Published online Oct 28, 2009. doi: 10.3748/wjg.15.5058
Revised: September 27, 2009
Accepted: October 4, 2009
Published online: October 28, 2009
AIM: To investigate the correlation between uridine diphosphate glucuronosyl transferase 1A1 (UGT1A1) gene polymorphisms and irinotecan-associated side effects and parameters of drug efficacy in patients with metastatic colorectal cancer (mCRC) receiving a low-dose weekly irinotecan chemotherapeutic regimen.
METHODS: Genotypes were retrospectively evaluated by gene scan analysis on the ABI 310 sequencer of the TATAA box in the promoter region of the UGT1A1 gene in blood samples from 105 patients who had received 1st line irinotecan-based chemotherapy for mCRC.
RESULTS: The distribution of the genotypes was as follows: wild type genotype (WT) (6/6) 39.0%, heterozygous genotype (6/7) 49.5%, and homozygous genotype (7/7) 9.5%. The overall response rate (OR) was similar between patients carrying the (6/7, 7/7) or the WT genotype (6/6) (44.3% vs 43.2%, P = 0.75). Neither time to progression [(TTP) 8.1 vs 8.2 mo, P = 0.97] nor overall survival [(OS) 21.2 vs 18.9 mo, P = 0.73] differed significantly in patients who carried the (6/6) when compared to the (6/7, 7/7) genotype. No significant differences in toxicity were observed: Grade 3 and 4 delayed diarrhoea [(6/7, 7/7) vs (6/6); 13.0% vs 6.2%, P = 0.08], treatment delays [(6/7, 7/7) vs (6/6); 25.1% vs 19.3%, P =0.24] or dose reductions [(6/7, 7/7) vs (6/6); 21.5% vs 27.2%, P = 0.07].
CONCLUSION: This analysis demonstrates the non-significant influence of the UGT1A1 gene polymorphism on efficacy and rate of irinotecan-associated toxicity in mCRC patients receiving low-dose irinotecan based chemotherapy.
-
Citation: Schulz C, Heinemann V, Schalhorn A, Moosmann N, Zwingers T, Boeck S, Giessen C, Stemmler HJ.
UGT1A1 gene polymorphism: Impact on toxicity and efficacy of irinotecan-based regimens in metastatic colorectal cancer. World J Gastroenterol 2009; 15(40): 5058-5066 - URL: https://www.wjgnet.com/1007-9327/full/v15/i40/5058.htm
- DOI: https://dx.doi.org/10.3748/wjg.15.5058
| Total (n = 103) | WT (6/6) (n = 41) | (6/7) (n = 52) | (7/7) (n = 10) | |
| Age (yr) median (range) | 64 (41-79) | 63 (41-76) | 65 (42-79) | 65 (52-74) |
| Gender (M/F) | 70.5/29.5 | 70.7/29.3 | 67.3/32.7 | 80.0-20.0 |
| Colon cancer | 59.0 | 51.2 | 63.5 | 80.0 |
| Rectal cancer | 41.0 | 48.8 | 36.5 | 20.0 |
| Adjuvant pre-treatment | 35.2 | 43.9 | 30.8 | 20.0 |
| KPS 70%-90% | 43.8 | 29.3 | 51.9 | 70.0 |
| KPS 100% | 56.2 | 70.7 | 48.1 | 30.0 |
| LDH ≤ 240 U/L | 55.2 | 61.0 | 48.1 | 70.0 |
| LDH > 240 U/L | 44.8 | 39.0 | 51.9 | 30.0 |
| UGT1A1 status | FOLFIRI | IROX | All |
| WT (6/6) | 19 (34.5) | 22 (44.0) | 41 (39.0) |
| (6/7) | 28 (50.9) | 24 (48.0) | 52 (49.5) |
| (7/7) | 7 (12.7) | 3 (6.0) | 10 (9.5) |
| (5/7) | 1 (1.8) | 1 (2.0) | 2 (1.9) |
| WT (6/6) | (6/7, 7/7) | All | χ² test [WT vs (6/7, 7/7)] | |
| CR | 3 (8.1) | 7 (11.5) | 10 (10.2) | |
| PR | 13 (35.1) | 20 (32.8) | 33 (33.7) | |
| SD | 11 (29.7) | 29 (47.5) | 40 (40.8) | |
| PD | 7 (18.9) | 5 (8.2) | 12 (12.2) | |
| NA | 3 (8.1) | - | 3 (3.1) | |
| OR | 16 (43.2) | 27 (44.3) | 43 (43.9) | P > 0.05 |
| DCR | 27 (73.0) | 56 (91.8) | 83 (84.7) | P < 0.05 |
| Toxicity WHO | mFOLFIRI | mIROX | χ² test | ||||||
| Grade 1 | Grade 2 | Grade 3 | Grade 4 | Grade 1 | Grade 2 | Grade 3 | Grade 4 | ||
| WT (6/6) | 2 (33.3) | 10 (33.3) | 6 (37.5) | 1 (33.3) | - | 17 (53.1) | 3 (23.1) | 2 (50.0) | P > 0.05 |
| (6/7) | 3 (50.0) | 16 (53.3) | 7 (43.8) | 2 (66.7) | 1 (100) | 14 (43.8) | 7 (53.8) | 2 (50) | P > 0.05 |
| (7/7) | 1 (16.7) | 4 (13.3) | 2 (12.5) | - | - | 1 (3.1) | 2 (15.4) | - | P > 0.05 |
| (5/7) | - | - | 1 (6.3) | - | - | - | 1 (7.7) | - | P > 0.05 |
| WT (6/6) | (6/7, 7/7) | χ² test | |||||
| Grade 0 | Grade 1-2 | Grade 3-4 | Grade 0 | Grade 1-2 | Grade 3-4 | ||
| Leucocytes | 69 (61.1) | 43 (38.1) | 1 (0.9) | 102 (52.8) | 85 (44.0) | 6 (3.1) | P > 0.05 |
| Neutropenic fever | 113 (100.0) | - | - | 191 (98.9) | 1 (0.5) | 1 (0.5) | P > 0.05 |
| Anaemia | 30 (26.5) | 80 (70.8) | 3 (2.7) | 47 (24.3) | 142 (73.6) | 4 (2.1) | P > 0.05 |
| Thrombocytes | 89 (78.8) | 23 (20.4) | 1 (0.9) | 157 (81.3) | 36 (18.7) | - | P > 0.05 |
| WT (6/6) | (6/7, 7/7) | χ² test | |||||
| Grade 0 | Grade 1-2 | Grade 3-4 | Grade 0 | Grade 1-2 | Grade 3-4 | ||
| Nausea | 35 (30.9) | 77 (68.1) | 1 (0.9) | 69 (35.8) | 122 (63.2) | 2 (1.0) | P > 0.05 |
| Vomiting | 77 (68.1) | 35 (31.0) | 1 (0.9) | 123 (63.7) | 69 (35.8) | 1 (0.5) | P > 0.05 |
| Diarrhoea early | 81 (71.7) | 30 (26.5) | 2 (1.8) | 149 (77.2) | 42 (21.8) | 2 (1.0) | P > 0.05 |
| Diarrhoea delayed | 51 (45.1) | 55 (48.7) | 7 (6.2) | 78 (40.4) | 90 (46.6) | 25 (13.0) | P > 0.05 |
| Mucositis | 97 (85.8) | 16 (14.2) | - | 160 (82.9) | 31 (16.1) | 2 (1.0) | P > 0.05 |
| WT (6/6) | (6/7, 7/7) | χ² test | |
| Dose reduction | 22 (19.3) | 48 (25.1) | P > 0.05 |
| Dose delayed | 31 (27.2) | 41 (21.5) | P > 0.05 |
Irinotecan (Camptosar®, CPT-11, Pfizer Oncology, New York, NY, US) is one of the most effective chemotherapeutic agents in the treatment of metastatic colorectal cancer (mCRC)[1,2]. Data from phase III studies indicated an improved clinical outcome of patients who had received an irinotecan-based regimen when compared to those who had received 5-FU/LV alone. As reported by Saltz et al[3], the irinotecan-based combination therapy not only improved the response rate (39% vs 21%), but also the progression free (PFS) (7.0 vs 4.3 mo; P = 0.004) and overall survival (OS) (14.8 vs 12.6 mo; P = 0.04).
Irinotecan is a semisynthetic derivative of camptothecin and acts as an inhibitor of intracellular topoisomerase-I[4]. In vivo, the pro-drug is metabolized by carboxylesterase into its active metabolite SN-38. SN-38 is inactivated by uridine disphosphate glucuronosyl transferase 1 (UGT1A1) into SN-38G which is excreted with bile[5].
The most common side effects of irinotecan include neutropenia, febrile neutropenia, nausea, alopecia and delayed diarrhoea which particularly often represents the main and dose-limiting toxicity. This major side effect is because betaglucuronidase in the bowel re-activates SN-38G into the active metabolite SN-38[6-8].
UGT1A1, an essential enzyme for the inactivation of SN-38, is also involved in the metabolism of bilirubin. Changes in the metabolism of bilirubin result in several clinical disorders. They range from the clinically harmless condition of mild jaundice to the deadly disease Crigler-Najjar that may have a lethal outcome in adolescence[9]. Patients with Gilbert’s syndrome present with a mild unconjugated hyperbilirubinaemia without any structural liver disease or haemolysis. The activity of UGT1A1 is reduced in these individuals compared to those without a comparable defect[10,11].
Molecular analyses have revealed that in the Caucasian population Gilbert’s syndrome is most commonly caused by a polymorphism in the UGT1A1 gene[12]. It consists of a TA insertion in the TATAA element of the 5’-promotor region. Genotypes are defined (6/6), (6/7) and (7/7) according to the number of TA repeats. Therefore patients with the wild type (WT) genotype (6/6) (33% in the Caucasian population) are homozygous with 6 repeats of the TA insertion. Patients with the (7/7) genotype are homozygous with 7 TA repeats while the heterozygous genotype (6/7) consists of 1 allele with 6 TA repeats and of 1 with 7 TA repeats. Patients being heterozygous or homozygous for this variant allele (also named UGT1A1*28) show a reduced expression of the UGT1A1 enzyme resulting in lower rates of bilirubin and SN-38 glucuronidation[13]. Compared to the wild type genotype (6/6) patients carrying the (7/7) genotype displayed a 70% reduction in transcriptional activity and are, by the attenuated expression of UGT1A1, theoretically predisposed to SN-38 associated side effects[14]. Apart from that, there are rare genotypes with less than five or more than seven TA repeats leading to variable enzyme levels.
A number of trials have provided evidence of an association between UGT1A1 gene polymorphism and increased toxicity in patients who received irinotecan[15-18]. Comparability and appliance of the results of these trials is difficult because of the various regimens and irinotecan dosages used. Due to the widespread use of irinotecan and the associated risk of severe side effects the question of defining subgroups of patients susceptible to irinotecan-related toxicities is of eminent clinical importance.
This retrospective analysis included a subgroup of 105 patients with mCRC who were treated with an irinotecan-based chemotherapy within a large prospective randomized multicenter phase III study which investigated the role of a low-dose irinotecan-based chemotherapy in patients with metastatic or advanced colorectal cancer (FIRE-trial)[19]. Patients underwent UGT1A1 genotyping in order to evaluate UGT1A1 as a predictor for drug efficacy and/or toxicity for patients with low-dose irinotecan-based chemotherapy and to provide a future tool for a patient tailored chemotherapy.
One hundred and five Caucasian patients with mCRC were included in this analysis. The study population represents a subgroup of patients treated within the FIRE-trial. In the FIRE-trial a total of 492 patients from 56 centres were included, of which 478 patients could be evaluated for efficacy and toxicity. After an amendment UGT1A1 genotyping was offered and 105 patients were included in this subgroup analysis within the ongoing main study. The local ethics committees of the participating centres approved both the study protocol of the FIRE-trial and the amendment for UGT1A1 genotyping. Since the trial was a multicentre trial, monitoring for consistency was done by an external monitoring expert (ClinAssess GmbH, Leverkusen, Germany). Patients gave written informed consent prior to any study-specific procedures.
Patients with known Gilbert’s syndrome or known DPD-deficiency were not allowed to enter the trial.
Patients within the FIRE-trial were randomly assigned to a modified FOLFIRI (mFOLFIRI) or modified IROX (mIROX) protocol. Randomisation was done by the external monitoring expert after an eligibility check. Inclusion criteria included serum bilirubin ≤ 1.25 or ≤ 1.5 of the upper institutional limits without or with hepatic metastasis. The modified FOLFIRI consisted of irinotecan 80 mg/m2 i.v. over 30 min, folinic acid 500 mg/m2 i.v. over 120 min, followed by 5-FU 2000 mg/m2 i.v. over 24 h weekly for 6 wk. The modified IROX consisted of oxaliplatin 85 mg/m2 i.v. over 120 min every 2 wk and irinotecan 80 mg/m2 i.v. over 30 min weekly. For both arms, treatment cycles were repeated on day 50. Pre-medication with atropine (s.c.) was given routinely to prevent acute anti-cholinergic syndrome. Antiemetics were given according to good clinical practice and the local standards of the participating centres (generally 5-HT3 antagonists). Treatment continuation was intended until progression, until inacceptable toxicity or confirmed complete response (CR).
Therapy was postponed for at least one week until bone marrow recovery or resolution of side effects (diarrhoea ≥ grade 1, mucositis ≥ grade 1, leucocytopenia ≥ grade 2, thrombocytopenia ≥ grade 1 or any other toxicity ≥ grade 2). In case of toxicity, defined as diarrhoea ≥ grade 3, mucositis ≥ grade 3, leucocytopenia grade 4, thrombocytopenia ≥ grade 3, severe obstipation ≥ 96 h or hand-foot-syndrome ≥ grade 3, a dose reduction of irinotecan and 5-FU to 80% was mandatory. Doses of irinotecan and oxaliplatin were reduced to 80% for obstipation ≥ grade 3. Moreover, a dose reduction of oxaliplatin was required in cases of persistent paraesthesia (dose reduction 25%), painful paraesthesia with duration of > 7 d (dose reduction 25%) or paraesthesia with functional impairment with duration of > 7 d (dose reduction 50%).
In case of any toxicity as described above, following cycles were begun at a reduced dose level as defined by the scheme above.
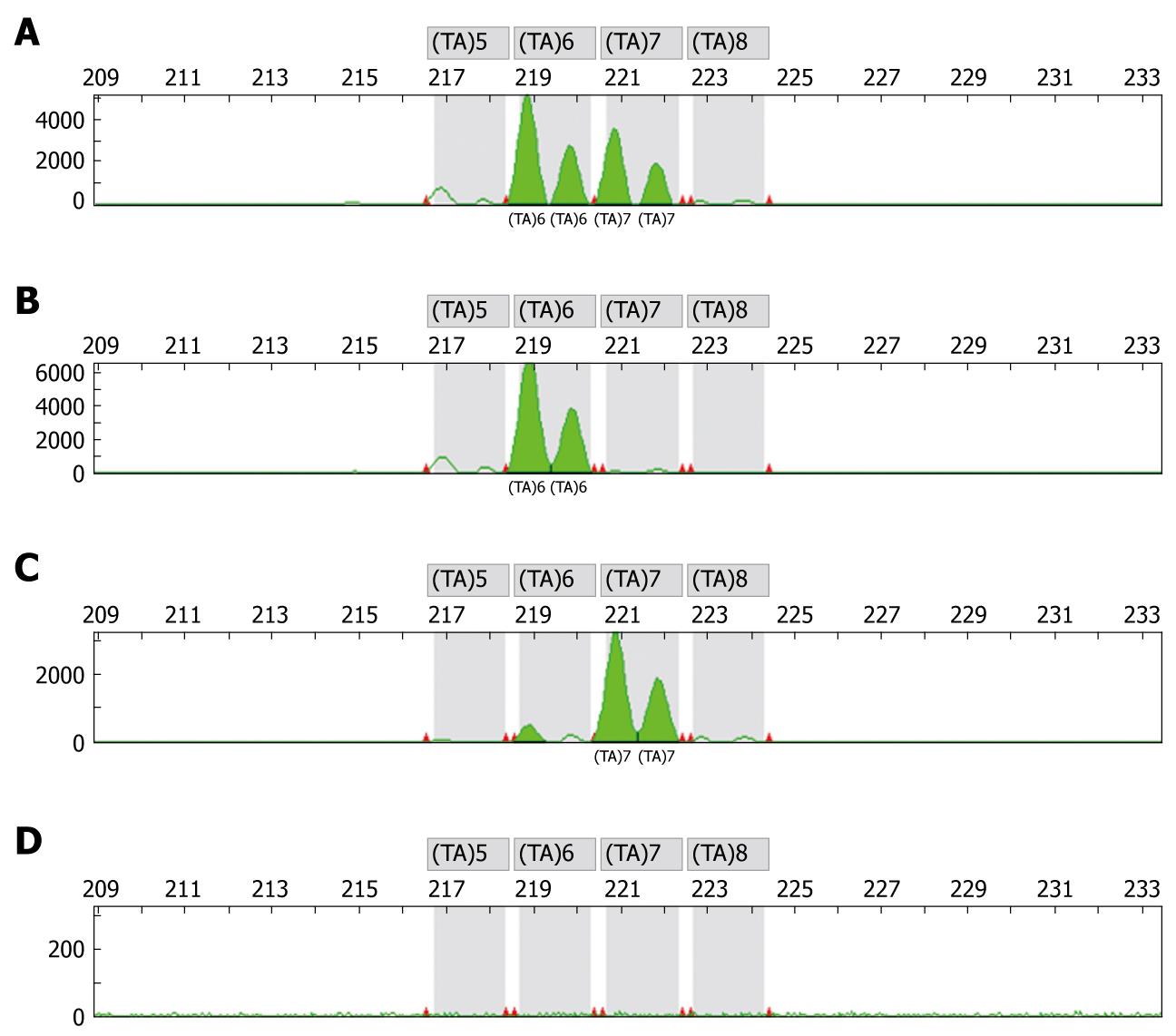
Genotyping studies were performed by an independent laboratory (OncoScreen GmbH, Jena, Germany). Prior to any chemotherapy blood samples were collected for DNA isolation and determination of genotypes. DNA extraction, preparation and genotyping was performed using the methods as previously described[12,14,16]. DNA was extracted from peripheral blood leucocytes using standard methods. A two-extra-nucleotide insertion (TA) within the TATA box resulting in the sequence (TA)7TAA (-39 to 53) was researched. Genotypes were identified by gene scan analysis on the ABI 310 sequencer of the TATAA box in the promoter region of the UGT1A1 gene. Figure 1 displays an electropherogram with typical overlapping peaks to distinguish the WT genotype from the heterozygous and homozygous genotype (Figure 1). The design of the primers to specifically amplify the TA expansion was done with support of GenBank AF297093.
Standard evaluation by history, physical examination and routine laboratory tests (including complete blood count, chemical profile and electrolyte determination) were performed prior to the beginning of each cycle. Drug administration, performance status and toxicity or adverse events were recorded after each cycle of chemotherapy. Toxicity was graded according to the NCI-CTC Classification (version 3.0). Imaging studies using computed tomography (CT) or magnetic resonance imaging (MRI) were performed prior to the beginning of each following cycle as well as for confirmation 4 wk after the end of chemotherapy. Data collection was monitored by an independent monitoring expert (ClinAssess GmbH, Leverkusen, Germany) and was done by an external data manager (Estimate GmbH, Augsburg, Germany).
Patients’ response was assessed by standard WHO criteria, as follows: complete response (CR) was defined as the disappearance of all known disease, documented by at least two observations not less than 4 wk apart, while partial response (PR) was defined as a decrease by at least 50% of the sum of the products of the largest perpendicular diameters of all measurable lesions, as determined by two observations not less than 4 wk apart. Stable disease (SD), lasting for at least 6 wk from the start of the study (i.e. first drug administration), was defined as a < 50% decrease and < 25% increase in the sum of the products of the largest perpendicular diameters of all measurable lesions. Progressive disease (PD) was defined a > 25% increase in the size of at least one bidimensional or unidimensional measurable lesion, or the appearance of a new lesion.
The primary endpoint of the FIRE-trial was time to progression (TTP). Secondary endpoints included response rate (RR), overall survival (OS), resectability rate, toxicity and quality of life. TTP was defined as the interval between the start of therapy and first documentation of disease progression. OS was measured from the date of starting treatment to the date of death from any cause (intent-to-treat). Probability of survival and TTP were estimated using the Kaplan-Meier method[20]. Statistical comparisons between different genotypes were determined by using the χ2-test. Differences in OS and TTP were analyzed using the log-rank test. A difference of P < 0.05 was considered statistically significant.
The median age was 64 years with a gender distribution of 70.5% male and 29.5% female patients. 59% of the patients suffered from metastatic colon and 41% from metastatic rectal cancer. Performance status and adjuvant pre-treatment was similar between WT and heterozygous genotype patients. Detailed patient characteristics are provided in Table 1. Patients in the main trial were stratified according to performance status, lactate dehydrogenase (LDH) and adjuvant pre-treatment. These patients were well balanced between the two treatment arms.
The distribution of the UGT1A1 genotypes was analysed in 105 of 478 patients evaluated within the FIRE-trial. The majority of the patients (49.5%) had the heterozygous genotype (6/7), 39.0% showed the WT genotype (6/6) and the homozygous genotype (7/7) was found in 9.5% of the analyzed patients. There were also two single cases of the rare genotype (5/7) which were not included for further evaluation due to the low frequency. Results of the distribution of the genotypes are given in Table 2.
The overall response rate (ORR = CR + PR) was similar between the WT genotype (6/6) and the (6/7, 7/7) genotypes (43.2% vs 44.3% P = 0.75). However, the disease control rate (DCR = CR + PR + SD) appeared to be lower within patients carrying the WT genotype (6/6) as compared to those patients with the (6/7, 7/7) genotypes (73.0% vs 91.8%, P = 0.008). Detailed response data are presented in Table 3.
There are no efficacy data for the 2 patients carrying the rare genotype (5/7). One patient quit the trial while the other died early during therapy.
Overall toxicity according to genotype and treatment arm is given in Table 4. The incidence of grade 3-4 toxicity did not differ significantly between the treatment arms mFOLFIRI and mIROX both for patients with the WT (6/6) and those carrying the (6/7), (7/7) and (5/7) genotypes.
Haematological toxicity was generally mild (grade 3 and 4 < 5%). Comparing the incidence of grade 1-2 and grade 3-4 haematotoxicity in patients with the WT (6/6) genotype to those with the (6/7, 7/7) genotypes, there were no significant differences regarding leucocytopenia, anaemia or thrombocytopenia (P > 0.05) (Table 5).
Regarding non-haematological toxicity, the incidence of grade 3-4 delayed diarrhoea appeared higher in patients with the (6/7, 7/7) genotypes compared to the WT (6/6) genotype even though this difference did not reach the statistical level of significance (13.0% vs 6.2%, P = 0.08) (Table 6).
Both cases with the rare genotype (5/7) were excluded from the statistical evaluation. However, they experienced grade 3-4 toxicity in the course of the treatment (delayed diarrhoea and leucocytopenia, per cycle analysis).
There were no significant differences regarding toxicity-related treatment discontinuations or dose adjustments. Dose delays in patients with the WT (6/6) were observed in 27.2%, compared to 21.5% in patients with the (6/7, 7/7) genotypes (P = 0.071). Dose reductions were more frequently observed in patients with the (6/7, 7/7) genotypes compared to patients who carried the WT (6/6) genotype (25.1% vs 19.3%, P = 0.24) (Table 7).
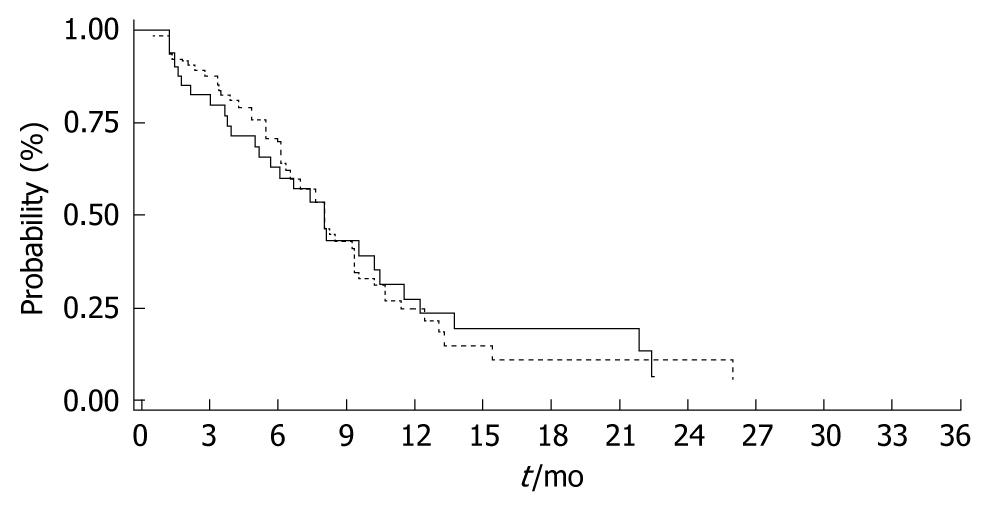
TTP was similar between the WT (6/6) and the (6/7, 7/7) genotypes with 8.1 mo and 8.2 mo respectively (P = 0.971) (Figure 2). Moreover, there was no difference in survival between the WT compared to the genotypes [WT (6/6) vs (6/7, 7/7); 21.2 mo vs 18.9 mo, P = 0.725] (Figure 3).
More than 50 genetic alterations of the UGT1A1 gene locus have previously been described[21]. Among the Caucasian population the UGT1A1*28 genotype (7/7) plays the most important role for the development of Gilbert’s syndrome and an increased toxicity related to irinotecan-based chemotherapy.
The distribution of the UGT1A1 genotype in this retrospective analysis was comparable to that described by Iyer and co-workers[15]. They reported a frequency of the WT (6/6) genotype in 45% of the patients (39% in the present analysis), the (6/7) genotype in 35% (49.5% in the present analysis), and the (7/7) genotype in 20% (9.5% in the present analysis) respectively[15]. Moreover, another analysis of 51 patients suffering from non-small cell lung cancer (NSCLC) showed a comparable distribution of the genotypes WT (6/6) 49%, (6/7) 36% and (7/7) 15%[22].
Prior to discussion of an association of toxicity and UGT1A1 genotype the following preliminary remarks are inevitable: patients randomized within the FIRE-trial received different irinotecan-based regimes. The incidence of delayed diarrhoea, neurotoxicity, leucocytopenia and thrombocytopenia was more frequent in the IROX arm compared to the FOLFIRI arm[19]. Differences in the frequency of adverse events might therefore partly influence the present analysis. Moreover, 7.3% of the patients (mainly with rectal cancer) had previously received locoregional radiotherapy prior to study entry. Local irradiation may also play a role in the development of severe diarrhoea or may at least worsen it, independent of the UGT1A1 genotype[23]. Finally, in analogy to the study of Ando et al[14], there is a certain bias by excluding patients with elevated pre-treatment bilirubin levels.
Among non-haematological toxicities, the incidence of grade 3 and 4 delayed diarrhoea was twice as high in our study population with a homozygous (7/7) or heterozygous (6/7) genotype compared to those carrying the wild type genotype WT (6/6), even though the difference did not reach the level of significance (13.0% vs 6.2%; P = 0.08). Nevertheless, this trend is supported by the data provided by Marcuello et al[23] who found a significant correlation of the UGT1A1 genotype and the frequency of severe delayed diarrhoea [WT (6/6) 17%, (6/7) 33%, (7/7) 70%; P = 0.005]. Moreover, Ando et al[14] reported on a 3.5-fold higher frequency of the UGT1A1*28 genotype in patients who suffered from severe diarrhoea and leucocytopenia during irinotecan-based chemotherapy. Taking the UGT1A1*28 genotype as a significant risk factor for the development of severe toxicity the authors suggest that UGT1A1 genotyping serves as a useful tool in predicting toxicity in patients receiving irinotecan. It is critical to state that the majority of these analyses were done retrospectively. In contrast, an exploratory analysis presented by Seymour et al[24] could not confirm such an association of UGT1A1 to irinotecan-related toxicity. Moreover comparability is hampered by the different doses of irinotecan applied within these trials. The interpretation of our data is limited by a slight imbalance with more patients with the (6/7) and (7/7) genotype being treated with the modified FOLFIRI protocol. Therefore, a tendency towards an increased rate of diarrhoea which might be associated with exposure to 5-FU can not be ruled out.
Comparing the incidence of severe haematotoxicity in patients with the WT (6/6) genotype to those with the (6/7, 7/7) genotype in our analysis, there were no significant differences regarding leucocytopenia, anaemia or thrombocytopenia (P > 0.05). The incidence of haematological toxicity was generally low, as expected from other trials with comparable doses of weekly scheduled irinotecan[25,26]. Nevertheless, these findings are in contrast to previously published results of patients with mCRC treated with IRIFUFOL or FOLFIRI[18]. In the study of Rouits et al[18] grade 3-4 neutropenia was significantly associated with the genotypes (6/7, 7/7). The results of a retrospective analysis of 128 Chinese patients with mCRC who received biweekly irinotecan indicate that the heterogenous and homogenous genotype UGT1A1 (6/7) and (7/7) predicts severe neutropenia and diarrhoea but not treatment efficacy[27]. Necessity for dose reduction was significantly associated with the (6/7) or (7/7) genotype (42.3% vs 12.7%; P < 0.01).
Moreover, Innocenti et al[16] also observed no case of grade 4 neutropenia among patients with the wild type genotype WT (6/6), whereas 50% among patients with the homozygous genotype (7/7), and 12.5% of those with the heterozygous genotype (6/7) experienced grade 4 neutropenia. In their study irinotecan was administered at a dose of 350 mg/m2 every 3 wk. Accordingly, data from Roth et al[28] indicated that the risk of severe neutropenia was higher among patients carrying the homogenous genotype (7/7). Interestingly, female sex was superior to UGT1A1 in predicting grade 4 neutropenia. Contrary, Marcuello et al[23] found an increased, but not significantly increased, haematological toxicity among patients with the genotypes (6/7, 7/7) during single-agent or combination chemotherapy with irinotecan given at a dosage of 350 mg/m2 every 3 wk or 180 mg/m2 every 2 wk.
The literature regarding UGT1A1 genotyping and irinotecan-related toxicity is heterogeneous and to some extent conflicting which may be partly explained by the retrospective character of most studies[14,16,18,23,24]. Another aspect contributing to the heterogeneity of the results is due to the variety of irinotecan dosages and schedules which also applies for the present study. Patients within our trial received dose reduced and modified FOLFIRI and IROX. Patients in both treatment arms received weekly scheduled irinotecan (80 mg/m2) in combination with 5-FU 2000 mg/m2 weekly or oxaliplatin 85 mg/m2 biweekly. We assume that the mild toxicity and the absent impact of UGT1A1 genotype on toxicity in our study are a result of the low irinotecan dose. Stewart et al[29] concluded from their study of low-dose protracted irinotecan in pediatric patients that UGT1A1 genotyping is not a useful prognostic factor in predicting toxicity. Hoskins et al[30] advised a genotype-tailored therapy only for those patients who receive irinotecan at higher doses. Patients carrying the genotype UGT1A1*28 who receive irinotecan up to a dose of 150 mg/m2 are at the same risk of experiencing severe neutropenia as any patients. Consequently, UGT1A1 genotyping is not generally recommended.
The response rate in the present analysis was similar between the WT (6/6) genotype and the (6/7, 7/7) genotypes (43.2% vs 44.3%, P > 0.05). Interestingly, the disease control rate appeared to be higher in patients with the (6/7, 7/7) genotypes compared to those with the WT (6/6) genotype (91.8% vs 73.0%, P = 0.008). However, this finding did not result in any advantage regarding TTP or OS.
Douillard et al[31] reported a TTP of 6.7 and an OS of 17.4 mo in previously untreated patients with mCRC who had received irinotecan and FU/FA. A large Italian study reported a comparable TTP of 7 mo and an OS of 14 mo for the FOLFIRI regimen given as 1st line chemotherapy for mCRC[32]. Ashley et al[33] have found a TTP of 6.7 mo and an overall survival of 17.3 mo in previously untreated patients with mCRC who had received an IROX regimen consisted of oxaliplatin 85 mg/m2 and irinotecan 200 mg/m2 every 3 wk. These data are supported by the findings of Goldberg et al[34] who found a similar TTP of 6.5 mo and an overall survival of 17.4 mo for patients who were treated with IROX. Comparing these data to those of our study a loss of activity due to the dose modifications with low-dose irinotecan dosage can be ruled out.
In a large prospective study conducted by Toffoli et al[35], the response rate was higher in mCRC patients with the homozygous genotype (7/7) compared to the wild type genotype WT (6/6). These patients experienced a slightly improved survival of approximately 2 mo (P > 0.05). One may argue that in patients with the (6/7) or (7/7) genotype there is less detoxification of SN-38 resulting in higher blood levels of the active compound, more anti-tumor effect and therefore a better response rate. However, Marcuello et al[23] observed no statistically significant impact of UGT1A1 gene polymorphism on response rate but a trend towards an improved OS in patients with the WT (6/6) genotype compared to patients with the (6/7) or (7/7) genotype (33 vs 21 mo; P = 0.09). This is partly explained by dose reductions in patients with homozygous or heterozygous genotypes (6/7, 7/7) because of severe diarrhoea. Another recent study reported on prospectively genotyped patients suffering from mCRC who had received either 1st line irinotecan/capecitabine or 2nd line single-agent irinotecan. Response rates, numbers of dose reductions and applied chemotherapy cycles were similar within the different genotypes[36].
An important factor contributing to irinotecan metabolism is the existence of several UGT1A1 isoforms and their distribution among different patient populations[37]. There exist an increasing number of reports on genetic variants of UGT1A1 as well as SNPs in the coding region of the gene locus potentially influencing drug metabolism. Moreover, other enzymes of the UGT1 family like UGT1A7 and UGT1A9 are also involved in the glucuronidation of SN-38[38-40]. Patients of African, Caucasian and Asian descent show a different gene frequency of the UGT1A1*28 gene variant[12]. When analysing UGT1A1 genotypes and irinotecan-related adverse events the different variants of the UGT1A1 gene locus among an ethnic population must be taken into consideration[41]. Moreover, there is strong evidence that several other individual factors apart from ethnic affiliation may influence the irinotecan metabolism[42].
In conclusion, UGT1A1 genotyping alone does not allow characterizing subgroups of patients who are at an increased risk of life threatening toxicity during low-dose irinotecan-based chemotherapy. Due to the incoherent findings of several small trials, a larger prospective phase III trial is warranted. The metabolism of irinotecan is highly complex with different UGT enzymes and drug transporters involved. Distinction between high-dose and low-dose irinotecan is of eminent importance when considering the use of UGT1A1 genotyping. There is a need for a diagnostic panel including the testing of multiple gene polymorphisms that more reliably predicts toxicity. On the other hand, beside a comprehensive patient education, and the escalation of supportive therapy, dose modifications and alternative treatment schedules may help to provide patients who are at high risk with a safe irinotecan-based chemotherapy.
Irinotecan is one of the most effective chemotherapeutic agents in the treatment of colorectal cancer. Among other side effects irinotecan can lead to neutropenia and delayed diarrhoea. The active metabolite of irinotecan, SN-38, is inactivated by uridine disphosphate glucoronosyl transferase 1 (UGT1A1), the same enzyme by which bilirubin is metabolised. Genetic polymorphisms of the UGT1A1 gene result in variable levels of the enzyme. According to the number of TA repeats in the enhancer region of the gene a wild type genotype (6/6) can be differentiated from the heterozygous genotype (6/7) and the homozygous genotype (7/7).
Preclinical and clinical studies have revealed a close dependency of the activity of UGT1A1 enzyme and the occurrence of irinotecan-associated side effects. Existing data are contradictory to some extent or are derived from heterogenous or small study populations. Therefore and against the background of other metabolic ways of toxification and detoxification the research hotspot is how to apply genetic testing of UGT1A1 gene polymorphisms to predict toxicity and efficacy of an irinotecan-based chemotherapy.
Recent studies have emphasized the impact of UGT1A1 genotyping to predict toxicity and outcome in patients undergoing an irinotecan-based chemotherapy. Distribution of the genotype in this study was well in line with data in the literature. No significant differences could be noticed between the homozygous and the heterozygous genotype compared to the wild type genotype in terms of efficacy, toxicity of higher grades, treatment delay or dose reduction. This is a retrospective study to report that UGT1A1 genotyping appeared not to be useful for predicting treatment efficacy and irinotecan-associated side effects in patients receiving low-dose irinotecan for mCRC. Furthermore, our analysis supports the idea that UGT1A1 genotyping should be considered when using irinotecan at a higher dosage.
By understanding the limitations of UGT1A1 genotyping and by understanding the complexity of irinotecan metabolism, this study may just add another piece of the puzzle for the future development of a patient-tailored chemotherapy with genotyping as one tool among others.
The chemotherapeutic agent irinotecan is metabolised by the enzyme UGT1A1, the same enzyme by which bilirubin is metabolised. Genetic polymorphisms resulting in a decreased amount of the enzyme can lead to enhanced irinotecan-associated toxicity. Pre-existing data have partly shown genetic subgroups of patients with different toxicity and treatment efficacy under chemotherapy with irinotecan.
The manuscript by Schulz et al describes the impact of UGT1A1 gene polymorphism on toxicity and efficacy of irinotecan-based regimens in metastatic colorectal cancer. The authors demonstrate that the genetic polymorphism of the UGT1A1 gene does not influence treatment efficacy. The manuscript is well written, the methods are adequately chosen.
Peer reviewers: Ulrike S Stein, PhD, Assistant Professor, Max-Delbrück-Center for Molecular Medicine, Robert-Rössle-Straße 10, 13125 Berlin, Germany; Tamara Cacev, MSc, Division of Molecular Medicine, Rudjer Boskovic Institute, Bijenicka c. 54, Zagreb 10000, Croatia
S- Editor Tian L L- Editor O'Neill M E- Editor Lin YP
| 1. | Garcia-Carbonero R, Supko JG. Current perspectives on the clinical experience, pharmacology, and continued development of the camptothecins. Clin Cancer Res. 2002;8:641-661. [Cited in This Article: ] |
| 2. | Ulukan H, Swaan PW. Camptothecins: a review of their chemotherapeutic potential. Drugs. 2002;62:2039-2057. [Cited in This Article: ] |
| 3. | Saltz LB, Cox JV, Blanke C, Rosen LS, Fehrenbacher L, Moore MJ, Maroun JA, Ackland SP, Locker PK, Pirotta N. Irinotecan plus fluorouracil and leucovorin for metastatic colorectal cancer. Irinotecan Study Group. N Engl J Med. 2000;343:905-914. [Cited in This Article: ] |
| 4. | Folprecht G, Köhne CH. The role of new agents in the treatment of colorectal cancer. Oncology. 2004;66:1-17. [Cited in This Article: ] |
| 5. | Kawato Y, Aonuma M, Hirota Y, Kuga H, Sato K. Intracellular roles of SN-38, a metabolite of the camptothecin derivative CPT-11, in the antitumor effect of CPT-11. Cancer Res. 1991;51:4187-4191. [Cited in This Article: ] |
| 6. | Vanhoefer U, Harstrick A, Achterrath W, Cao S, Seeber S, Rustum YM. Irinotecan in the treatment of colorectal cancer: clinical overview. J Clin Oncol. 2001;19:1501-1518. [Cited in This Article: ] |
| 7. | Araki E, Ishikawa M, Iigo M, Koide T, Itabashi M, Hoshi A. Relationship between development of diarrhea and the concentration of SN-38, an active metabolite of CPT-11, in the intestine and the blood plasma of athymic mice following intraperitoneal administration of CPT-11. Jpn J Cancer Res. 1993;84:697-702. [Cited in This Article: ] |
| 8. | Gupta E, Lestingi TM, Mick R, Ramirez J, Vokes EE, Ratain MJ. Metabolic fate of irinotecan in humans: correlation of glucuronidation with diarrhea. Cancer Res. 1994;54:3723-3725. [Cited in This Article: ] |
| 9. | Bosma PJ. Inherited disorders of bilirubin metabolism. J Hepatol. 2003;38:107-117. [Cited in This Article: ] |
| 10. | Monaghan G, Ryan M, Seddon R, Hume R, Burchell B. Genetic variation in bilirubin UPD-glucuronosyltransferase gene promoter and Gilbert's syndrome. Lancet. 1996;347:578-581. [Cited in This Article: ] |
| 11. | Miners JO, McKinnon RA, Mackenzie PI. Genetic polymorphisms of UDP-glucuronosyltransferases and their functional significance. Toxicology. 2002;181-182:453-456. [Cited in This Article: ] |
| 12. | Beutler E, Gelbart T, Demina A. Racial variability in the UDP-glucuronosyltransferase 1 (UGT1A1) promoter: a balanced polymorphism for regulation of bilirubin metabolism? Proc Natl Acad Sci USA. 1998;95:8170-8174. [Cited in This Article: ] |
| 13. | Iyer L, Hall D, Das S, Mortell MA, Ramírez J, Kim S, Di Rienzo A, Ratain MJ. Phenotype-genotype correlation of in vitro SN-38 (active metabolite of irinotecan) and bilirubin glucuronidation in human liver tissue with UGT1A1 promoter polymorphism. Clin Pharmacol Ther. 1999;65:576-582. [Cited in This Article: ] |
| 14. | Ando Y, Saka H, Ando M, Sawa T, Muro K, Ueoka H, Yokoyama A, Saitoh S, Shimokata K, Hasegawa Y. Polymorphisms of UDP-glucuronosyltransferase gene and irinotecan toxicity: a pharmacogenetic analysis. Cancer Res. 2000;60:6921-6926. [Cited in This Article: ] |
| 15. | Iyer L, Das S, Janisch L, Wen M, Ramírez J, Karrison T, Fleming GF, Vokes EE, Schilsky RL, Ratain MJ. UGT1A1*28 polymorphism as a determinant of irinotecan disposition and toxicity. Pharmacogenomics J. 2002;2:43-47. [Cited in This Article: ] |
| 16. | Innocenti F, Undevia SD, Iyer L, Chen PX, Das S, Kocherginsky M, Karrison T, Janisch L, Ramírez J, Rudin CM. Genetic variants in the UDP-glucuronosyltransferase 1A1 gene predict the risk of severe neutropenia of irinotecan. J Clin Oncol. 2004;22:1382-1388. [Cited in This Article: ] |
| 17. | Côté JF, Kirzin S, Kramar A, Mosnier JF, Diebold MD, Soubeyran I, Thirouard AS, Selves J, Laurent-Puig P, Ychou M. UGT1A1 polymorphism can predict hematologic toxicity in patients treated with irinotecan. Clin Cancer Res. 2007;13:3269-3275. [Cited in This Article: ] |
| 18. | Rouits E, Boisdron-Celle M, Dumont A, Guérin O, Morel A, Gamelin E. Relevance of different UGT1A1 polymorphisms in irinotecan-induced toxicity: a molecular and clinical study of 75 patients. Clin Cancer Res. 2004;10:5151-5159. [Cited in This Article: ] |
| 19. | Schalhorn A, Fischer von Weikersthal L, Quietzsch D, Maubach P, Oruzio D, Lambertz H, Weigang-Koehler K, Schulze M, Schlag R, Heinemann V. Phase III trial of irinotecan plus oxaliplatin (IROX) versus irinotecan plus 5-FU/folinic acid (FOLFIRI) as first-line treatment of metastatic colorectal cancer (CRC): The FIRE-TRIAL. ASCO Annual Meeting Proceedings, 2005: Abstract No. 3516. . [Cited in This Article: ] |
| 20. | Kaplan EL, Meier P. Nonparametric estimation from incomplete observations. J Am Stat Assoc. 1959;53:457-481. [Cited in This Article: ] |
| 21. | Kadakol A, Ghosh SS, Sappal BS, Sharma G, Chowdhury JR, Chowdhury NR. Genetic lesions of bilirubin uridine-diphosphoglucuronate glucuronosyltransferase (UGT1A1) causing Crigler-Najjar and Gilbert syndromes: correlation of genotype to phenotype. Hum Mutat. 2000;16:297-306. [Cited in This Article: ] |
| 22. | Font A, Sánchez JM, Tarón M, Martinez-Balibrea E, Sánchez JJ, Manzano JL, Margelí M, Richardet M, Barnadas A, Abad A. Weekly regimen of irinotecan/docetaxel in previously treated non-small cell lung cancer patients and correlation with uridine diphosphate glucuronosyltransferase 1A1 (UGT1A1) polymorphism. Invest New Drugs. 2003;21:435-443. [Cited in This Article: ] |
| 23. | Marcuello E, Altés A, Menoyo A, Del Rio E, Gómez-Pardo M, Baiget M. UGT1A1 gene variations and irinotecan treatment in patients with metastatic colorectal cancer. Br J Cancer. 2004;91:678-682. [Cited in This Article: ] |
| 24. | Seymour MT, Braun MS, Richman SD, Daly C, Thompson LC, Meade A, Parmar M, Allan JM, Selby P, Quirke P. Association of molecular markers with toxicity outcomes in a randomized trial of chemotherapy for advanced colorectal cancer (FOCUS). ASCO Annual Meeting Proceedings Part I (Abstract No. 2022). J Clin Oncol. 2006;24:18S. [Cited in This Article: ] |
| 25. | Carlini LE, Meropol NJ, Bever J, Andria ML, Hill T, Gold P, Rogatko A, Wang H, Blanchard RL. UGT1A7 and UGT1A9 polymorphisms predict response and toxicity in colorectal cancer patients treated with capecitabine/irinotecan. Clin Cancer Res. 2005;11:1226-1236. [Cited in This Article: ] |
| 26. | Massacesi C, Terrazzino S, Marcucci F, Rocchi MB, Lippe P, Bisonni R, Lombardo M, Pilone A, Mattioli R, Leon A. Uridine diphosphate glucuronosyl transferase 1A1 promoter polymorphism predicts the risk of gastrointestinal toxicity and fatigue induced by irinotecan-based chemotherapy. Cancer. 2006;106:1007-1016. [Cited in This Article: ] |
| 27. | Liu CY, Chen PM, Chiou TJ, Liu JH, Lin JK, Lin TC, Chen WS, Jiang JK, Wang HS, Wang WS. UGT1A1*28 polymorphism predicts irinotecan-induced severe toxicities without affecting treatment outcome and survival in patients with metastatic colorectal carcinoma. Cancer. 2008;112:1932-1940. [Cited in This Article: ] |
| 28. | Roth AD, Yan P, Dietrich D, Fiocca R, Bodoky G, Labianca R, Cunningham D, Van Cutsem E, Bosman F, Tejpar S. Is the UGT1A*28 homozygosity the strongest predictor for severe hematoxicity in patients with 5-fluorouracil (5-FU)-irinotecan (IRI)? Results of the PETACC 3 - EORTC 40993 - SAKK 60/00 trial comparing IRI/5-FU/folinic acid (FA) to 5-FU/FA in stage II - III colon cancer (COC) patients (Abstract No. 4036). J Clin Oncol. 2008;112:26. [Cited in This Article: ] |
| 29. | Stewart CF, Panetta JC, O'Shaughnessy MA, Throm SL, Fraga CH, Owens T, Liu T, Billups C, Rodriguez-Galindo C, Gajjar A. UGT1A1 promoter genotype correlates with SN-38 pharmacokinetics, but not severe toxicity in patients receiving low-dose irinotecan. J Clin Oncol. 2007;25:2594-2600. [Cited in This Article: ] |
| 30. | Hoskins JM, Goldberg RM, Qu P, Ibrahim JG, McLeod HL. UGT1A1*28 genotype and irinotecan-induced neutropenia: dose matters. J Natl Cancer Inst. 2007;99:1290-1295. [Cited in This Article: ] |
| 31. | Douillard JY, Cunningham D, Roth AD, Navarro M, James RD, Karasek P, Jandik P, Iveson T, Carmichael J, Alakl M. Irinotecan combined with fluorouracil compared with fluorouracil alone as first-line treatment for metastatic colorectal cancer: a multicentre randomised trial. Lancet. 2000;355:1041-1047. [Cited in This Article: ] |
| 32. | Colucci G, Gebbia V, Paoletti G, Giuliani F, Caruso M, Gebbia N, Cartenì G, Agostara B, Pezzella G, Manzione L. Phase III randomized trial of FOLFIRI versus FOLFOX4 in the treatment of advanced colorectal cancer: a multicenter study of the Gruppo Oncologico Dell'Italia Meridionale. J Clin Oncol. 2005;23:4866-4875. [Cited in This Article: ] |
| 33. | Ashley AC, Sargent DJ, Alberts SR, Grothey A, Campbell ME, Morton RF, Fuchs CS, Ramanathan RK, Williamson SK, Findlay BP. Updated efficacy and toxicity analysis of irinotecan and oxaliplatin (IROX) : intergroup trial N9741 in first-line treatment of metastatic colorectal cancer. Cancer. 2007;110:670-677. [Cited in This Article: ] |
| 34. | Goldberg RM, Sargent DJ, Morton RF, Fuchs CS, Ramanathan RK, Williamson SK, Findlay BP, Pitot HC, Alberts SR. A randomized controlled trial of fluorouracil plus leucovorin, irinotecan, and oxaliplatin combinations in patients with previously untreated metastatic colorectal cancer. J Clin Oncol. 2004;22:23-30. [Cited in This Article: ] |
| 35. | Toffoli G, Cecchin E, Corona G, Russo A, Buonadonna A, D'Andrea M, Pasetto LM, Pessa S, Errante D, De Pangher V. The role of UGT1A1*28 polymorphism in the pharmacodynamics and pharmacokinetics of irinotecan in patients with metastatic colorectal cancer. J Clin Oncol. 2006;24:3061-3068. [Cited in This Article: ] |
| 36. | Kweekel DM, Gelderblom H, Van der Straaten T, Antonini NF, Punt CJ, Guchelaar HJ. UGT1A1*28 genotype and irinotecan dosage in patients with metastatic colorectal cancer: a Dutch Colorectal Cancer Group study. Br J Cancer. 2008;99:275-282. [Cited in This Article: ] |
| 37. | Mercke Odeberg J, Andrade J, Holmberg K, Hoglund P, Malmqvist U, Odeberg J. UGT1A polymorphisms in a Swedish cohort and a human diversity panel, and the relation to bilirubin plasma levels in males and females. Eur J Clin Pharmacol. 2006;62:829-837. [Cited in This Article: ] |
| 38. | Iyer L, Ratain MJ. Clinical pharmacology of camptothecins. Cancer Chemother Pharmacol. 1998;42 Suppl:S31-S43. [Cited in This Article: ] |
| 39. | Ciotti M, Basu N, Brangi M, Owens IS. Glucuronidation of 7-ethyl-10-hydroxycamptothecin (SN-38) by the human UDP-glucuronosyltransferases encoded at the UGT1 locus. Biochem Biophys Res Commun. 1999;260:199-202. [Cited in This Article: ] |
| 40. | Tukey RH, Strassburg CP, Mackenzie PI. Pharmacogenomics of human UDP-glucuronosyltransferases and irinotecan toxicity. Mol Pharmacol. 2002;62:446-450. [Cited in This Article: ] |
| 41. | Zhang A, Xing Q, Qin S, Du J, Wang L, Yu L, Li X, Xu L, Xu M, Feng G. Intra-ethnic differences in genetic variants of the UGT-glucuronosyltransferase 1A1 gene in Chinese populations. Pharmacogenomics J. 2007;7:333-338. [Cited in This Article: ] |
| 42. | Nagar S, Blanchard RL. Pharmacogenetics of uridine diphosphoglucuronosyltransferase (UGT) 1A family members and its role in patient response to irinotecan. Drug Metab Rev. 2006;38:393-409. [Cited in This Article: ] |