occur most frequently in the A-B interdomain which overlaps with the HBsAg “a” determinant region, in which mutations can lead to simultaneous viral immune escape (Figure 3). In conclusion, the presence of baseline preexisting RT mutations can affect drug treatment outcomes and disease progression in populations by modulation of viral fitness and host-immune responses.
Manuscript source: Invited manuscript
Specialty type: Gastroenterology and hepatology
Country of origin: South Korea
Peer-review report classification
Grade A (Excellent): 0
Grade B (Very good): 0
Grade C (Good): C, C
Grade D (Fair): 0
Grade E (Poor): 0
P- Reviewer: Quarleri J, Tamori A S- Editor: Gong ZM L- Editor: A E- Editor: Huang Y
| 1. | Beasley RP, Hwang LY, Lee GC, Lan CC, Roan CH, Huang FY, Chen CL. Prevention of perinatally transmitted hepatitis B virus infections with hepatitis B immune globulin and hepatitis B vaccine. Lancet. 1983;2:1099-1102. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 564] [Cited by in F6Publishing: 580] [Article Influence: 14.1] [Reference Citation Analysis (0)] |
| 2. | GBD 2013 Mortality and Causes of Death Collaborators. Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990-2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet. 2015;385:117-171. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 4902] [Cited by in F6Publishing: 4886] [Article Influence: 542.9] [Reference Citation Analysis (0)] |
| 3. | Nevens F, Main J, Honkoop P, Tyrrell DL, Barber J, Sullivan MT, Fevery J, De Man RA, Thomas HC. Lamivudine therapy for chronic hepatitis B: a six-month randomized dose-ranging study. Gastroenterology. 1997;113:1258-1263. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 211] [Cited by in F6Publishing: 221] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 4. | Marcellin P, Chang TT, Lim SG, Tong MJ, Sievert W, Shiffman ML, Jeffers L, Goodman Z, Wulfsohn MS, Xiong S. Adefovir dipivoxil for the treatment of hepatitis B e antigen-positive chronic hepatitis B. N Engl J Med. 2003;348:808-816. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1060] [Cited by in F6Publishing: 1107] [Article Influence: 52.7] [Reference Citation Analysis (0)] |
| 5. | Rivkin A. Entecavir: a new nucleoside analogue for the treatment of chronic hepatitis B. Drugs Today (Barc). 2007;43:201-220. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 11] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 6. | Matthews SJ. Telbivudine for the management of chronic hepatitis B virus infection. Clin Ther. 2007;29:2635-2653. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 41] [Cited by in F6Publishing: 42] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 7. | Jenh AM, Thio CL, Pham PA. Tenofovir for the treatment of hepatitis B virus. Pharmacotherapy. 2009;29:1212-1227. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 23] [Cited by in F6Publishing: 26] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 8. | Song ZL, Cui YJ, Zheng WP, Teng DH, Zheng H. Diagnostic and therapeutic progress of multi-drug resistance with anti-HBV nucleos(t)ide analogues. World J Gastroenterol. 2012;18:7149-7157. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 15] [Cited by in F6Publishing: 18] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 9. | Shi H, Han Z, Liu J, Xue J, Zhang S, Zhu Z, Xia J, Huang M. Comparing Efficacy of Lamivudine, Adefovir Dipivoxil, Telbivudine, and Entecavir in Treating Nucleoside Analogues Naïve for HBeAg-Negative Hepatitis B with Medium Hepatitis B Virus (HBV) DNA Levels. Med Sci Monit. 2017;23:5230-5236. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 7] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 10. | Nowak MA, Bonhoeffer S, Hill AM, Boehme R, Thomas HC, McDade H. Viral dynamics in hepatitis B virus infection. Proc Natl Acad Sci U S A. 1996;93:4398-4402. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 708] [Cited by in F6Publishing: 547] [Article Influence: 19.5] [Reference Citation Analysis (0)] |
| 11. | Caligiuri P, Cerruti R, Icardi G, Bruzzone B. Overview of hepatitis B virus mutations and their implications in the management of infection. World J Gastroenterol. 2016;22:145-154. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 102] [Cited by in F6Publishing: 92] [Article Influence: 11.5] [Reference Citation Analysis (1)] |
| 12. | Selabe SG, Lukhwareni A, Song E, Leeuw YG, Burnett RJ, Mphahlele MJ. Mutations associated with lamivudine-resistance in therapy-naïve hepatitis B virus (HBV) infected patients with and without HIV co-infection: implications for antiretroviral therapy in HBV and HIV co-infected South African patients. J Med Virol. 2007;79:1650-1654. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 35] [Cited by in F6Publishing: 40] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 13. | Rodriguez C, Chevaliez S, Bensadoun P, Pawlotsky JM. Characterization of the dynamics of hepatitis B virus resistance to adefovir by ultra-deep pyrosequencing. Hepatology. 2013;58:890-901. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 48] [Cited by in F6Publishing: 52] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 14. | Tenney DJ, Rose RE, Baldick CJ, Pokornowski KA, Eggers BJ, Fang J, Wichroski MJ, Xu D, Yang J, Wilber RB. Long-term monitoring shows hepatitis B virus resistance to entecavir in nucleoside-naïve patients is rare through 5 years of therapy. Hepatology. 2009;49:1503-1514. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 615] [Cited by in F6Publishing: 653] [Article Influence: 43.5] [Reference Citation Analysis (0)] |
| 15. | Zhang Y, Lian JQ, Li Y, Wang JP, Huang CX, Bai XF, Wang JP. Telbivudine plus adefovir therapy for chronic hepatitis B patients with virological breakthrough or genotypic resistance to telbivudine. Eur J Gastroenterol Hepatol. 2013;25:814-819. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12] [Cited by in F6Publishing: 13] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 16. | Strasfeld L, Chou S. Antiviral drug resistance: mechanisms and clinical implications. Infect Dis Clin North Am. 2010;24:809-833. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 48] [Cited by in F6Publishing: 54] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 17. | Sheldon J, Rodès B, Zoulim F, Bartholomeusz A, Soriano V. Mutations affecting the replication capacity of the hepatitis B virus. J Viral Hepat. 2006;13:427-434. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 97] [Cited by in F6Publishing: 86] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 18. | Stuyver LJ, Locarnini SA, Lok A, Richman DD, Carman WF, Dienstag JL, Schinazi RF. Nomenclature for antiviral-resistant human hepatitis B virus mutations in the polymerase region. Hepatology. 2001;33:751-757. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 307] [Cited by in F6Publishing: 321] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 19. | Sheldon J, Soriano V. Hepatitis B virus escape mutants induced by antiviral therapy. J Antimicrob Chemother. 2008;61:766-768. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 96] [Cited by in F6Publishing: 102] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 20. | Zheng J, Zeng Z, Zhang D, Yu Y, Wang F, Pan CQ. Prevalence and significance of Hepatitis B reverse transcriptase mutants in different disease stages of untreated patients. Liver Int. 2012;32:1535-1542. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 22] [Cited by in F6Publishing: 23] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 21. | Liu BM, Li T, Xu J, Li XG, Dong JP, Yan P, Yang JX, Yan L, Gao ZY, Li WP. Characterization of potential antiviral resistance mutations in hepatitis B virus reverse transcriptase sequences in treatment-naïve Chinese patients. Antiviral Res. 2010;85:512-519. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 69] [Cited by in F6Publishing: 75] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 22. | Solmone M, Vincenti D, Prosperi MC, Bruselles A, Ippolito G, Capobianchi MR. Use of massively parallel ultradeep pyrosequencing to characterize the genetic diversity of hepatitis B virus in drug-resistant and drug-naive patients and to detect minor variants in reverse transcriptase and hepatitis B S antigen. J Virol. 2009;83:1718-1726. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 129] [Cited by in F6Publishing: 139] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 23. | Tan YW, Ge GH, Zhao W, Gan JH, Zhao Y, Niu ZL, Zhang DJ, Chen L, Yu XJ, Yang LJ. YMDD motif mutations in chronic hepatitis B antiviral treatment naïve patients: a multi-center study. Braz J Infect Dis. 2012;16:250-255. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 14] [Cited by in F6Publishing: 14] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 24. | Fung SK, Mazzulli T, El-Kashab M, Sherman M, Popovic V, Sablon E. Lamivudine-Resistant Mutation among Treatment-Naive Hepatitis B Patients Is Common and May Be Associated with Treatment Failure. Hepatology. 2008;48:703a-703a. [Cited in This Article: ] |
| 25. | Mirandola S, Campagnolo D, Bortoletto G, Franceschini L, Marcolongo M, Alberti A. Large-scale survey of naturally occurring HBV polymerase mutations associated with anti-HBV drug resistance in untreated patients with chronic hepatitis B. J Viral Hepat. 2011;18:e212-e216. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 37] [Cited by in F6Publishing: 34] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 26. | Vutien P, Trinh HN, Garcia RT, Nguyen HA, Levitt BS, Nguyen K, da Silveira E, Daugherty T, Ahmed A, Garcia G. Mutations in HBV DNA polymerase associated with nucleos(t)ide resistance are rare in treatment-naive patients. Clin Gastroenterol Hepatol. 2014;12:1363-1370. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 22] [Cited by in F6Publishing: 24] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 27. | Zhao Y, Wu J, Sun L, Liu G, Li B, Zheng Y, Li X, Tao J. Prevalence of mutations in HBV DNA polymerase gene associated with nucleos(t)ide resistance in treatment-naive patients with Chronic Hepatitis B in Central China. Braz J Infect Dis. 2016;20:173-178. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 10] [Cited by in F6Publishing: 11] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 28. | Akarsu M, Sengonul A, Tankurt E, Sayiner AA, Topalak O, Akpinar H, Abacioglu YH. YMDD motif variants in inactive hepatitis B carriers detected by Inno-Lipa HBV DR assay. J Gastroenterol Hepatol. 2006;21:1783-1788. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 30] [Cited by in F6Publishing: 29] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 29. | Ciftci S, Keskin F, Cakiris A, Akyuz F, Pinarbasi B, Abaci N, Dincer E, Badur S, Kaymakoglu S, Ustek D. Analysis of potential antiviral resistance mutation profiles within the HBV reverse transcriptase in untreated chronic hepatitis B patients using an ultra-deep pyrosequencing method. Diagn Microbiol Infect Dis. 2014;79:25-30. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 13] [Cited by in F6Publishing: 14] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 30. | Zhang X, Li M, Xi H, Zhang R, Chen J, Zhang Y, Xu X. Pre-existing mutations related to tenofovir in chronic hepatitis B patients with long-term nucleos(t)ide analogue drugs treatment by ultra-deep pyrosequencing. Oncotarget. 2016;7:70264-70275. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 7] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 31. | Wasityastuti W, Yano Y, Widasari DI, Yamani LN, Ratnasari N, Heriyanto DS, Okada R, Tanahashi T, Murakami Y, Azuma T. Different Variants in Reverse Transcriptase Domain Determined by Ultra-deep Sequencing in Treatment-naïve and Treated Indonesian Patients Infected with Hepatitis B Virus. Kobe J Med Sci. 2016;62:E1-E8. [PubMed] [Cited in This Article: ] |
| 32. | Zöllner B, Sterneck M, Wursthorn K, Petersen J, Schröter M, Laufs R, Feucht HH. Prevalence, incidence, and clinical relevance of the reverse transcriptase V207I mutation outside the YMDD motif of the hepatitis B virus polymerase during lamivudine therapy. J Clin Microbiol. 2005;43:2503-2505. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 18] [Cited by in F6Publishing: 18] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 33. | Kim JE, Lee SY, Kim H, Kim KJ, Choe WH, Kim BJ. Naturally occurring mutations in the reverse transcriptase region of hepatitis B virus polymerase from treatment-naïve Korean patients infected with genotype C2. World J Gastroenterol. 2017;23:4222-4232. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 17] [Cited by in F6Publishing: 18] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 34. | Kobayashi S, Ide T, Sata M. Detection of YMDD motif mutations in some lamivudine-untreated asymptomatic hepatitis B virus carriers. J Hepatol. 2001;34:584-586. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 86] [Cited by in F6Publishing: 95] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 35. | Lee CZ, Lee HS, Huang GT, Yang PM, Sheu JC. Detection of YMDD mutation using mutant-specific primers in chronic hepatitis B patients before and after lamivudine treatment. World J Gastroenterol. 2006;12:5301-5305. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 11] [Cited by in F6Publishing: 13] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 36. | Tunçbilek S, Köse S, Elaldi A, Akman S. Lamivudine resistance in untreated chronic hepatitis B patients in Turkey. Turk J Gastroenterol. 2008;19:99-103. [PubMed] [Cited in This Article: ] |
| 37. | Huang CJ, Wu CF, Lan CY, Sung FY, Lin CL, Liu CJ, Liu HF, Yu MW. Impact of genetic heterogeneity in polymerase of hepatitis B virus on dynamics of viral load and hepatitis B progression. PLoS One. 2013;8:e70169. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 10] [Cited by in F6Publishing: 11] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 38. | Yamani LN, Yano Y, Utsumi T, Wasityastuti W, Rinonce HT, Widasari DI, Juniastuti , Lusida MI, Soetjipto , Hayashi Y. Profile of Mutations in the Reverse Transcriptase and Overlapping Surface Genes of Hepatitis B Virus (HBV) in Treatment-Naïve Indonesian HBV Carriers. Jpn J Infect Dis. 2017;70:647-655. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12] [Cited by in F6Publishing: 14] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 39. | Wang LP, Han FZ, Duan HL, Ji F, Yan XB, Fan YC, Wang K. Hepatitis B virus pre-existing drug resistant mutation is related to the genotype and disease progression. J Infect Dev Countr. 2017;11:727-732. [DOI] [Cited in This Article: ] [Cited by in Crossref: 4] [Cited by in F6Publishing: 4] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 40. | Ismail AM, Samuel P, Eapen CE, Kannangai R, Abraham P. Antiviral resistance mutations and genotype-associated amino acid substitutions in treatment-naïve hepatitis B virus-infected individuals from the Indian subcontinent. Intervirology. 2012;55:36-44. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 28] [Cited by in F6Publishing: 32] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 41. | Pacheco SR, Dos Santos MIMA, Stocker A, Zarife MAS, Schinoni MI, Paraná R, Dos Reis MG, Silva LK. Genotyping of HBV and tracking of resistance mutations in treatment-naïve patients with chronic hepatitis B. Infect Drug Resist. 2017;10:201-207. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12] [Cited by in F6Publishing: 13] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 42. | Fan J, Zhang Y, Xiong H, Wang Y, Guo X. Nucleotide analogue-resistant mutations in hepatitis B viral genomes found in hepatitis B patients. J Gen Virol. 2015;96:663-670. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 11] [Cited by in F6Publishing: 12] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 43. | Li X, Liu Y, Zhao P, Wang Y, Chen L, Xin S, Zhang XX, Xu D. Investigation into drug-resistant mutations of HBV from 845 nucleoside/nucleotide analogue-naive Chinese patients with chronic HBV infection. Antivir Ther. 2015;20:141-147. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12] [Cited by in F6Publishing: 14] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 44. | Singla B, Chakraborti A, Sharma BK, Kapil S, Chawla YK, Arora SK, Das A, Dhiman RK, Duseja A. Hepatitis B virus reverse transcriptase mutations in treatment Naïve chronic hepatitis B patients. J Med Virol. 2013;85:1155-1162. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 17] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 45. | Nguyen MH, Garcia RT, Trinh HN, Nguyen HA, Nguyen KK, Nguyen LH, Levitt B. Prevalence of hepatitis B virus DNA polymerase mutations in treatment-naïve patients with chronic hepatitis B. Aliment Pharmacol Ther. 2009;30:1150-1158. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 17] [Cited by in F6Publishing: 16] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 46. | Xu J, Wu B, Wang JH, Huang L, Wang DY, Zhao L, Zhao GP, Wang Y. Pre-existing mutations in reverse transcriptase of hepatitis B virus in treatment-naive Chinese patients with chronic hepatitis B. PLoS One. 2015;10:e0117429. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 13] [Cited by in F6Publishing: 16] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 47. | Qian F, Zou W, Qin J, Li D. Naturally occurring genotypic drug-resistant mutations of HBV in Huzhou, China: a single-center study. Infect Drug Resist. 2017;10:507-509. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3] [Cited by in F6Publishing: 3] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 48. | Matsuda M, Suzuki F, Suzuki Y, Tsubota A, Akuta N, Hosaka T, Someya T, Kobayashi M, Saitoh S, Arase Y. Low rate of YMDD motif mutations in polymerase gene of hepatitis B virus in chronically infected patients not treated with lamivudine. J Gastroenterol. 2004;39:34-40. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 20] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 49. | Pollicino T, Isgrò G, Di Stefano R, Ferraro D, Maimone S, Brancatelli S, Squadrito G, Di Marco V, Craxì A, Raimondo G. Variability of reverse transcriptase and overlapping S gene in hepatitis B virus isolates from untreated and lamivudine-resistant chronic hepatitis B patients. Antivir Ther. 2009;14:649-654. [PubMed] [Cited in This Article: ] |
| 50. | Mahabadi M, Norouzi M, Alavian SM, Samimirad K, Azad TM, Saberfar E, Mahmoodi M, Ramezani F, Karimzadeh H, Malekzadeh R. Drug-related mutational patterns in hepatitis B virus (HBV) reverse transcriptase proteins from Iranian treatment-naïve chronic HBV patients. Hepat Mon. 2013;13:e6712. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 19] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 51. | Salpini R, Svicher V, Cento V, Gori C, Bertoli A, Scopelliti F, Micheli V, Cappiello T, Spanò A, Rizzardini G. Characterization of drug-resistance mutations in HBV D-genotype chronically infected patients, naïve to antiviral drugs. Antiviral Res. 2011;92:382-385. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 18] [Cited by in F6Publishing: 18] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 52. | Margeridon-Thermet S, Shulman NS, Ahmed A, Shahriar R, Liu T, Wang C, Holmes SP, Babrzadeh F, Gharizadeh B, Hanczaruk B. Ultra-deep pyrosequencing of hepatitis B virus quasispecies from nucleoside and nucleotide reverse-transcriptase inhibitor (NRTI)-treated patients and NRTI-naive patients. J Infect Dis. 2009;199:1275-1285. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 170] [Cited by in F6Publishing: 180] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 53. | Han Y, Huang LH, Liu CM, Yang S, Li J, Lin ZM, Kong XF, Yu DM, Zhang DH, Jin GD. Characterization of hepatitis B virus reverse transcriptase sequences in Chinese treatment naive patients. J Gastroenterol Hepatol. 2009;24:1417-1423. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 33] [Cited by in F6Publishing: 30] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 54. | Panigrahi R, Biswas A, De BK, Chakrabarti S, Chakravarty R. Characterization of antiviral resistance mutations among the Eastern Indian Hepatitis B virus infected population. Virol J. 2013;10:56. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12] [Cited by in F6Publishing: 14] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 55. | Altindis M, Aslan FG, Koroglu M, Eren A, Demir L, Uslan MI, Aslan S, Ozdemir M, Baykan M. Hepatitis B Virus Carrying Drug-resistance Compensatory Mutations in Chronically Infected Treatment-naive Patients. Viral Hepat J. 2016;22:103-107. [DOI] [Cited in This Article: ] [Cited by in Crossref: 2] [Cited by in F6Publishing: 2] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 56. | Amini-Bavil-Olyaee S, Hosseini SY, Sabahi F, Alavian SM. Hepatitis B virus (HBV) genotype and YMDD motif mutation profile among patients infected with HBV and untreated with lamivudine. Int J Infect Dis. 2008;12:83-87. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 20] [Cited by in F6Publishing: 22] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 57. | Jardi R, Rodriguez-Frias F, Schaper M, Ruiz G, Elefsiniotis I, Esteban R, Buti M. Hepatitis B virus polymerase variants associated with entecavir drug resistance in treatment-naive patients. J Viral Hepat. 2007;14:835-840. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 3] [Cited by in F6Publishing: 12] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 58. | Li XG, Liu BM, Xu J, Liu XE, Ding H, Li T. Discrepancy of potential antiviral resistance mutation profiles within the HBV reverse transcriptase between nucleos(t)ide analogue-untreated and -treated patients with chronic hepatitis B in a hospital in China. J Med Virol. 2012;84:207-216. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 20] [Cited by in F6Publishing: 21] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 59. | Lampertico P, Vigano M, Facchetti F, Puoti M, Minola E, Suter F, Brunetto M, Coco B, Fargion S, Fotta E. Effectiveness of entecavir for the treatment of NUC-naive chronic hepatitis B patients: A large multicenter cohort study in clinical practice. Hepatology. 2008;48:707A-708A. [Cited in This Article: ] |
| 60. | Ramezani A, Velayati AA, Roshan MR, Gachkar L, Banifazl M, Keyvani H, Aghakhani A. Rate of YMDD motif mutants in lamivudine-untreated Iranian patients with chronic hepatitis B virus infection. Int J Infect Dis. 2008;12:252-255. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 11] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 61. | Ghosh S, Mondal RK, Banerjee P, Nandi M, Sarkar S, Das K, Santra A, Banerjee S, Chowdhury A, Datta S. Tracking the naturally occurring mutations across the full-length genome of hepatitis B virus of genotype D in different phases of chronic e-antigen-negative infection. Clin Microbiol Infect. 2012;18:E412-E418. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 23] [Cited by in F6Publishing: 25] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 62. | Ludwig AD, Goebel T, Adams O, Baumann N, Hauck K, Fey H, Hengel H, Haussinger D, Erhardt A. Primary Resistance Mutations against Nucleos(t)ide Analogues in Treatment Name Patients with HBV-Infection. Hepatology. 2008;48:701A-701A. [Cited in This Article: ] |
| 63. | Hamidi-Fard M, Makvandi M, Samarbaf-Zadeh A, Hajiani E, Shayesteh A, Masjedizadeh A. Mutation analysis of hepatitis B virus reverse transcriptase region among untreated chronically infected patients in Ahvaz city (South-West of Iran). Indian J Med Microbiol. 2013;31:360-365. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 7] [Cited by in F6Publishing: 7] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 64. | Villa E, Lei B, Taliani G, Graziosi A, Critelli R, Luongo M, Gennari W, Bianchini M, Ferretti I. Pretreatment with pegylated interferon prevents emergence of lamivudine mutants in lamivudine-naive patients: a pilot study. Antivir Ther. 2009;14:1081-1087. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 4] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 65. | Mantovani N, Cicero M, Santana LC, Silveira C, do Carmo EP, Abrão PR, Diaz RS, Caseiro MM, Komninakis SV. Detection of lamivudine-resistant variants and mutations related to reduced antigenicity of HBsAg in individuals from the cities of Santos and São Paulo, Brazil. Virol J. 2013;10:320. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 10] [Cited by in F6Publishing: 11] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 66. | Aberle SW, Kletzmayr J, Watschinger B, Schmied B, Vetter N, Puchhammer-Stöckl E. Comparison of sequence analysis and the INNO-LiPA HBV DR line probe assay for detection of lamivudine-resistant hepatitis B virus strains in patients under various clinical conditions. J Clin Microbiol. 2001;39:1972-1974. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 47] [Cited by in F6Publishing: 49] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 67. | Feeney F, Fanning LJ, Horgan M. Baseline genotypic resistance in untreated hepatitis B virus infection. Gastroenterology. 2007;132:35A. [Cited in This Article: ] |
| 68. | Masaadeh HA, Hayajneh WA, Alqudah EA. Hepatitis B virus genotypes and lamivudine resistance mutations in Jordan. World J Gastroenterol. 2008;14:7231-7234. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 17] [Cited by in F6Publishing: 21] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 69. | Lok AS, Zoulim F, Locarnini S, Bartholomeusz A, Ghany MG, Pawlotsky JM, Liaw YF, Mizokami M, Kuiken C; Hepatitis B Virus Drug Resistance Working Group. Antiviral drug-resistant HBV: standardization of nomenclature and assays and recommendations for management. Hepatology. 2007;46:254-265. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 359] [Cited by in F6Publishing: 385] [Article Influence: 22.6] [Reference Citation Analysis (0)] |
| 70. | Shaw T, Bartholomeusz A, Locarnini S. HBV drug resistance: mechanisms, detection and interpretation. J Hepatol. 2006;44:593-606. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 168] [Cited by in F6Publishing: 177] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 71. | Langley DR, Walsh AW, Baldick CJ, Eggers BJ, Rose RE, Levine SM, Kapur AJ, Colonno RJ, Tenney DJ. Inhibition of hepatitis B virus polymerase by entecavir. J Virol. 2007;81:3992-4001. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 141] [Cited by in F6Publishing: 150] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 72. | Locarnini S. Primary resistance, multidrug resistance, and cross-resistance pathways in HBV as a consequence of treatment failure. Hepatol Int. 2008;2:147-151. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 110] [Cited by in F6Publishing: 115] [Article Influence: 7.2] [Reference Citation Analysis (0)] |
| 73. | Zhang Q, Liao Y, Cai B, Li Y, Li L, Zhang J, An Y, Wang L. Incidence of natural resistance mutations in naïve chronic hepatitis B patients: a systematic review and meta-analysis. J Gastroenterol Hepatol. 2015;30:252-261. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 23] [Cited by in F6Publishing: 23] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 74. | Lai CL, Leung N, Teo EK, Tong M, Wong F, Hann HW, Han S, Poynard T, Myers M, Chao G. A 1-year trial of telbivudine, lamivudine, and the combination in patients with hepatitis B e antigen-positive chronic hepatitis B. Gastroenterology. 2005;129:528-536. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 359] [Cited by in F6Publishing: 367] [Article Influence: 19.3] [Reference Citation Analysis (0)] |
| 75. | Fu L, Cheng YC. Role of additional mutations outside the YMDD motif of hepatitis B virus polymerase in L(-)SddC (3TC) resistance. Biochem Pharmacol. 1998;55:1567-1572. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 102] [Cited by in F6Publishing: 107] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 76. | Wakil SM, Kazim SN, Khan LA, Raisuddin S, Parvez MK, Guptan RC, Thakur V, Hasnain SE, Sarin SK. Prevalence and profile of mutations associated with lamivudine therapy in Indian patients with chronic hepatitis B in the surface and polymerase genes of hepatitis B virus. J Med Virol. 2002;68:311-318. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 36] [Cited by in F6Publishing: 40] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 77. | Yang H, Westland CE, Delaney WE 4th, Heathcote EJ, Ho V, Fry J, Brosgart C, Gibbs CS, Miller MD, Xiong S. Resistance surveillance in chronic hepatitis B patients treated with adefovir dipivoxil for up to 60 weeks. Hepatology. 2002;36:464-473. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 141] [Cited by in F6Publishing: 143] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 78. | Ahn SH, Park YK, Park ES, Kim JH, Kim DH, Lim KH, Jang MS, Choe WH, Ko SY, Sung IK. The impact of the hepatitis B virus polymerase rtA181T mutation on replication and drug resistance is potentially affected by overlapping changes in surface gene. J Virol. 2014;88:6805-6818. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 29] [Cited by in F6Publishing: 29] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 79. | Schildgen O, Sirma H, Funk A, Olotu C, Wend UC, Hartmann H, Helm M, Rockstroh JK, Willems WR, Will H. Variant of hepatitis B virus with primary resistance to adefovir. N Engl J Med. 2006;354:1807-1812. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 167] [Cited by in F6Publishing: 182] [Article Influence: 10.1] [Reference Citation Analysis (0)] |
| 80. | Kramvis A, Kew M, François G. Hepatitis B virus genotypes. Vaccine. 2005;23:2409-2423. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 256] [Cited by in F6Publishing: 255] [Article Influence: 13.4] [Reference Citation Analysis (0)] |
| 81. | Schaefer S. Hepatitis B virus taxonomy and hepatitis B virus genotypes. World J Gastroenterol. 2007;13:14-21. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 243] [Cited by in F6Publishing: 229] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 82. | Sunbul M. Hepatitis B virus genotypes: global distribution and clinical importance. World J Gastroenterol. 2014;20:5427-5434. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 264] [Cited by in F6Publishing: 263] [Article Influence: 26.3] [Reference Citation Analysis (2)] |
| 83. | Mirandola S, Sebastiani G, Rossi C, Velo E, Erne EM, Vario A, Tempesta D, Romualdi C, Campagnolo D, Alberti A. Genotype-specific mutations in the polymerase gene of hepatitis B virus potentially associated with resistance to oral antiviral therapy. Antiviral Res. 2012;96:422-429. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 22] [Cited by in F6Publishing: 21] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 84. | Damerow H, Yuen L, Wiegand J, Walker C, Bock CT, Locarnini S, Tillmann HL. Mutation pattern of lamivudine resistance in relation to hepatitis B genotypes: hepatitis B genotypes differ in their lamivudine resistance associated mutation pattern. J Med Virol. 2010;82:1850-1858. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 28] [Cited by in F6Publishing: 29] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 85. | Colonno RJ, Rose R, Baldick CJ, Levine S, Pokornowski K, Yu CF, Walsh A, Fang J, Hsu M, Mazzucco C. Entecavir resistance is rare in nucleoside naïve patients with hepatitis B. Hepatology. 2006;44:1656-1665. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 279] [Cited by in F6Publishing: 292] [Article Influence: 16.2] [Reference Citation Analysis (0)] |
| 86. | Ciancio A, Smedile A, Rizzetto M, Lagget M, Gerin J, Korba B. Identification of HBV DNA sequences that are predictive of response to lamivudine therapy. Hepatology. 2004;39:64-73. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 34] [Cited by in F6Publishing: 36] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 87. | Chu CJ, Keeffe EB, Han SH, Perrillo RP, Min AD, Soldevila-Pico C, Carey W, Brown RS Jr, Luketic VA, Terrault N, Lok AS; U. S. HBV Epidemiology Study Group. Prevalence of HBV precore/core promoter variants in the United States. Hepatology. 2003;38:619-628. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 163] [Cited by in F6Publishing: 171] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 88. | Kirishima T, Okanoue T, Daimon Y, Itoh Y, Nakamura H, Morita A, Toyama T, Minami M. Detection of YMDD mutant using a novel sensitive method in chronic liver disease type B patients before and during lamivudine treatment. J Hepatol. 2002;37:259-265. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 67] [Cited by in F6Publishing: 73] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 89. | Zhu B, Wang T, Wei X, Zhuo Y, Liu A, Zhang G. Accumulation of mutations in reverse transcriptase of hepatitis B virus is associated with liver disease severity in treatment-naïve Chinese patients with chronic hepatitis B. Adv Clin Exp Med. 2017;26:1123-1129. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 6] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 90. | Fukai K, Zhang KY, Imazeki F, Kurihara T, Mikata R, Yokosuka O. Association between lamivudine sensitivity and the number of substitutions in the reverse transcriptase region of the hepatitis B virus polymerase. J Viral Hepat. 2007;14:661-666. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 12] [Cited by in F6Publishing: 11] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 91. | Wang YX, Xu X, Luo C, Ma ZM, Jiang HL, Ding JP, Wen YM. A putative new domain target for anti-hepatitis B virus: residues flanking hepatitis B virus reverse transcriptase residue 306 (rtP306). J Med Virol. 2007;79:676-682. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 9] [Cited by in F6Publishing: 11] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 92. | Pollicino T, Cacciola I, Saffioti F, Raimondo G. Hepatitis B virus PreS/S gene variants: pathobiology and clinical implications. J Hepatol. 2014;61:408-417. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 175] [Cited by in F6Publishing: 186] [Article Influence: 18.6] [Reference Citation Analysis (0)] |
| 93. | Huang ZM, Huang QW, Qin YQ, He YZ, Qin HJ, Zhou YN, Xu X, Huang MJ. YMDD mutations in patients with chronic hepatitis B untreated with antiviral medicines. World J Gastroenterol. 2005;11:867-870. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 27] [Cited by in F6Publishing: 31] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 94. | Nishijima N, Marusawa H, Ueda Y, Takahashi K, Nasu A, Osaki Y, Kou T, Yazumi S, Fujiwara T, Tsuchiya S. Dynamics of hepatitis B virus quasispecies in association with nucleos(t)ide analogue treatment determined by ultra-deep sequencing. PLoS One. 2012;7:e35052. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 65] [Cited by in F6Publishing: 68] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 95. | Sugauchi F, Orito E, Ichida T, Kato H, Sakugawa H, Kakumu S, Ishida T, Chutaputti A, Lai CL, Ueda R. Hepatitis B virus of genotype B with or without recombination with genotype C over the precore region plus the core gene. J Virol. 2002;76:5985-5992. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 218] [Cited by in F6Publishing: 228] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 96. | Yang J, Xing K, Deng R, Wang J, Wang X. Identification of Hepatitis B virus putative intergenotype recombinants by using fragment typing. J Gen Virol. 2006;87:2203-2215. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 46] [Cited by in F6Publishing: 50] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 97. | Rehermann B, Nascimbeni M. Immunology of hepatitis B virus and hepatitis C virus infection. Nat Rev Immunol. 2005;5:215-229. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 1202] [Cited by in F6Publishing: 1174] [Article Influence: 61.8] [Reference Citation Analysis (0)] |
| 98. | Shi W, Carr MJ, Dunford L, Zhu C, Hall WW, Higgins DG. Identification of novel inter-genotypic recombinants of human hepatitis B viruses by large-scale phylogenetic analysis. Virology. 2012;427:51-59. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 37] [Cited by in F6Publishing: 38] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 99. | Shi W, Zhu C, Zheng W, Carr MJ, Higgins DG, Zhang Z. Subgenotype reclassification of genotype B hepatitis B virus. BMC Gastroenterol. 2012;12:116. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 17] [Cited by in F6Publishing: 19] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 100. | Liu B, Yang JX, Yan L, Zhuang H, Li T. Novel HBV recombinants between genotypes B and C in 3’-terminal reverse transcriptase (RT) sequences are associated with enhanced viral DNA load, higher RT point mutation rates and place of birth among Chinese patients. Infect Genet Evol. 2018;57:26-35. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 4] [Cited by in F6Publishing: 10] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 101. | Lee SY, Lee SH, Kim JE, Kim H, Kim K, Kook YH, Kim BJ. Identification of Novel A2/C2 Inter-Genotype Recombinants of Hepatitis B Virus from a Korean Chronic Patient Co-Infected with Both Genotype A2 and C2. Int J Mol Sci. 2017;18:pii: E737. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 4] [Cited by in F6Publishing: 4] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 102. | Pan D, Zhou B, Yang J, Hou J. [Identification of hepatitis B virus intergenotype recombination in Chinese patients]. Nan Fang Yi Ke Da Xue Xue Bao. 2014;34:1436-1442. [PubMed] [Cited in This Article: ] |
| 103. | Simmonds P, Midgley S. Recombination in the genesis and evolution of hepatitis B virus genotypes. J Virol. 2005;79:15467-15476. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 177] [Cited by in F6Publishing: 187] [Article Influence: 10.4] [Reference Citation Analysis (0)] |
| 104. | Burnett RJ, François G, Kew MC, Leroux-Roels G, Meheus A, Hoosen AA, Mphahlele MJ. Hepatitis B virus and human immunodeficiency virus co-infection in sub-Saharan Africa: a call for further investigation. Liver Int. 2005;25:201-213. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 93] [Cited by in F6Publishing: 104] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 105. | Thio CL, Seaberg EC, Skolasky R Jr, Phair J, Visscher B, Muñoz A, Thomas DL; Multicenter AIDS Cohort Study. HIV-1, hepatitis B virus, and risk of liver-related mortality in the Multicenter Cohort Study (MACS). Lancet. 2002;360:1921-1926. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 760] [Cited by in F6Publishing: 739] [Article Influence: 33.6] [Reference Citation Analysis (0)] |
| 106. | Audsley J, Arrifin N, Yuen LK, Ayres A, Crowe SM, Bartholomeusz A, Locarnini SA, Mijch A, Lewin SR, Sasadeusz J. Prolonged use of tenofovir in HIV/hepatitis B virus (HBV)-coinfected individuals does not lead to HBV polymerase mutations and is associated with persistence of lamivudine HBV polymerase mutations. HIV Med. 2009;10:229-235. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 25] [Cited by in F6Publishing: 29] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 107. | Quarleri J, Moretti F, Bouzas MB, Laufer N, Carrillo MG, Giuliano SF, Pérez H, Cahn P, Salomon H. Hepatitis B virus genotype distribution and its lamivudine-resistant mutants in HIV-coinfected patients with chronic and occult hepatitis B. AIDS Res Hum Retroviruses. 2007;23:525-531. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 29] [Cited by in F6Publishing: 31] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 108. | Makondo E, Bell TG, Kramvis A. Genotyping and molecular characterization of hepatitis B virus from human immunodeficiency virus-infected individuals in southern Africa. PLoS One. 2012;7:e46345. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 36] [Cited by in F6Publishing: 38] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 109. | Selabe SG, Song E, Burnett RJ, Mphahlele MJ. Frequent detection of hepatitis B virus variants associated with lamivudine resistance in treated South African patients infected chronically with different HBV genotypes. J Med Virol. 2009;81:996-1001. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 17] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 110. | Thio CL, Smeaton L, Saulynas M, Hwang H, Saravanan S, Kulkarni S, Hakim J, Nyirenda M, Iqbal HS, Lalloo UG. Characterization of HIV-HBV coinfection in a multinational HIV-infected cohort. AIDS. 2013;27:191-201. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 68] [Cited by in F6Publishing: 70] [Article Influence: 6.4] [Reference Citation Analysis (1)] |
| 111. | Archampong TN, Boyce CL, Lartey M, Sagoe KW, Obo-Akwa A, Kenu E, Blackard JT, Kwara A. HBV genotypes and drug resistance mutations in antiretroviral treatment-naive and treatment-experienced HBV-HIV-coinfected patients. Antivir Ther. 2017;22:13-20. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 16] [Cited by in F6Publishing: 8] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 112. | Warner N, Locarnini S, Kuiper M, Bartholomeusz A, Ayres A, Yuen L, Shaw T. The L80I substitution in the reverse transcriptase domain of the hepatitis B virus polymerase is associated with lamivudine resistance and enhanced viral replication in vitro. Antimicrob Agents Chemother. 2007;51:2285-2292. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 76] [Cited by in F6Publishing: 82] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 113. | Shi YH, Shi CH. Molecular characteristics and stages of chronic hepatitis B virus infection. World J Gastroenterol. 2009;15:3099-3105. [PubMed] [DOI] [Cited in This Article: ] [Cited by in CrossRef: 72] [Cited by in F6Publishing: 66] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 114. | Yin F, Xie Y, Fan H, Zhang J, Guo Z. Mutations in hepatitis B virus polymerase are associated with the postoperative survival of hepatocellular carcinoma patients. PLoS One. 2017;12:e0189730. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 7] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 115. | Li H, Jia J, Wang M, Wang H, Gu X, Fang M, Gao C. F221Y mutation in hepatitis B virus reverse transcriptase is associated with hepatocellular carcinoma prognosis following liver resection. Mol Med Rep. 2017;15:3292-3300. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 6] [Cited by in F6Publishing: 7] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 116. | Wu Y, Gan Y, Gao F, Zhao Z, Jin Y, Zhu Y, Sun Z, Wu H, Chen T, Wang J. Novel natural mutations in the hepatitis B virus reverse transcriptase domain associated with hepatocellular carcinoma. PLoS One. 2014;9:e94864. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 13] [Cited by in F6Publishing: 15] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 117. | Kao JH, Chen PJ, Lai MY, Chen DS. Hepatitis B genotypes correlate with clinical outcomes in patients with chronic hepatitis B. Gastroenterology. 2000;118:554-559. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 684] [Cited by in F6Publishing: 668] [Article Influence: 27.8] [Reference Citation Analysis (0)] |
| 118. | Lee CM, Chen CH, Lu SN, Tung HD, Wang JH, Chen TM, Huang CC. Hepatitis B virus genotypes and the progression of chronic liver disease. J Hepatol. 2002;36:238. [DOI] [Cited in This Article: ] |
| 119. | Fujie H, Moriya K, Shintani Y, Yotsuyanagi H, Iino S, Koike K. Hepatitis B virus genotypes and hepatocellular carcinoma in Japan. Gastroenterology. 2001;120:1564-1565. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 70] [Cited by in F6Publishing: 69] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 120. | Tsubota A, Arase Y, Ren F, Tanaka H, Ikeda K, Kumada H. Genotype may correlate with liver carcinogenesis and tumor characteristics in cirrhotic patients infected with hepatitis B virus subtype adw. J Med Virol. 2001;65:257-265. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 85] [Cited by in F6Publishing: 89] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 121. | Zhong YW, Li J, Song HB, Duan ZP, Dong Y, Xing XY, Li XD, Gu ML, Han YK, Zhu SS. Virologic and clinical characteristics of HBV genotypes/subgenotypes in 487 Chinese pediatric patients with CHB. BMC Infect Dis. 2011;11:262. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 15] [Cited by in F6Publishing: 15] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 122. | Chan HL, Hui AY, Wong ML, Tse AM, Hung LC, Wong VW, Sung JJ. Genotype C hepatitis B virus infection is associated with an increased risk of hepatocellular carcinoma. Gut. 2004;53:1494-1498. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 369] [Cited by in F6Publishing: 366] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 123. | Kwon H, Lok AS. Hepatitis B therapy. Nat Rev Gastroenterol Hepatol. 2011;8:275-284. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 191] [Cited by in F6Publishing: 195] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 124. | Datta S, Chatterjee S, Veer V, Chakravarty R. Molecular biology of the hepatitis B virus for clinicians. J Clin Exp Hepatol. 2012;2:353-365. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 40] [Cited by in F6Publishing: 50] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 125. | Avellón A, Echevarria JM. Frequency of hepatitis B virus ‘a’ determinant variants in unselected Spanish chronic carriers. J Med Virol. 2006;78:24-36. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 82] [Cited by in F6Publishing: 88] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 126. | Svicher V, Gori C, Trignetti M, Visca M, Micheli V, Bernassola M, Salpini R, Gubertini G, Longo R, Niero F. The profile of mutational clusters associated with lamivudine resistance can be constrained by HBV genotypes. J Hepatol. 2009;50:461-470. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 34] [Cited by in F6Publishing: 34] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 127. | Rhee SY, Margeridon-Thermet S, Nguyen MH, Liu TF, Kagan RM, Beggel B, Verheyen J, Kaiser R, Shafer RW. Hepatitis B virus reverse transcriptase sequence variant database for sequence analysis and mutation discovery. Antiviral Res. 2010;88:269-275. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 41] [Cited by in F6Publishing: 41] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 128. | Ji D, Liu Y, Li L, Xu Z, Si LL, Dai JZ, Li X, Wang L, Yao Z, Xin SJ. The rtL229 substitutions in the reverse transcriptase region of hepatitis B virus (HBV) polymerase are potentially associated with lamivudine resistance as a compensatory mutation. J Clin Virol. 2012;54:66-72. [PubMed] [DOI] [Cited in This Article: ] [Cited by in Crossref: 31] [Cited by in F6Publishing: 31] [Article Influence: 2.6] [Reference Citation Analysis (0)] |